We narrowed to 14,618 results for: RING
-
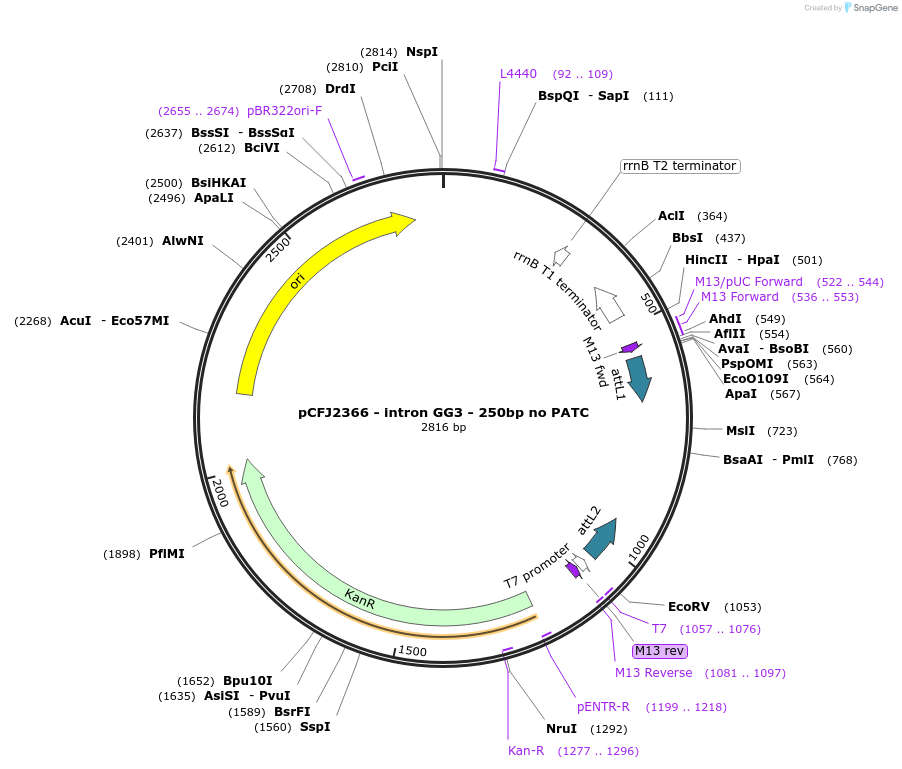
Plasmid#159885PurposeGolden Gate compatible intron with no PATCs for negative controlsDepositorInsertintron GG3 - 250bp no PATC
ExpressionWormMutationNot applicableAvailable SinceDec. 17, 2020AvailabilityAcademic Institutions and Nonprofits only -
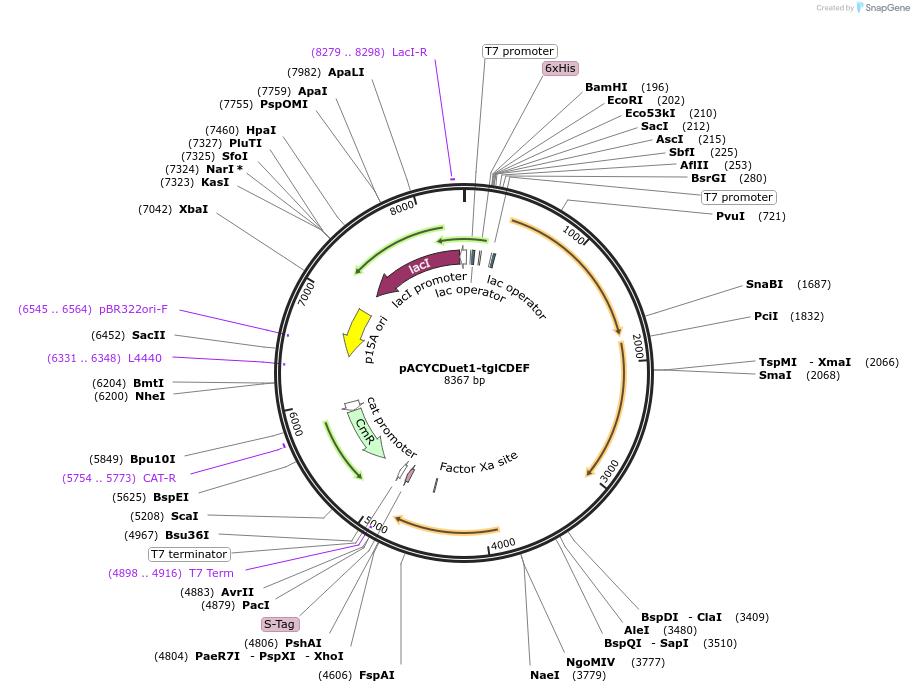
pACYCDuet1-tglCDEF
Plasmid#134677PurposeExpresses TglC, TglD, TglE, TglF in E. ColiDepositorInsertsTglC
TglD
TglE
TglF
ExpressionBacterialPromoterT7Available SinceJuly 17, 2020AvailabilityAcademic Institutions and Nonprofits only -
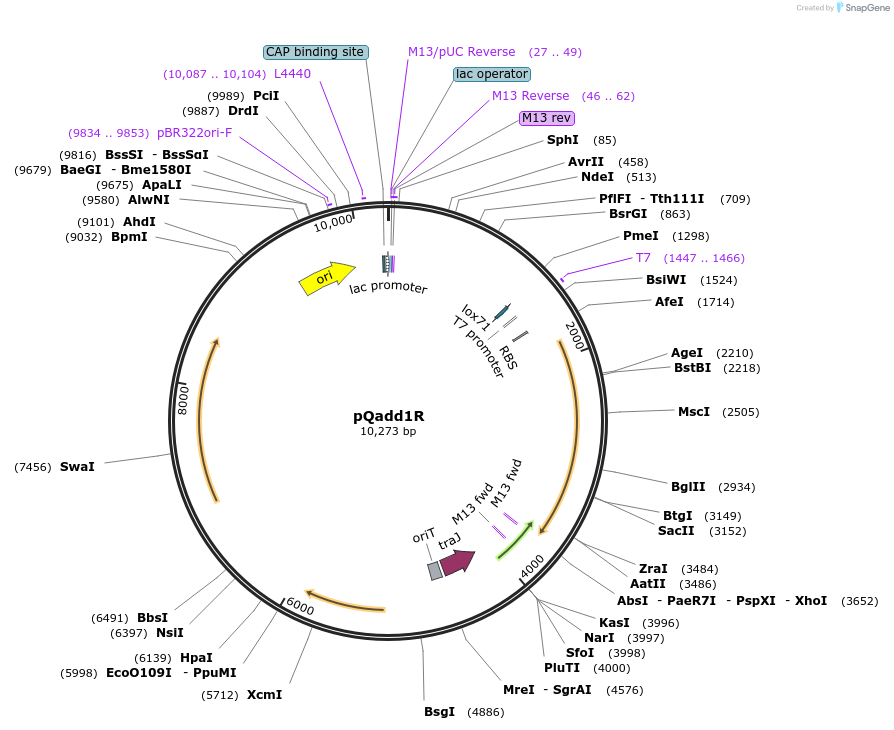
pQadd1R
Plasmid#135661PurposeClostridial vector, encoding a targetron carrying two lox511/71 and loxFAS/66 sites in tandem, in the reverse (antisense) orientation. Retarget intron to insert double-lox site at desired location.DepositorInsertIntron containing lox site
UseCre/LoxExpressionBacterialPromoterpFerAvailable SinceJuly 15, 2020AvailabilityAcademic Institutions and Nonprofits only -
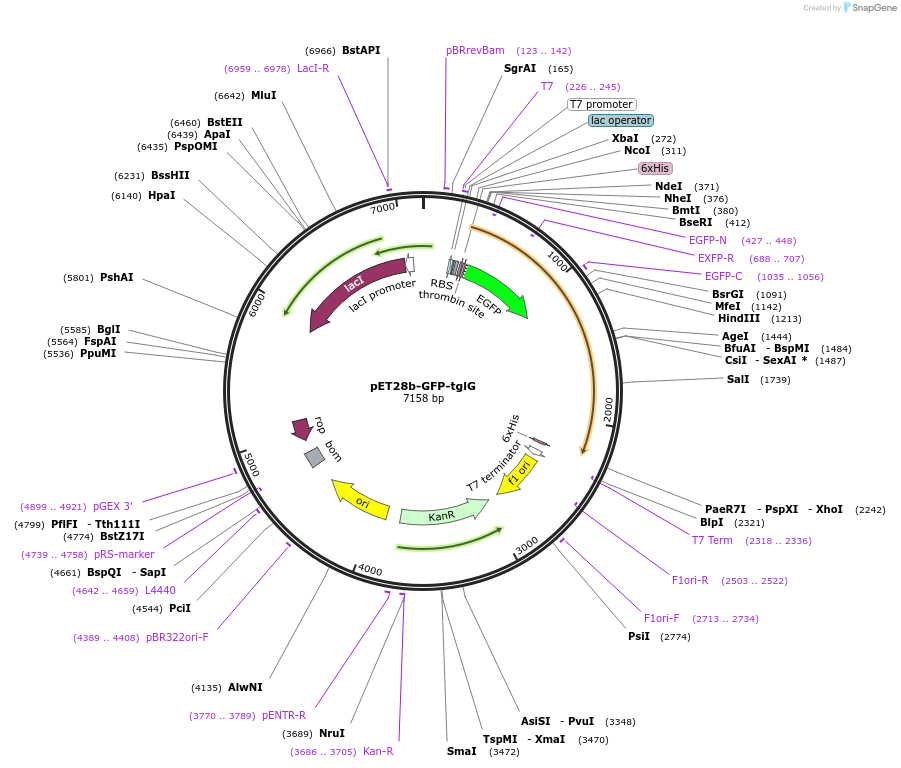
pET28b-GFP-tglG
Plasmid#134679PurposeExpresses His6-GFP-TglG fusion in E. ColiDepositorInsertTglG
TagsHis6-GFPExpressionBacterialPromoterT7Available SinceJuly 14, 2020AvailabilityAcademic Institutions and Nonprofits only -
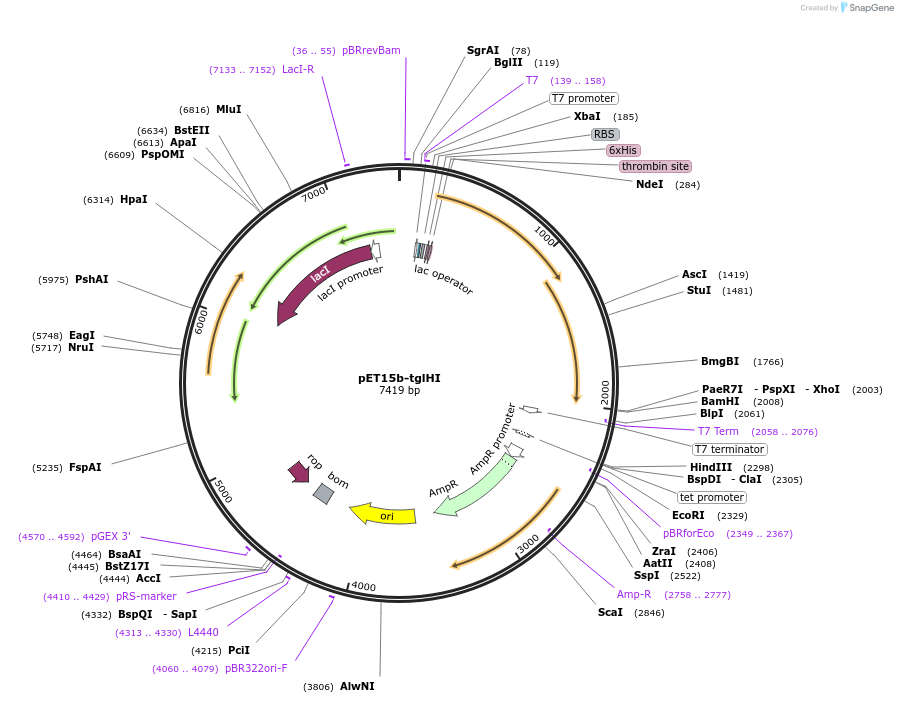
pET15b-tglHI
Plasmid#134672PurposeExpresses His6-TglH and TglI in E. ColiDepositorInsertsTglH
TglI
TagsHis6ExpressionBacterialPromoterT7Available SinceJuly 9, 2020AvailabilityAcademic Institutions and Nonprofits only -
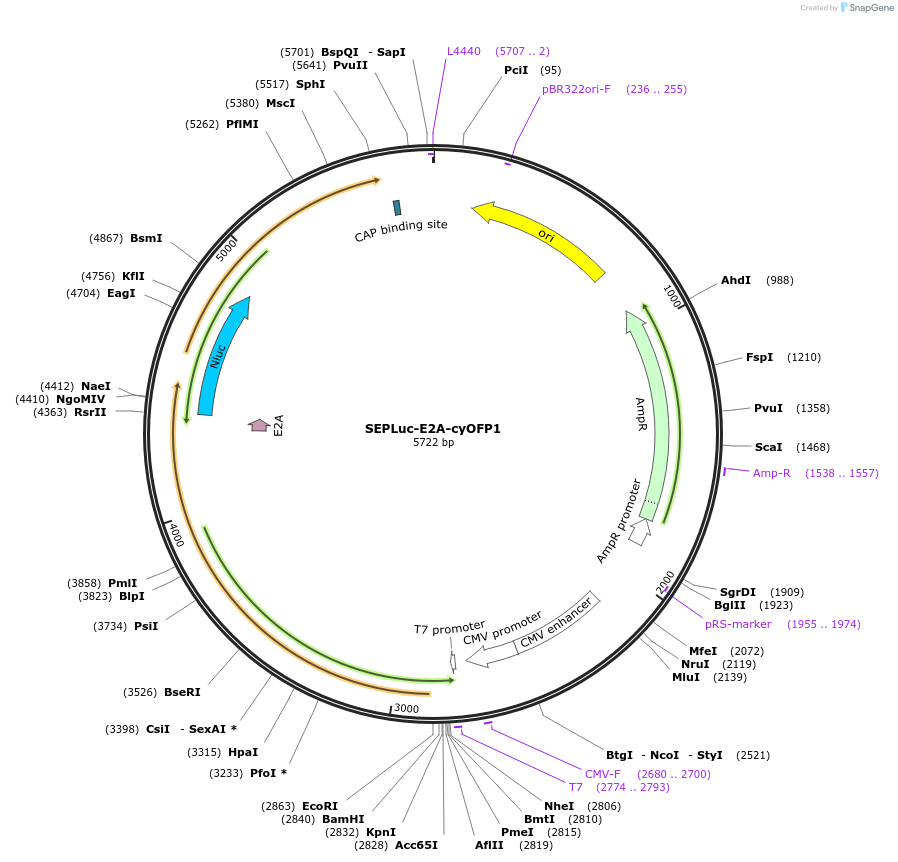
SEPLuc-E2A-cyOFP1
Plasmid#141084Purposeprecursor of the bioluminescent reporter pHLuc, for detecting extracellular pH changes in tumor acidosisDepositorInsertSEPLuc-E2A-cyOFP1
UseLentiviralPromoterCMVAvailable SinceJune 25, 2020AvailabilityAcademic Institutions and Nonprofits only -
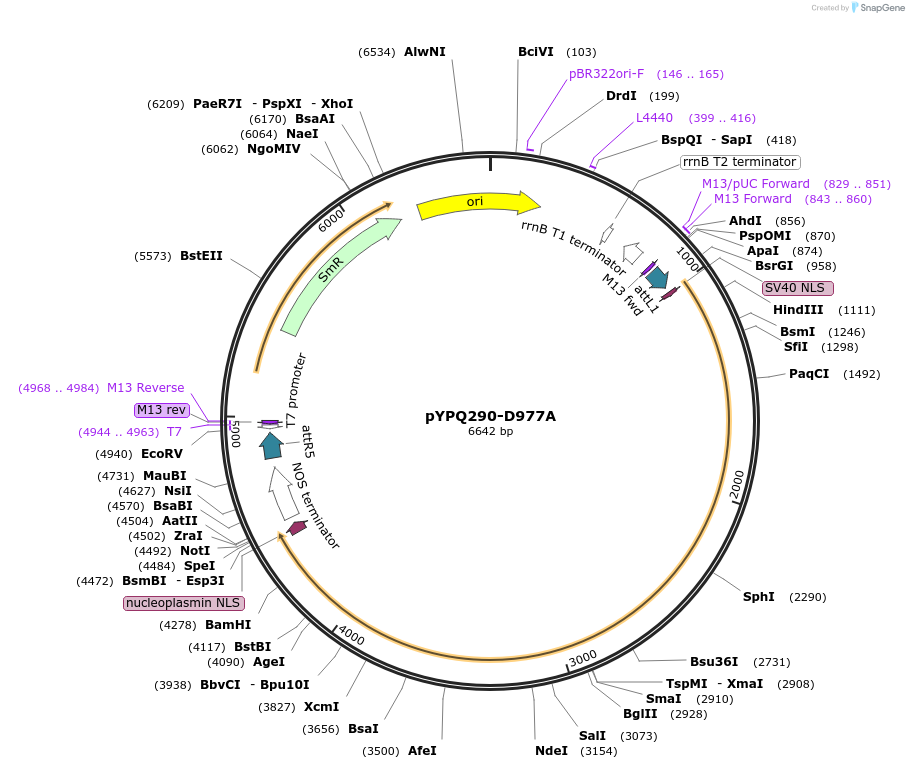
pYPQ290-D977A
Plasmid#129674PurposeGateway entry clone for CRISPR-Cas12b systemsDepositorInsertAacCas12b
UseCRISPR; Gateway compatible aaccas12b entry cloneExpressionPlantMutationD977A; AacCas12b is rice codon optimizedAvailable SinceMarch 18, 2020AvailabilityAcademic Institutions and Nonprofits only -
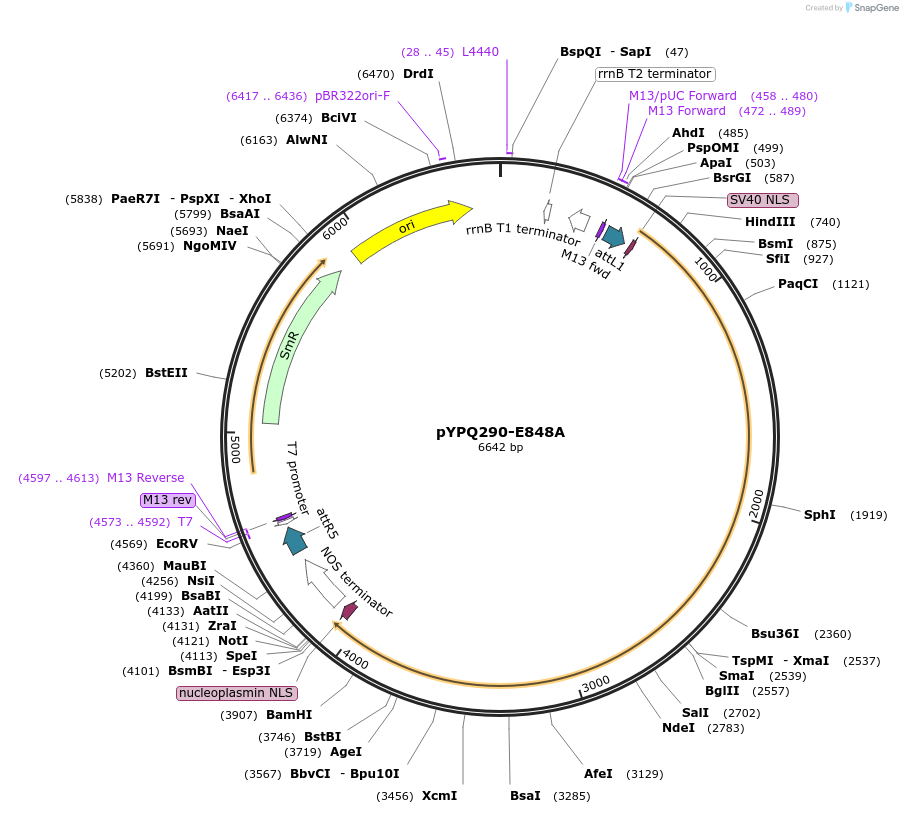
pYPQ290-E848A
Plasmid#129675PurposeGateway entry clone for CRISPR-Cas12b systemsDepositorInsertAacCas12b
UseCRISPR; Gateway compatible aaccas12b entry cloneTagsE848A; AacCas12b is rice codon optimizedExpressionPlantAvailable SinceMarch 18, 2020AvailabilityAcademic Institutions and Nonprofits only -
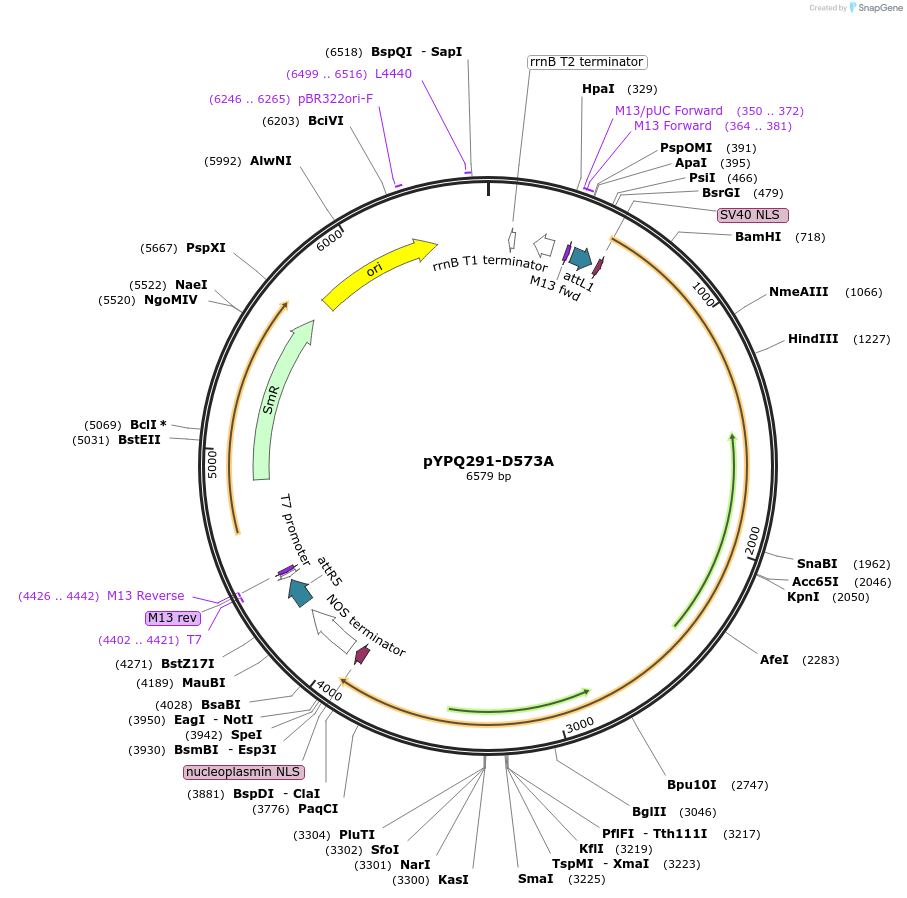
pYPQ291-D573A
Plasmid#129676PurposeGateway entry clone for CRISPR-Cas12b systemsDepositorInsertBthCas12b
UseCRISPR; Gateway compatible bthcas12b entry cloneExpressionPlantMutationD573A; BthCas12b is rice codon optimizedAvailable SinceMarch 18, 2020AvailabilityAcademic Institutions and Nonprofits only -
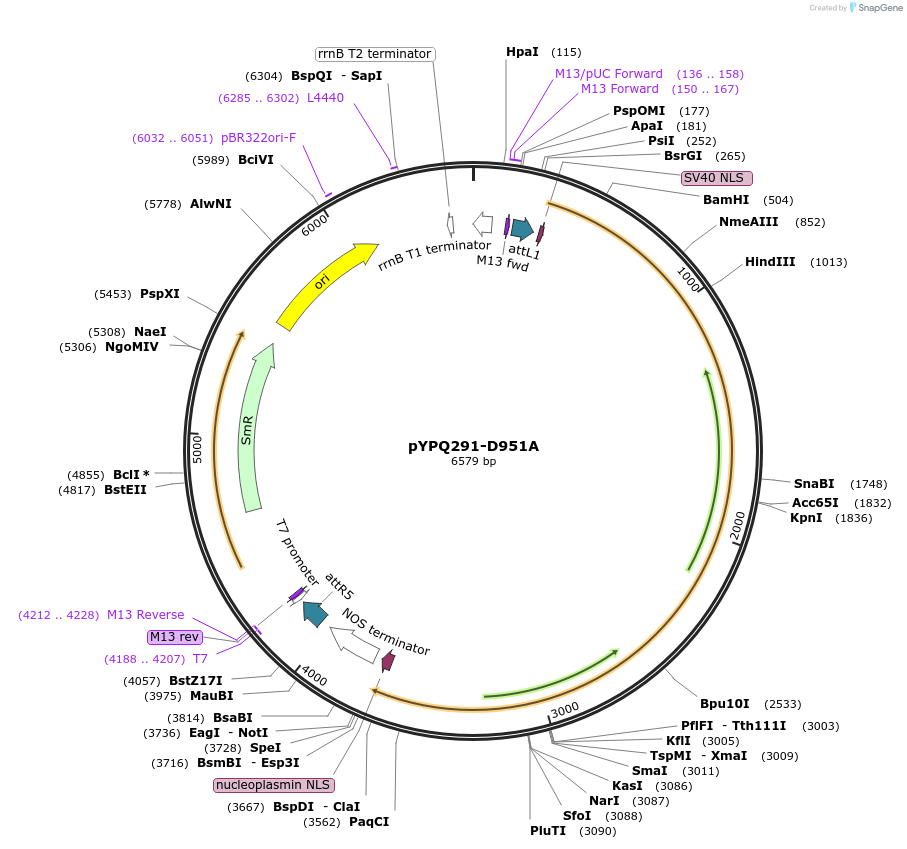
pYPQ291-D951A
Plasmid#129677PurposeGateway entry clone for CRISPR-Cas12b systemsDepositorInsertBthCas12b
UseCRISPR; Gateway compatible bthcas12b entry cloneExpressionPlantMutationD951A; BthCas12b is rice codon optimizedAvailable SinceMarch 18, 2020AvailabilityAcademic Institutions and Nonprofits only -
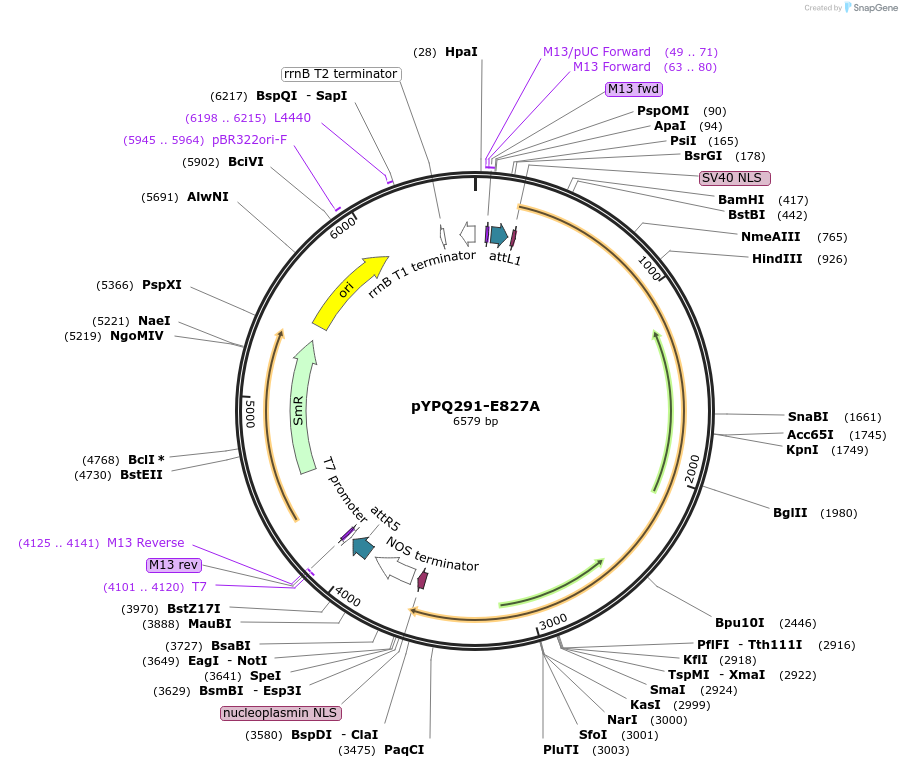
pYPQ291-E827A
Plasmid#129678PurposeGateway entry clone for CRISPR-Cas12b systemsDepositorInsertBthCas12b
UseCRISPR; Gateway compatible bthcas12b entry cloneExpressionPlantMutationE827A; BthCas12b is rice codon optimizedAvailable SinceMarch 18, 2020AvailabilityAcademic Institutions and Nonprofits only -
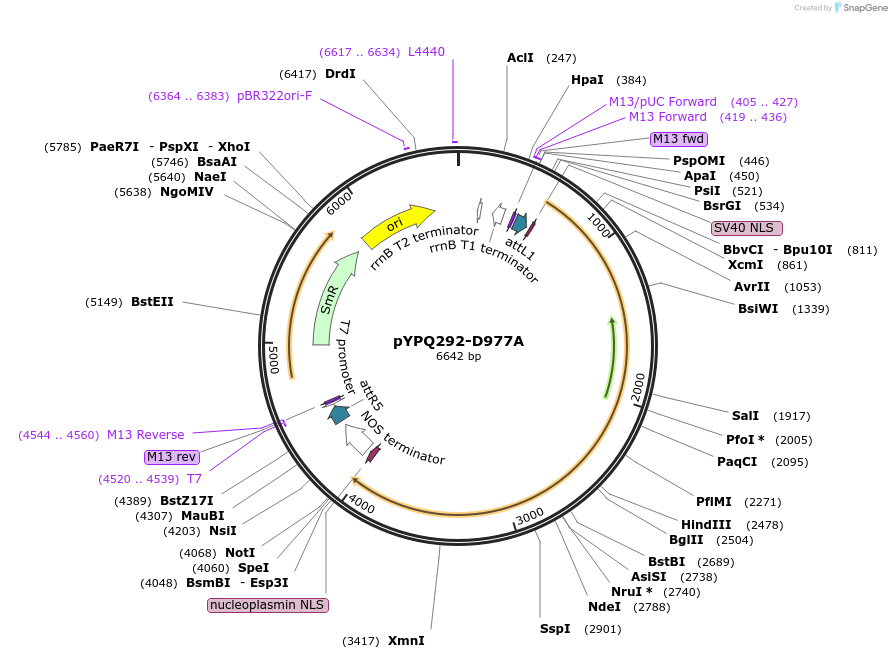
pYPQ292-D977A
Plasmid#129680PurposeGateway entry clone for CRISPR-Cas12b systemsDepositorInsertAaCas12b
UseCRISPR; Gateway compatible aacas12b entry cloneExpressionPlantMutationD977A; AaCas12b is rice codon optimizedAvailable SinceMarch 18, 2020AvailabilityAcademic Institutions and Nonprofits only -
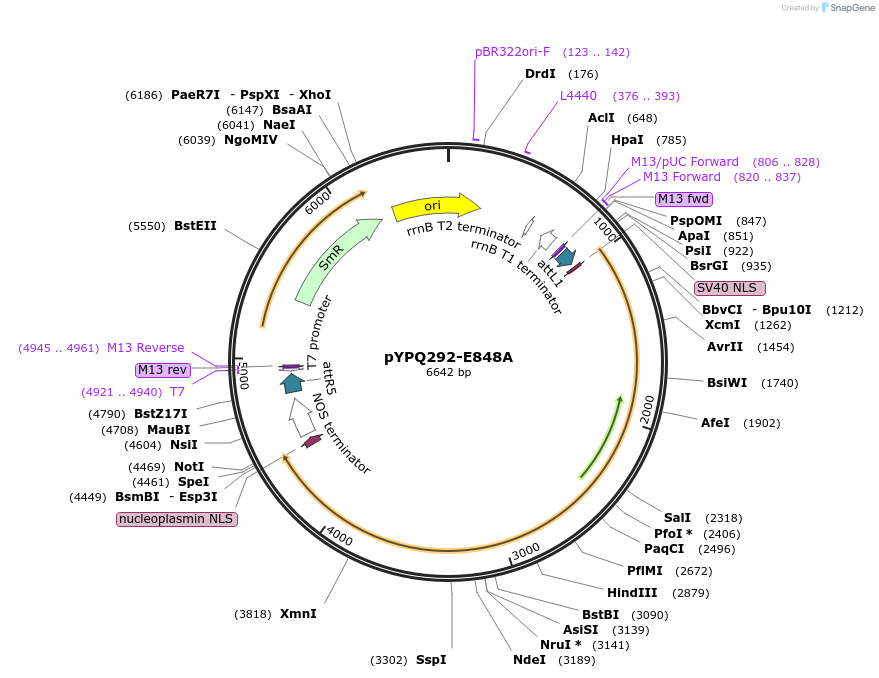
pYPQ292-E848A
Plasmid#129681PurposeGateway entry clone for CRISPR-Cas12b systemsDepositorInsertAaCas12b
UseCRISPR; Gateway compatible aacas12b entry cloneExpressionPlantMutationE848A; AaCas12b is rice codon optimizedAvailable SinceMarch 18, 2020AvailabilityAcademic Institutions and Nonprofits only -
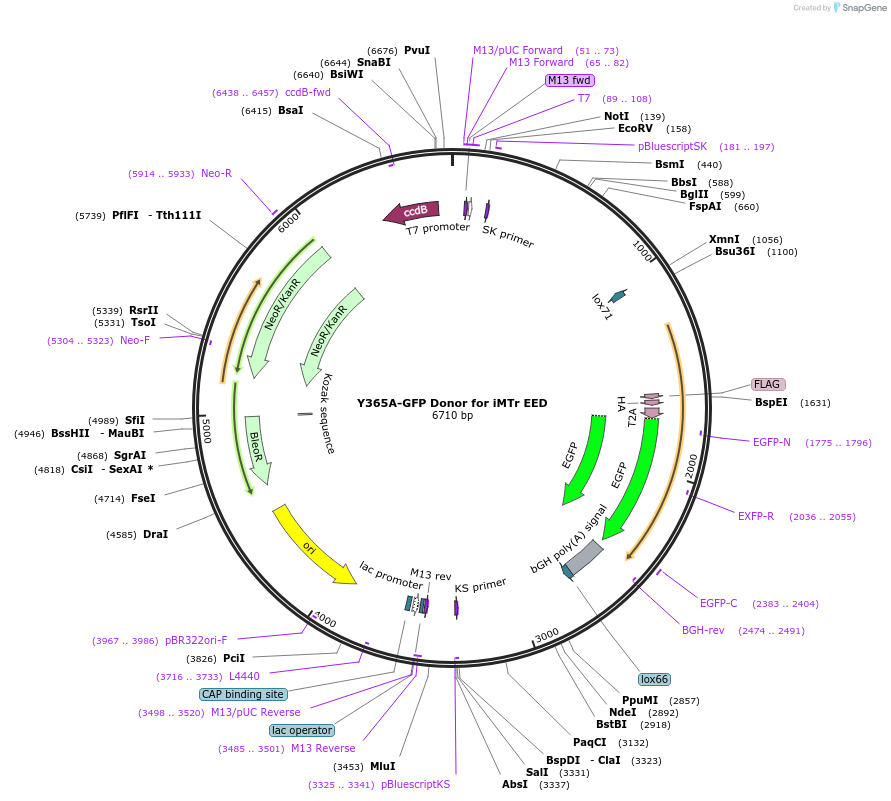
Y365A-GFP Donor for iMTr EED
Plasmid#131325PurposeDonor to generate i-MT-r systemDepositorInserti-MT-r.Donor
UseCRISPR and Mouse TargetingAvailable SinceMarch 6, 2020AvailabilityAcademic Institutions and Nonprofits only -
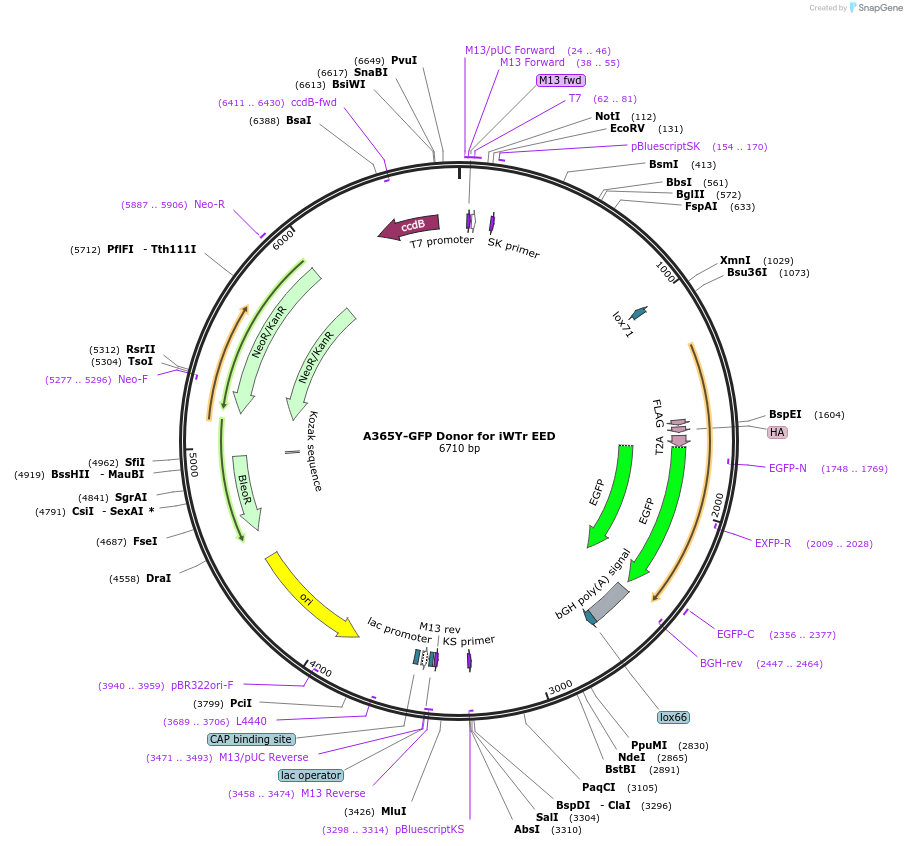
A365Y-GFP Donor for iWTr EED
Plasmid#131324PurposeDonor to generate i-WT-r systemDepositorInserti-WT-r.Donor
UseCRISPR and Mouse TargetingAvailable SinceMarch 6, 2020AvailabilityAcademic Institutions and Nonprofits only -
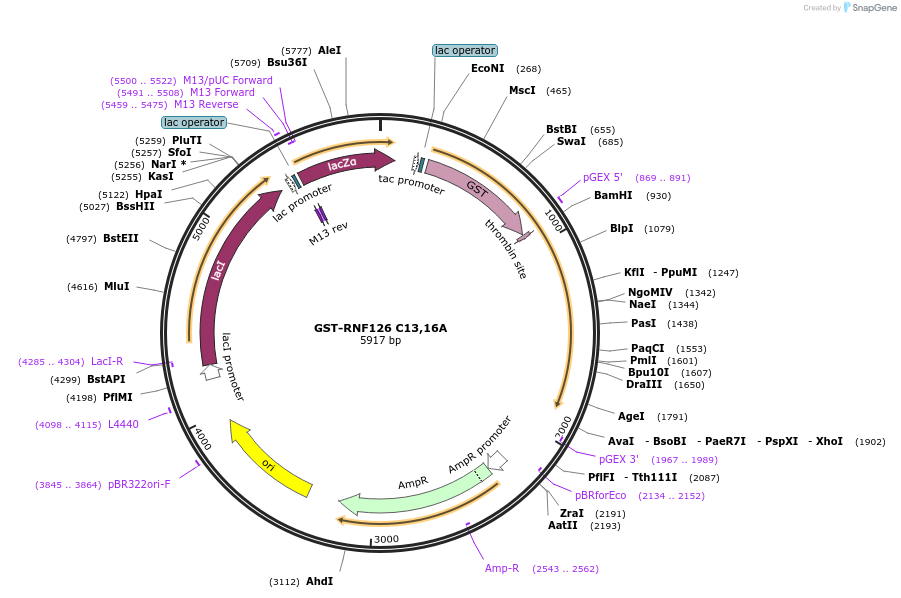
GST-RNF126 C13,16A
Plasmid#138645PurposeBacterial expression of GST-RNF126 ZnF mutant (C13,16A) fusion proteinDepositorAvailable SinceFeb. 27, 2020AvailabilityAcademic Institutions and Nonprofits only -
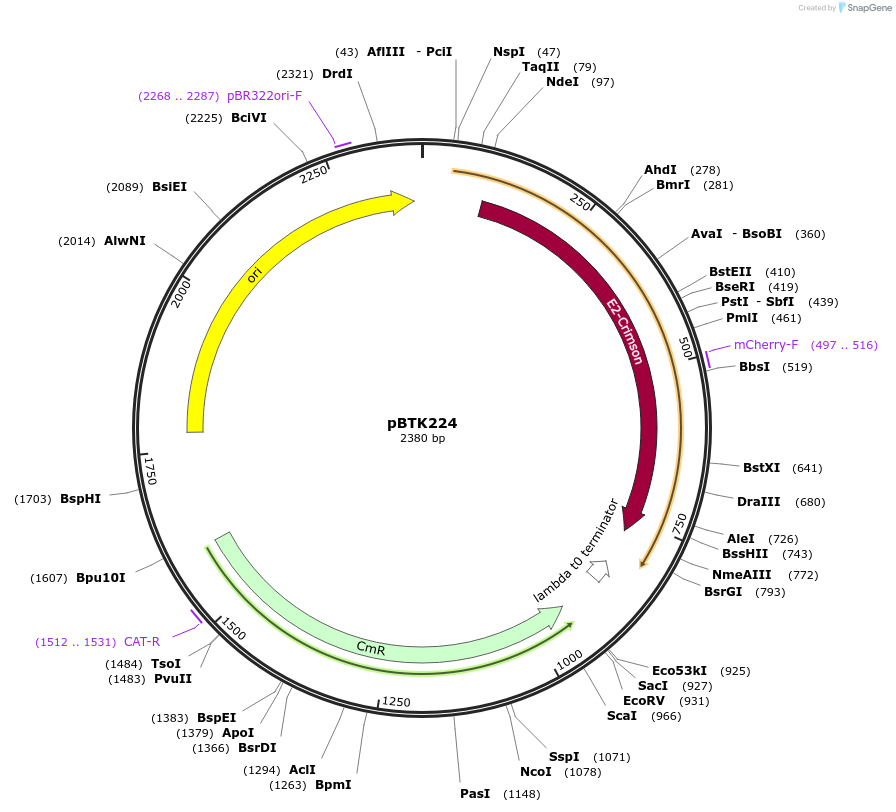
pBTK224
Plasmid#110590PurposeBTK Type 3 coding sequence for golden gate assemblyDepositorInsertE2-Crimson
UseSynthetic BiologyAvailable SinceFeb. 20, 2020AvailabilityIndustry, Academic Institutions, and Nonprofits -
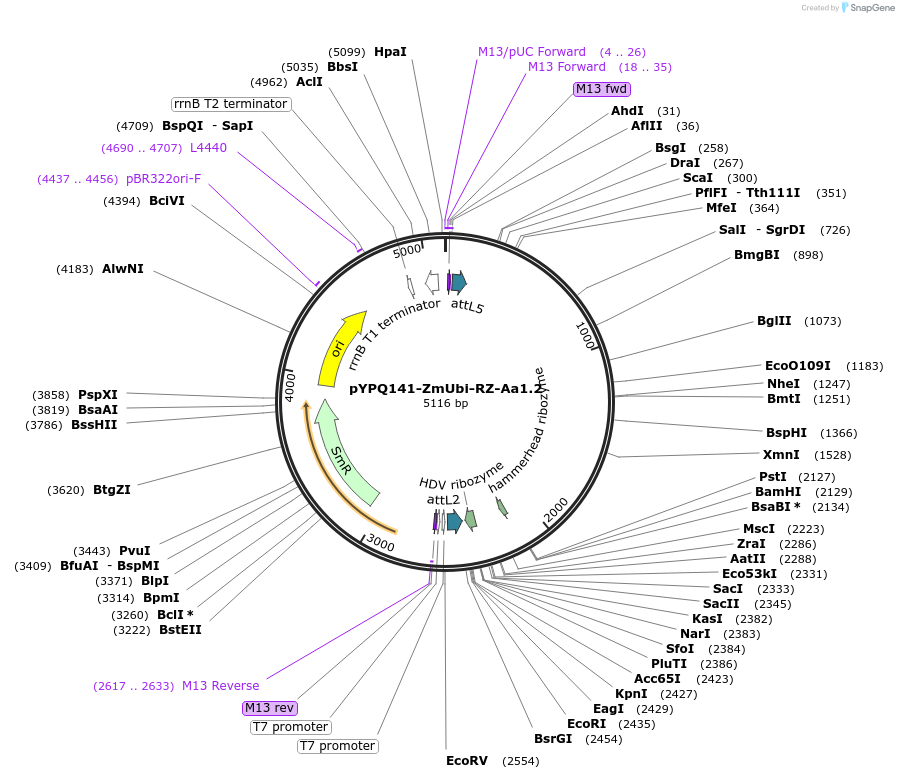
pYPQ141-ZmUbi-RZ-Aa1.2
Plasmid#136373PurposeGateway entry clone for AaCas12b sgRNA expression under ZmUbi promoter with ribozyme processing; sgRNA scaffold 1.2 is usedDepositorTypeEmpty backboneUseCRISPRExpressionPlantAvailable SinceFeb. 18, 2020AvailabilityAcademic Institutions and Nonprofits only -
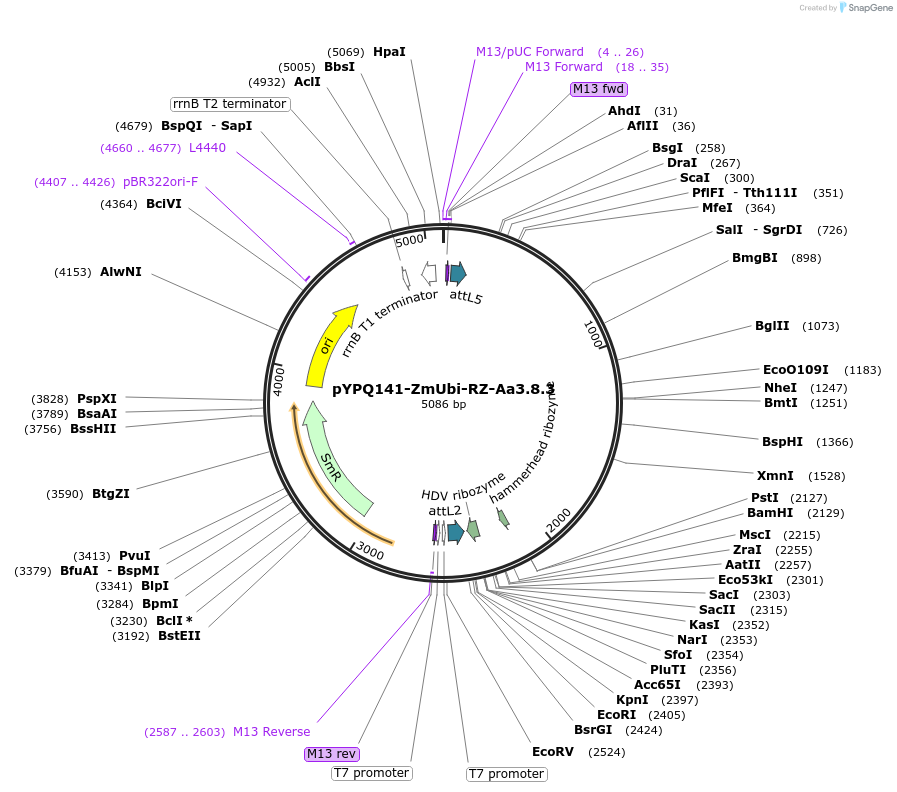
pYPQ141-ZmUbi-RZ-Aa3.8.3
Plasmid#136374PurposeGateway entry clone for AaCas12b sgRNA expression under ZmUbi promoter with ribozyme processing; sgRNA scaffold 3.8 with MS2 at the third stem loop is usedDepositorTypeEmpty backboneUseCRISPRExpressionPlantAvailable SinceFeb. 18, 2020AvailabilityAcademic Institutions and Nonprofits only -
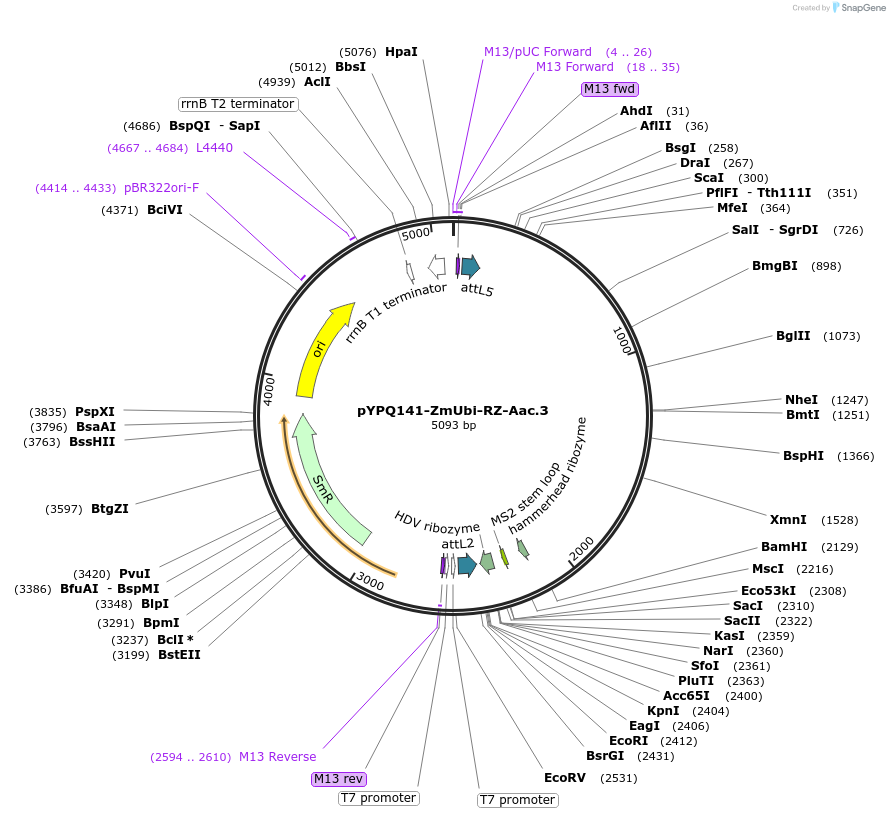
pYPQ141-ZmUbi-RZ-Aac.3
Plasmid#136377PurposeGateway entry clone for AacCas12b sgRNA expression under ZmUbi promoter with ribozyme processing; sgRNA scaffold with MS2 at the third stem loop is usedDepositorTypeEmpty backboneUseCRISPRExpressionPlantAvailable SinceFeb. 18, 2020AvailabilityAcademic Institutions and Nonprofits only -
pYPQ141-ZmUbi-RZ-Aa1.2.3
Plasmid#136372PurposeGateway entry clone for AaCas12b sgRNA expression under ZmUbi promoter with ribozyme processing; sgRNA scaffold 1.2 with MS2 at the third stem loop is usedDepositorTypeEmpty backboneUseCRISPRExpressionPlantAvailable SinceFeb. 18, 2020AvailabilityAcademic Institutions and Nonprofits only -
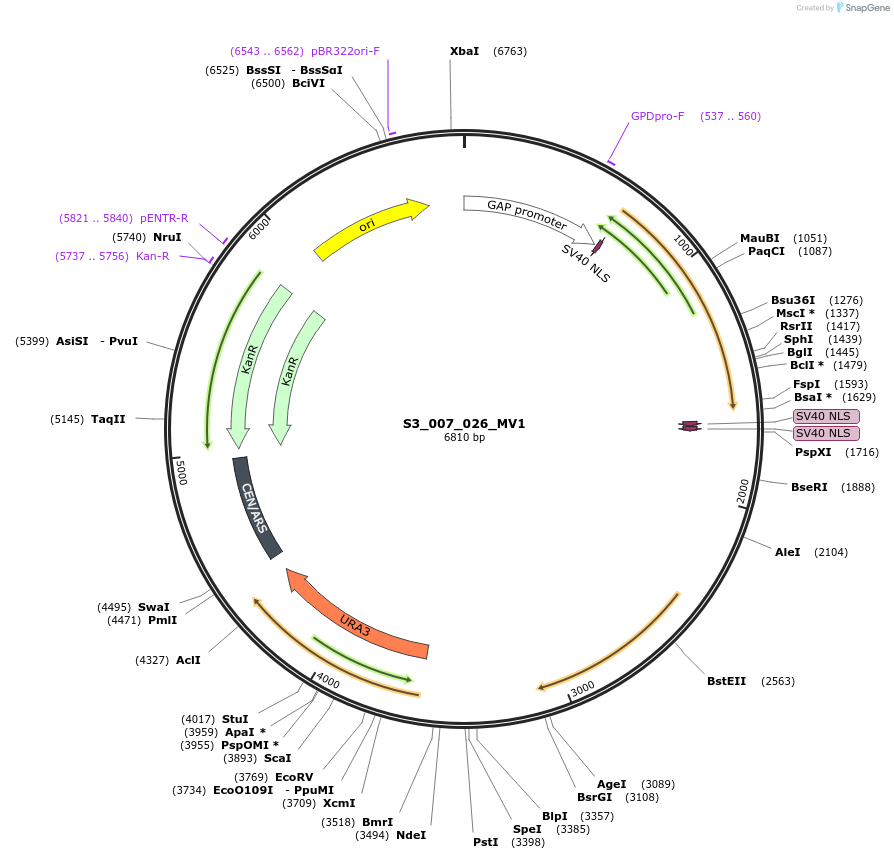
S3_007_026_MV1
Plasmid#134419PurposepYTKMV1-pTDH3_ NLS_FdeR-N_2NLS _tPGK1-pGPM1-fdeO_mcherry_tTDH1DepositorInsertsmCherry
NLS_FdeR-N_2xNLS
TagsnoneExpressionYeastMutationnonePromoterpGMP1 and pTDH3Available SinceFeb. 10, 2020AvailabilityAcademic Institutions and Nonprofits only -
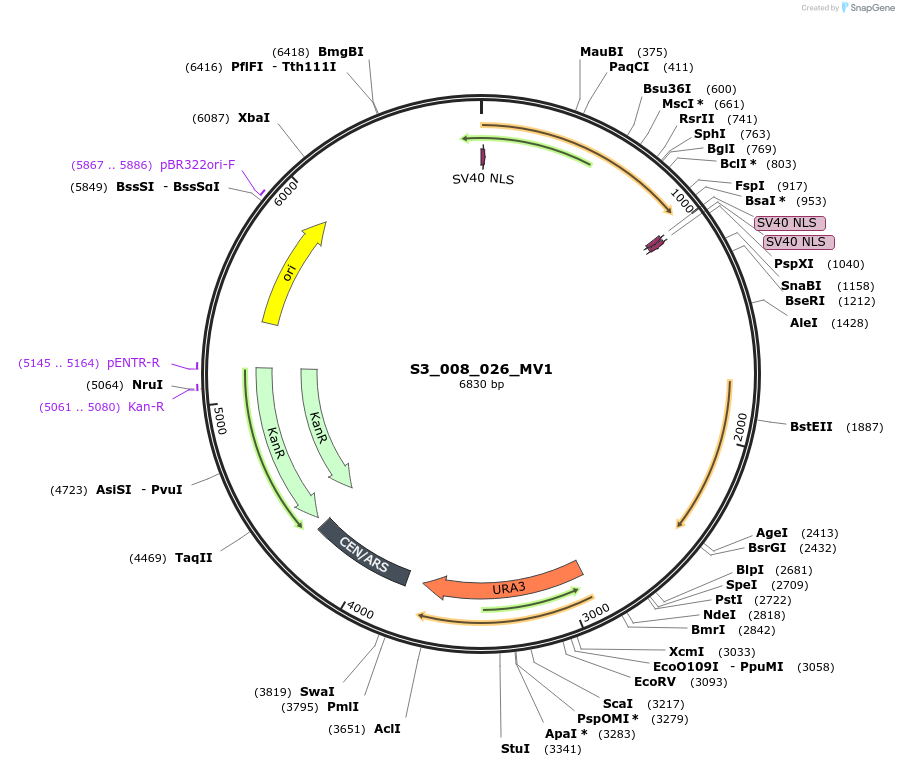
S3_008_026_MV1
Plasmid#134420PurposepYTKMV1-pRPL18B_ NLS_FdeR_N_2NLS _tPGK1-pGPM1-fdeO_mcherry_tTDH1DepositorInsertsmCherry
NLS_FdeR-N_2xNLS
TagsnoneExpressionYeastMutationnonePromoterpGMP1 and pRPL18BAvailable SinceFeb. 10, 2020AvailabilityAcademic Institutions and Nonprofits only -
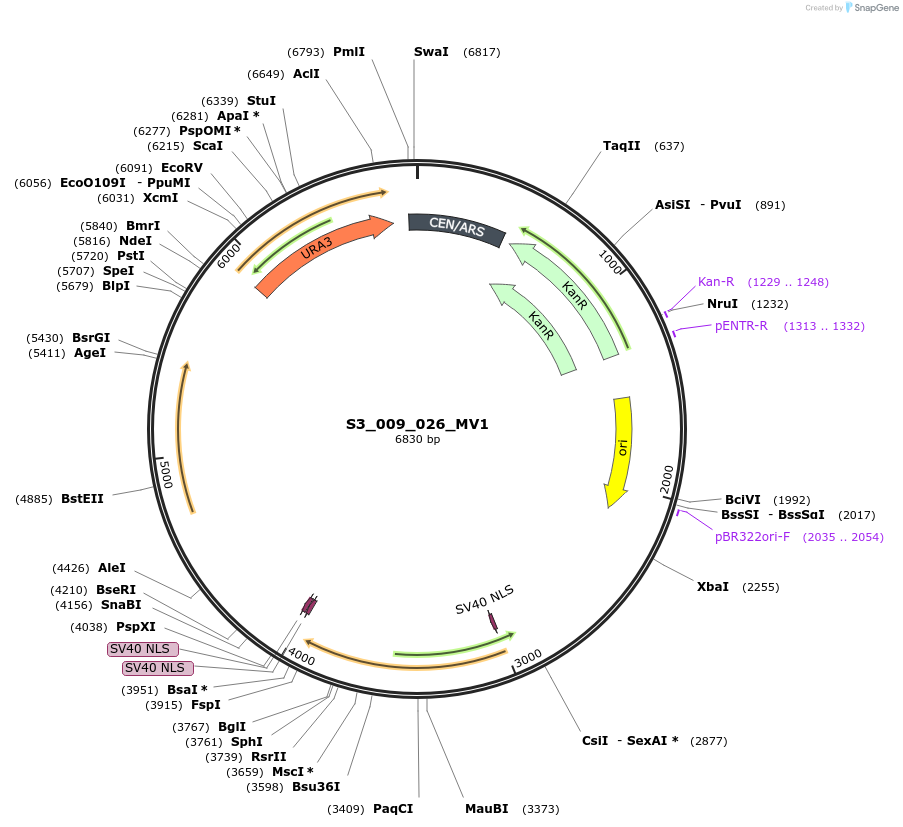
S3_009_026_MV1
Plasmid#134421PurposepYTKMV1-pREV1_ NLS_FdeR-N_2NLS-pGPM1-fdeO_mcherry_tTDH1DepositorInsertsmCherry
NLS_FdeR-N_2xNLS
TagsnoneExpressionYeastMutationnonePromoterpGMP1 and pREV1Available SinceFeb. 10, 2020AvailabilityAcademic Institutions and Nonprofits only -
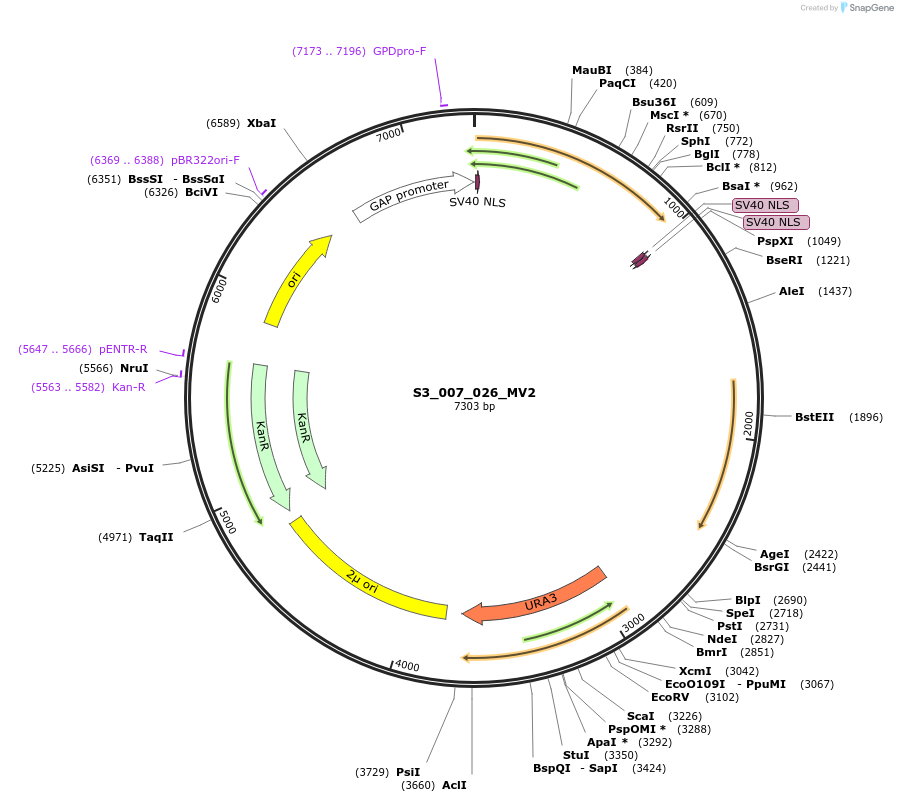
S3_007_026_MV2
Plasmid#134424PurposepYTKMV2-pTDH3_ NLS_FdeR-N_2NLS _tPGK1-pGPM1-fdeO_mcherry_tTDH1DepositorInsertsmCherry
NLS_FdeR-N_2xNLS
TagsnoneExpressionYeastMutationnonePromoterpGMP1 and pTDH3Available SinceFeb. 10, 2020AvailabilityAcademic Institutions and Nonprofits only -
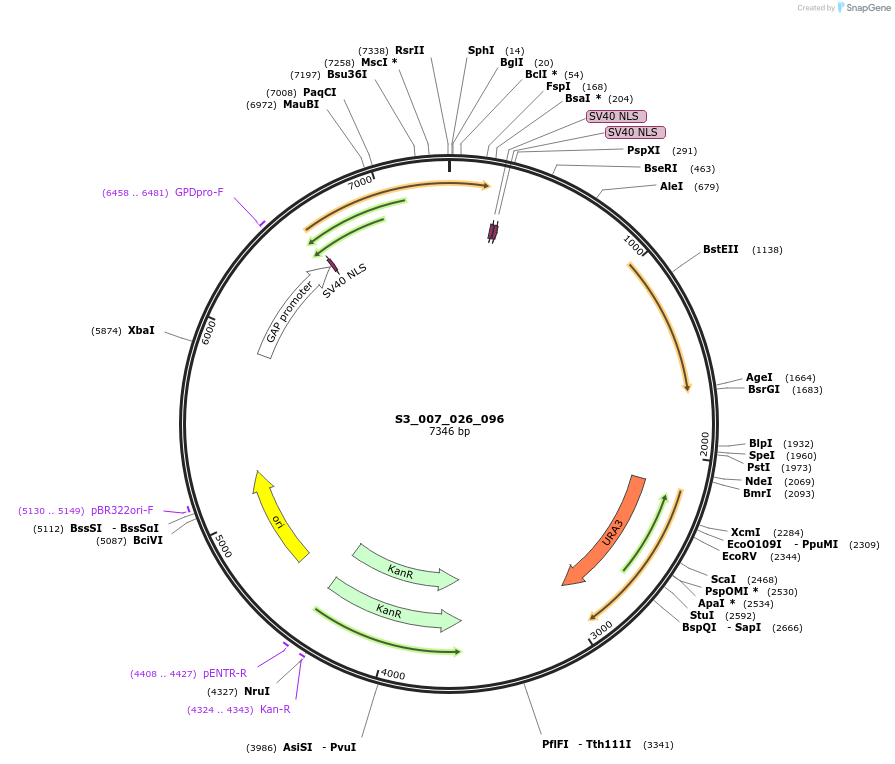
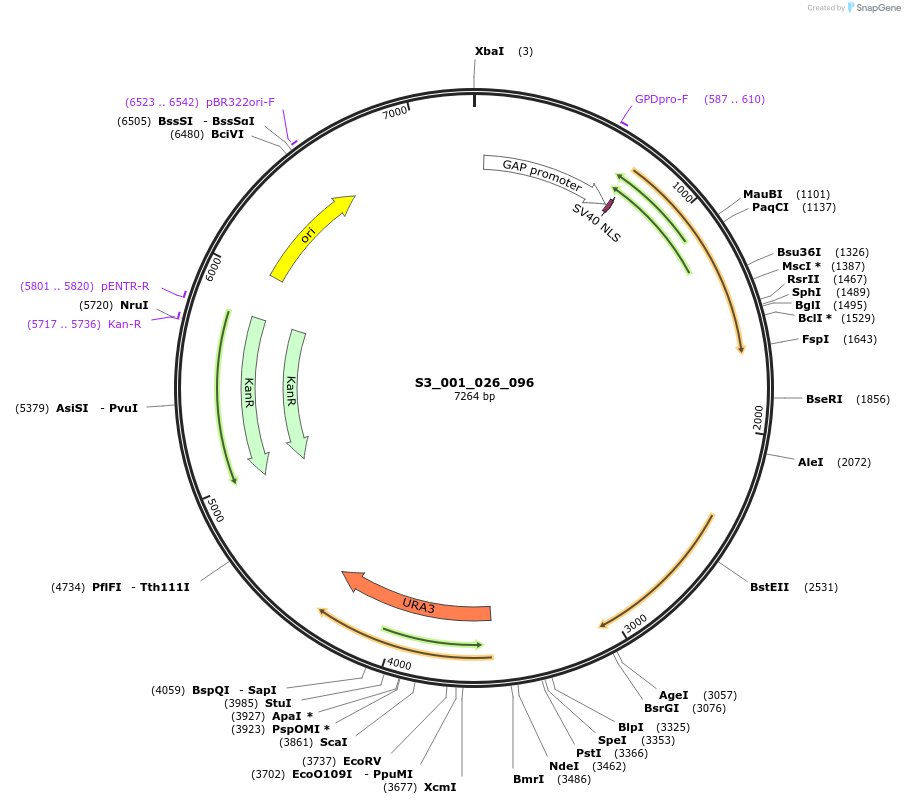
S3_007_026_096
Plasmid#134410PurposepYTK096-pTDH3_ NLS_FdeR-N_2NLS _tPGK1-pGPM1-fdeO_mcherry_tTDH1DepositorInsertsmCherry
NLS_FdeR-N_2xNLS
TagsnoneExpressionYeastMutationnonePromoterpGPM1 and pTDH3Available SinceFeb. 10, 2020AvailabilityAcademic Institutions and Nonprofits only -
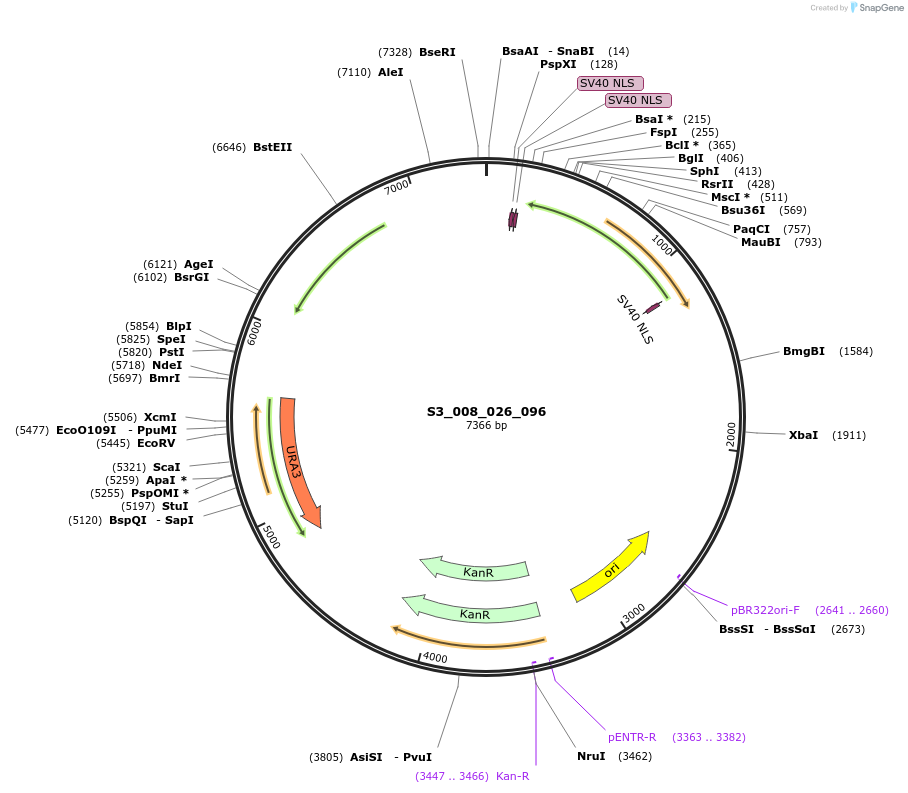
S3_008_026_096
Plasmid#134411PurposepYTK096-pRPL18B_ NLS_FdeR_N_2NLS _tPGK1-pGPM1-fdeO_mcherry_tTDH1DepositorInsertsmCherry
NLS_FdeR-N_2xNLS
TagsnoneExpressionYeastMutationnonePromoterpGPM1 and pRPL18BAvailable SinceFeb. 10, 2020AvailabilityAcademic Institutions and Nonprofits only -
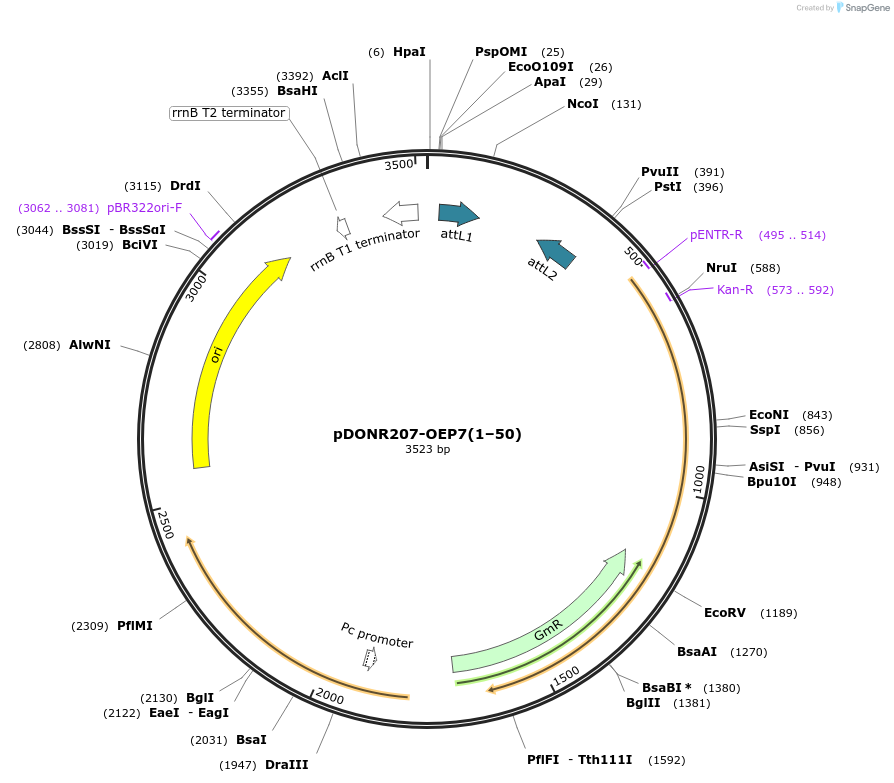
pDONR207-OEP7(1–50)
Plasmid#100601PurposeGateway entry vector for AtOEP7(1–50)DepositorInsertOEP7(1–50) (OEP7 Mustard Weed)
UseGateway entry vectorAvailable SinceFeb. 5, 2020AvailabilityAcademic Institutions and Nonprofits only -
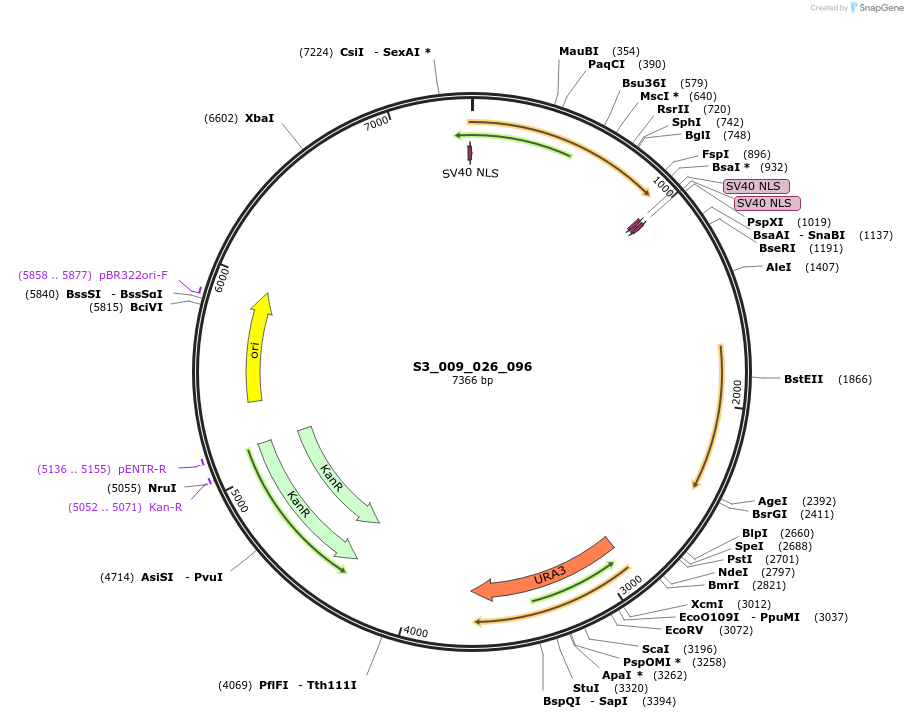
S3_009_026_096
Plasmid#134412PurposepYTK096-pREV1_ NLS_FdeR-N_2NLS-pGPM1-fdeO_mcherry_tTDH1DepositorInsertsmCherry
NLS_FdeR-N_2xNLS
TagsnoneExpressionYeastMutationnonePromoterpGMP1 and pREV1Available SinceDec. 19, 2019AvailabilityAcademic Institutions and Nonprofits only -
S3_001_026_096
Plasmid#134402PurposepYTK096-pTDH3_NLS_FdeR-N_tPGK1-pGPM1-fdeO_mcherry_tTDH1DepositorInsertsmCherry
NLS_FdeR-N
TagsnoneExpressionYeastMutationnonePromoterpGPM1 and pTDH3Available SinceDec. 19, 2019AvailabilityAcademic Institutions and Nonprofits only -
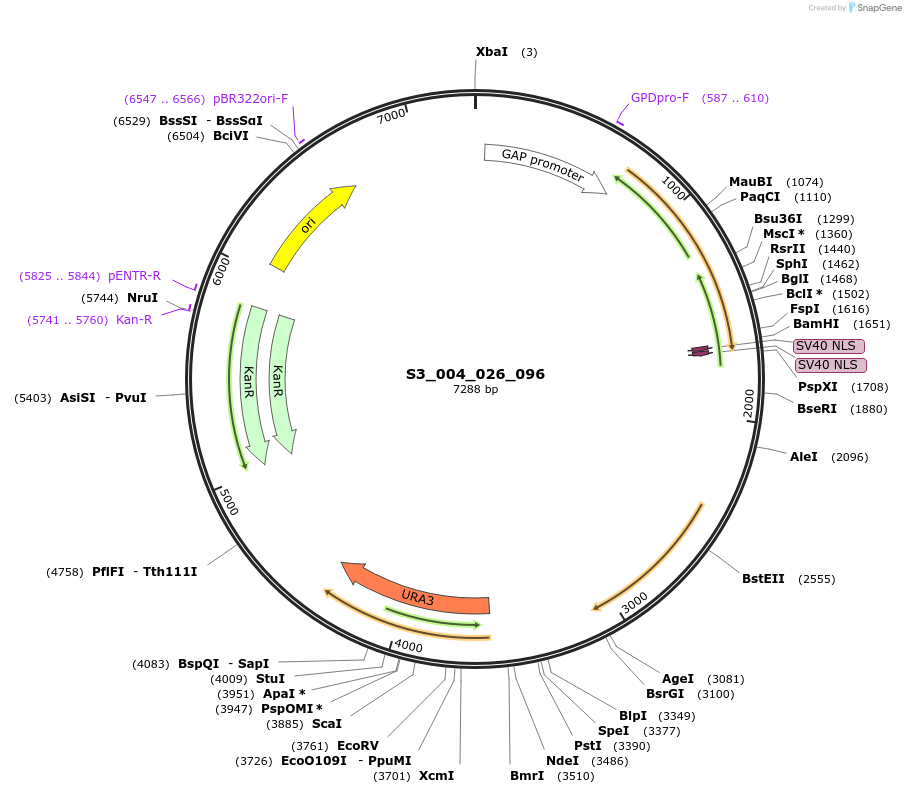
S3_004_026_096
Plasmid#134407PurposepYTK096-pTDH3_FdeR-N_2NLS_tPGK1-pGPM1-fdeO_mcherry_tTDH1DepositorInsertsmCherry
FdeR-N_2xNLS
TagsnoneExpressionYeastMutationnonePromoterpGPM1 and pTDH3Available SinceDec. 19, 2019AvailabilityAcademic Institutions and Nonprofits only -
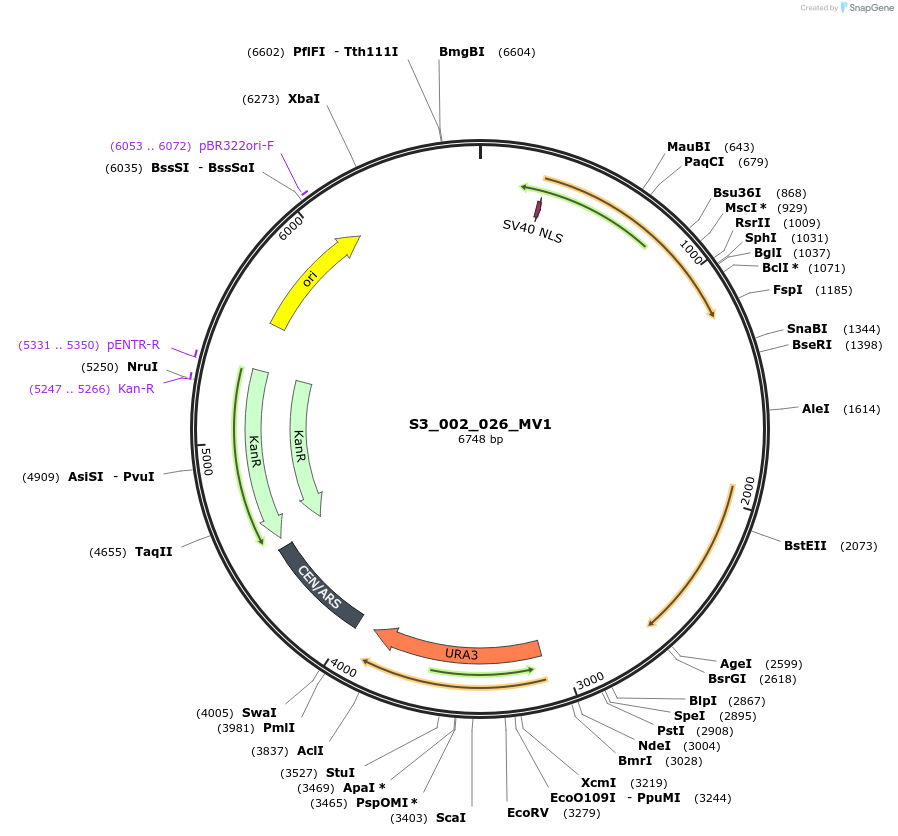
S3_002_026_MV1
Plasmid#134414PurposepYTKMV1-pRPL18B_NLS_FdeR-N_tPGK1-pGPM1-fdeO_mcherry_tTDH1DepositorInsertsmCherry
NLS_FdeR-N
TagsnoneExpressionYeastMutationnonePromoterpGMP1 and pRPL18BAvailable SinceDec. 19, 2019AvailabilityAcademic Institutions and Nonprofits only -
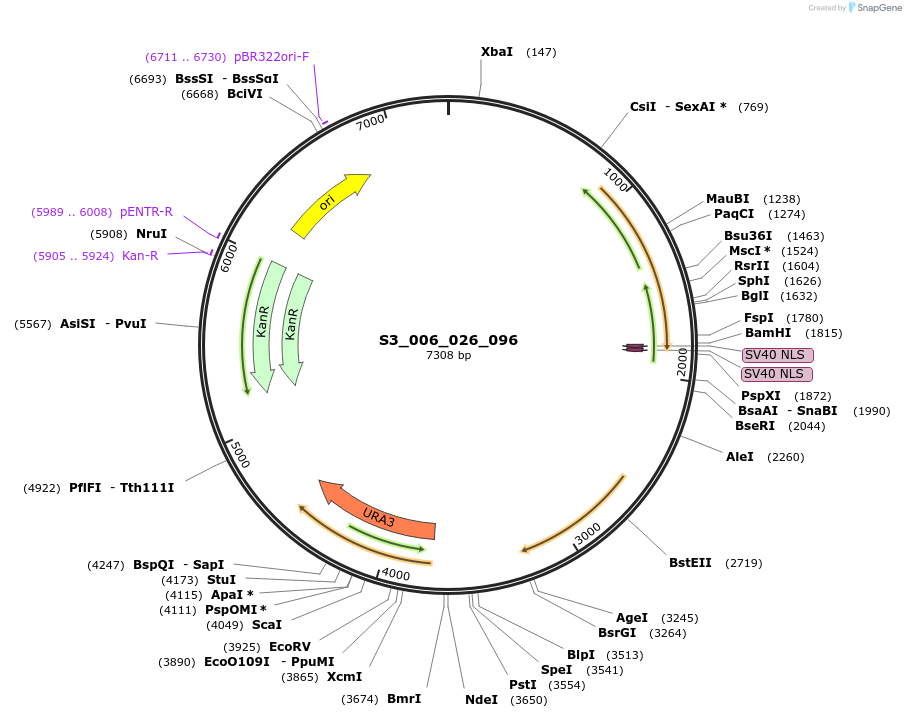
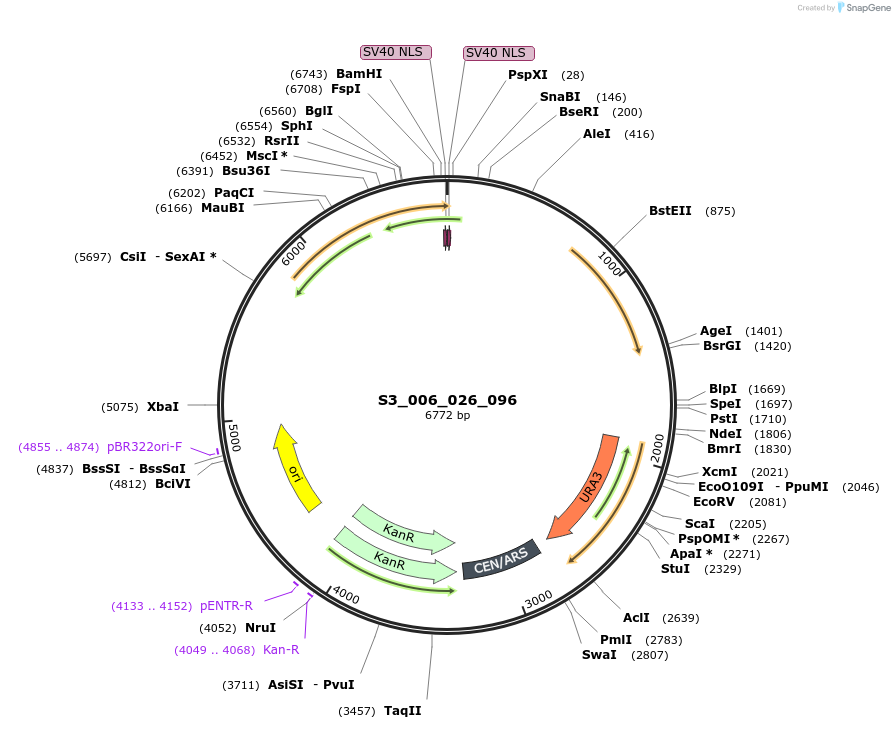
S3_006_026_096
Plasmid#134409PurposepYTK096-pREV1_FdeR-N_2NLS_tPGK1- pGPM1-fdeO_mcherry_tTDH1DepositorInsertsmCherry
FdeR-N_2xNLS
TagsnoneExpressionYeastMutationnonePromoterpGPM1 and pREV1Available SinceDec. 19, 2019AvailabilityAcademic Institutions and Nonprofits only -
S3_006_026_096
Plasmid#134418PurposepYTK096-pREV1_FdeR-N_2NLS_tPGK1- pGPM1-fdeO_mcherry_tTDH1DepositorInsertsmCherry
FdeR-N_2xNLS
TagsnoneExpressionYeastMutationnonePromoterpGMP1 and pREV1Available SinceDec. 18, 2019AvailabilityAcademic Institutions and Nonprofits only -
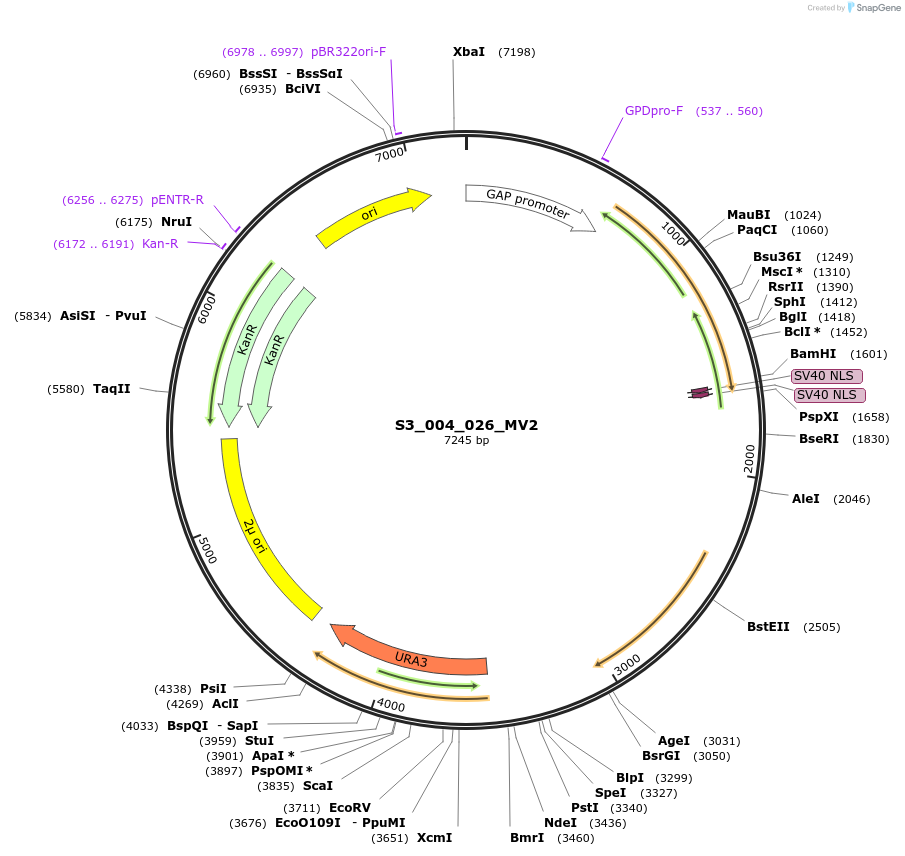
S3_004_026_MV2
Plasmid#134423PurposepYTKMV2-pTDH3_FdeR-N_2NLS_tPGK1-pGPM1-fdeO_mcherry_tTDH1DepositorInsertsmCherry
FdeR-N_2xNLS
TagsnoneExpressionYeastMutationnonePromoterpGMP1 and pTDH3Available SinceDec. 18, 2019AvailabilityAcademic Institutions and Nonprofits only -
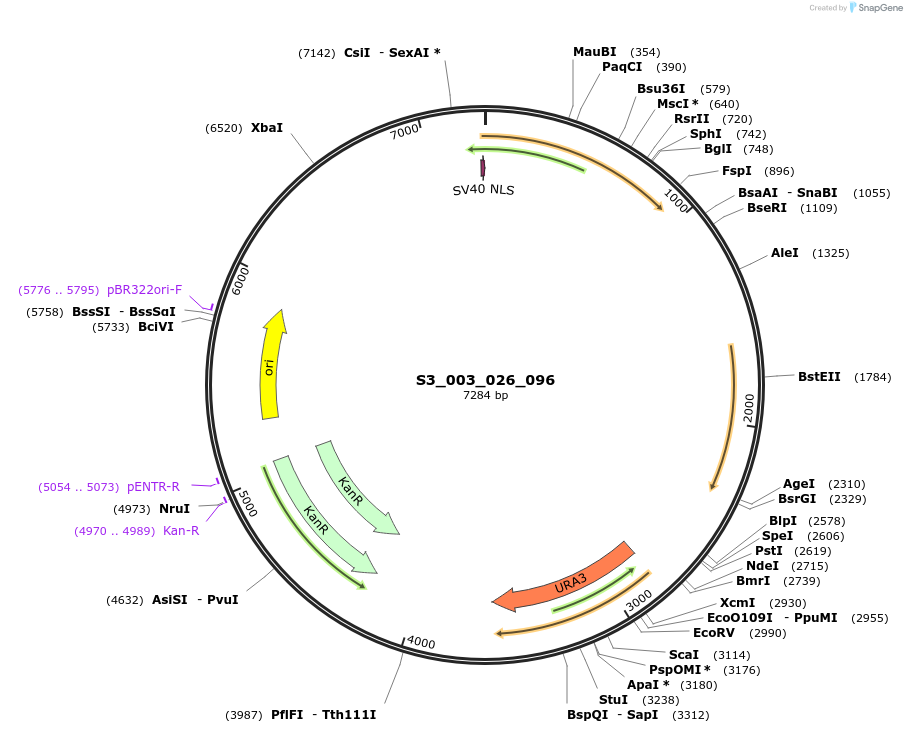
S3_003_026_096
Plasmid#134406PurposepYTK096-pREV1_NLS_FdeR-N_tPGK1-pGPM1-fdeO_mcherry_tTDH1DepositorInsertsmCherry
NLS_FdeR-N
TagsnoneExpressionYeastMutationnonePromoterpGPM1 and pREV1Available SinceDec. 18, 2019AvailabilityAcademic Institutions and Nonprofits only -
S3_005_026_MV1
Plasmid#134417PurposepYTKMV1-pRPL18B_FdeR-N_2NLS_tPGK1-pGPM1-fdeO_mcherry_tTDH1DepositorInsertsmCherry
FdeR-N_2xNLS
TagsnoneExpressionYeastMutationnonePromoterpGMP1 and pRPL18BAvailable SinceDec. 18, 2019AvailabilityAcademic Institutions and Nonprofits only -
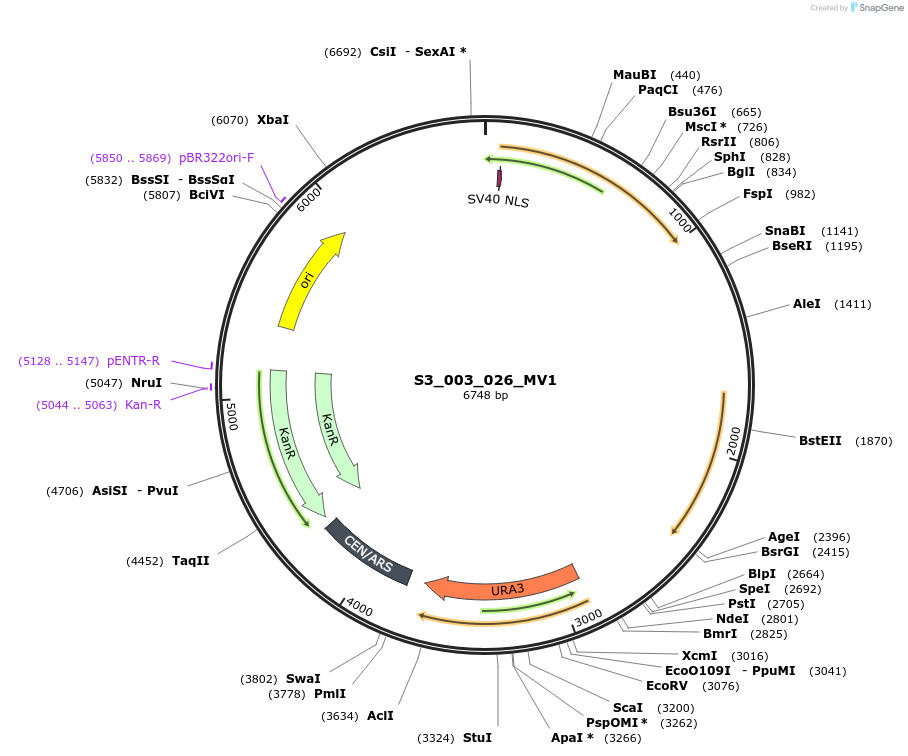
S3_003_026_MV1
Plasmid#134415PurposepYTKMV1-pREV1_NLS_FdeR-N_tPGK1-pGPM1-fdeO_mcherry_tTDH1DepositorInsertsmCherry
NLS_FdeR-N
TagsnoneExpressionYeastMutationnonePromoterpGMP1 and pREV1Available SinceDec. 18, 2019AvailabilityAcademic Institutions and Nonprofits only -
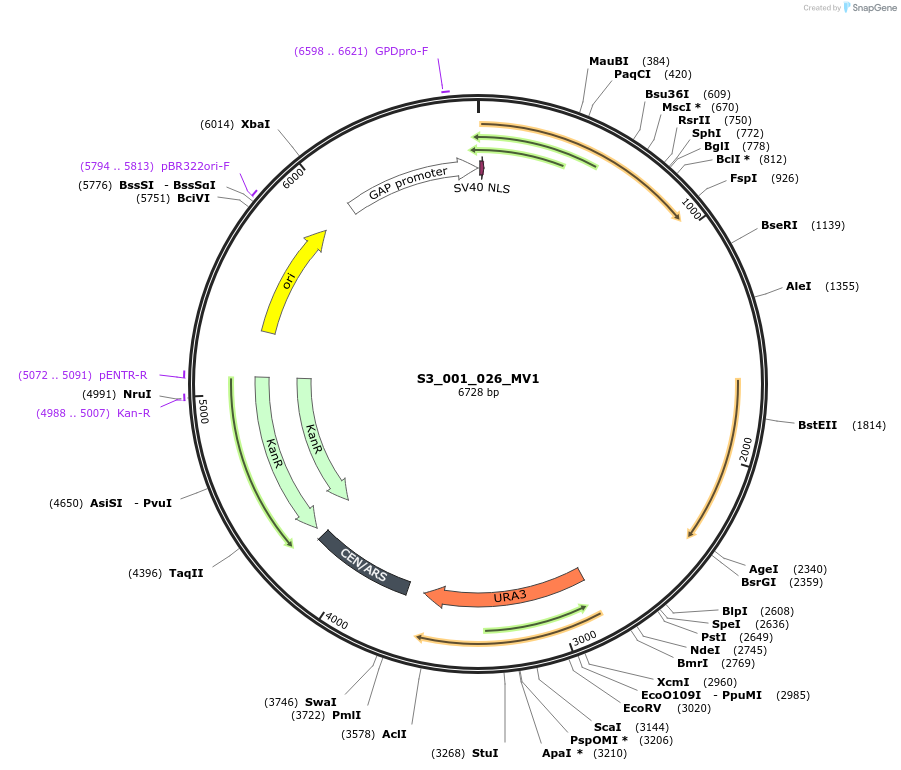
S3_001_026_MV1
Plasmid#134413PurposepYTKMV1-pTDH3_NLS_FdeR-N_tPGK1-pGPM1-fdeO_mcherry_tTDH1DepositorInsertsmCherry
NLS_FdeR-N
TagsnoneExpressionYeastMutationnonePromoterpGPM1 and pTDH3Available SinceDec. 18, 2019AvailabilityAcademic Institutions and Nonprofits only -
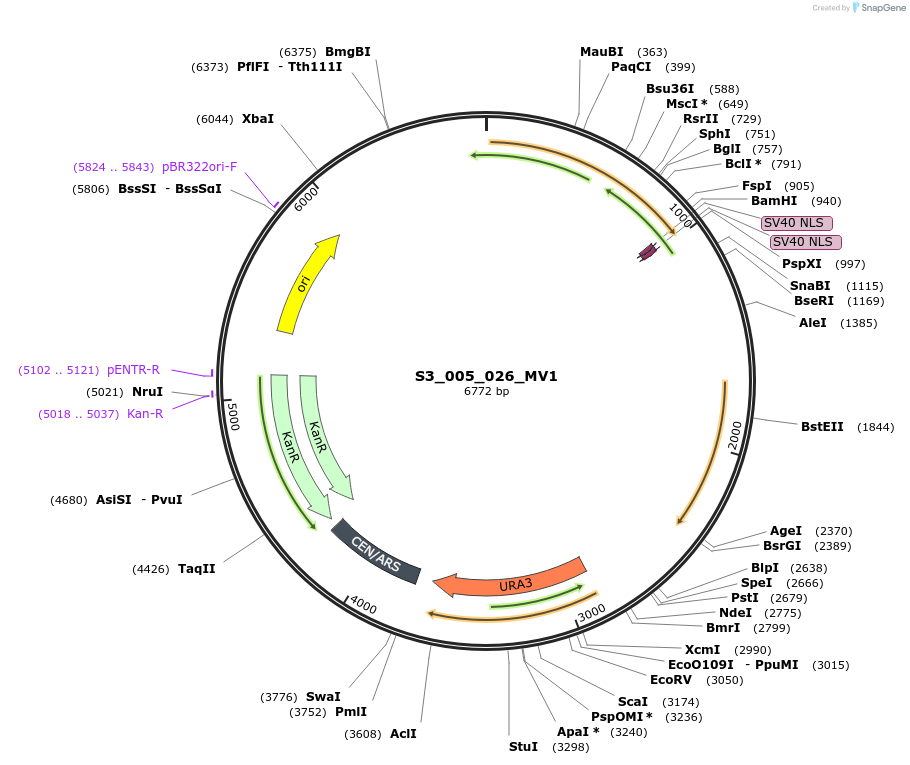
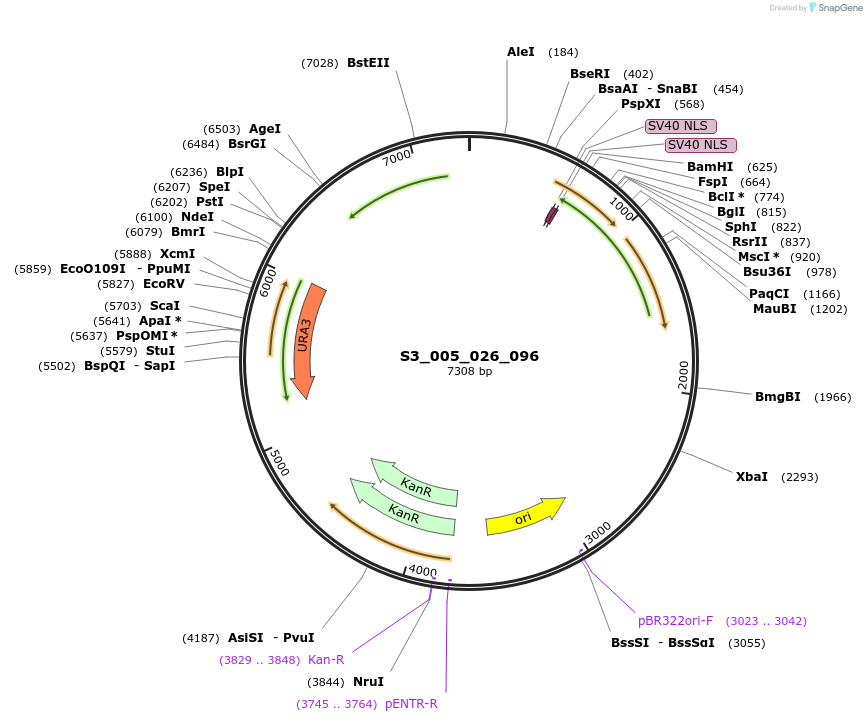
S3_005_026_096
Plasmid#134408PurposepYTK096-pRPL18B_FdeR-N_2NLS_tPGK1-pGPM1-fdeO_mcherry_tTDH1DepositorInsertsmCherry
FdeR-N_2xNLS
TagsnoneExpressionYeastMutationnonePromoterpGPM1 and pRPL18BAvailable SinceDec. 18, 2019AvailabilityAcademic Institutions and Nonprofits only -
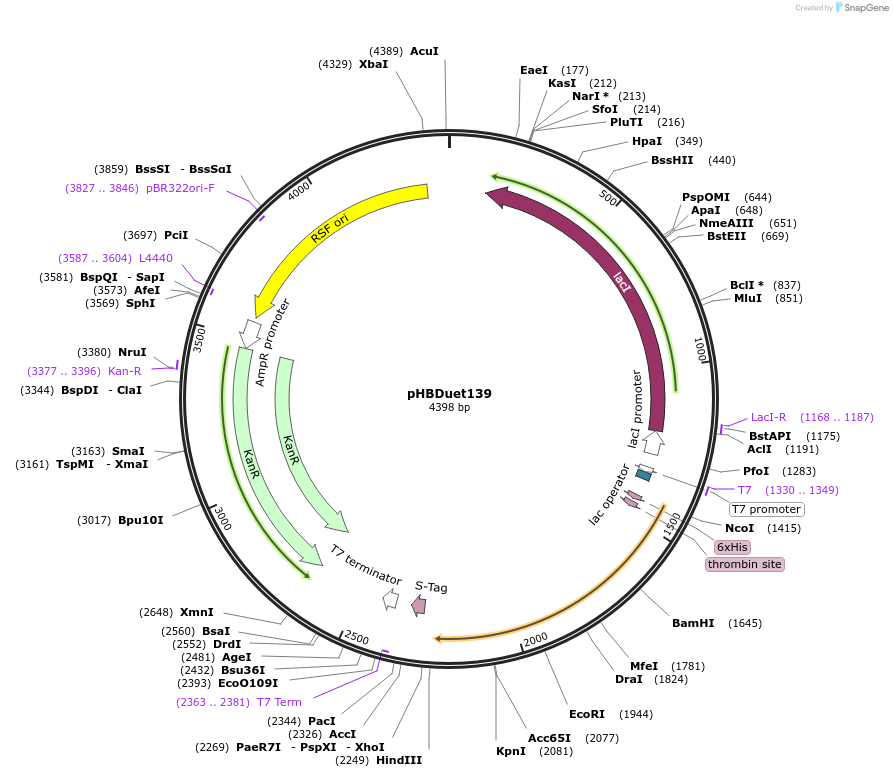
pHBDuet139
Plasmid#121913PurposeCharge-Introduced NpuDnaB mini-inteinDepositorInsertCI-NpuDnaBΔ290 intein
TagsGB1 and His6-GB1ExpressionBacterialMutationDeletion of 290-residue endonuclease domain, I53K…PromoterT7Available SinceDec. 2, 2019AvailabilityAcademic Institutions and Nonprofits only -
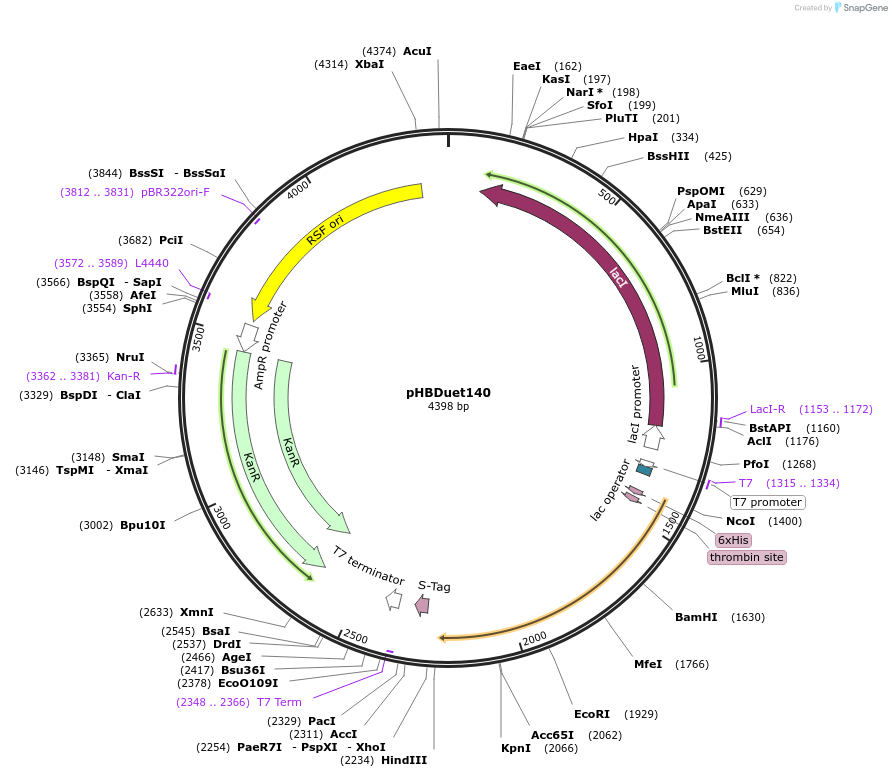
pHBDuet140
Plasmid#121914PurposeOrthogonal NpuDnaB mini-intein inteinDepositorInsertOth-NpuDnaBΔ290 intein
TagsGB1 and His6-GB1ExpressionBacterialMutationDeletion of 290-residue endonuclease domain, I53K…PromoterT7Available SinceDec. 2, 2019AvailabilityAcademic Institutions and Nonprofits only -
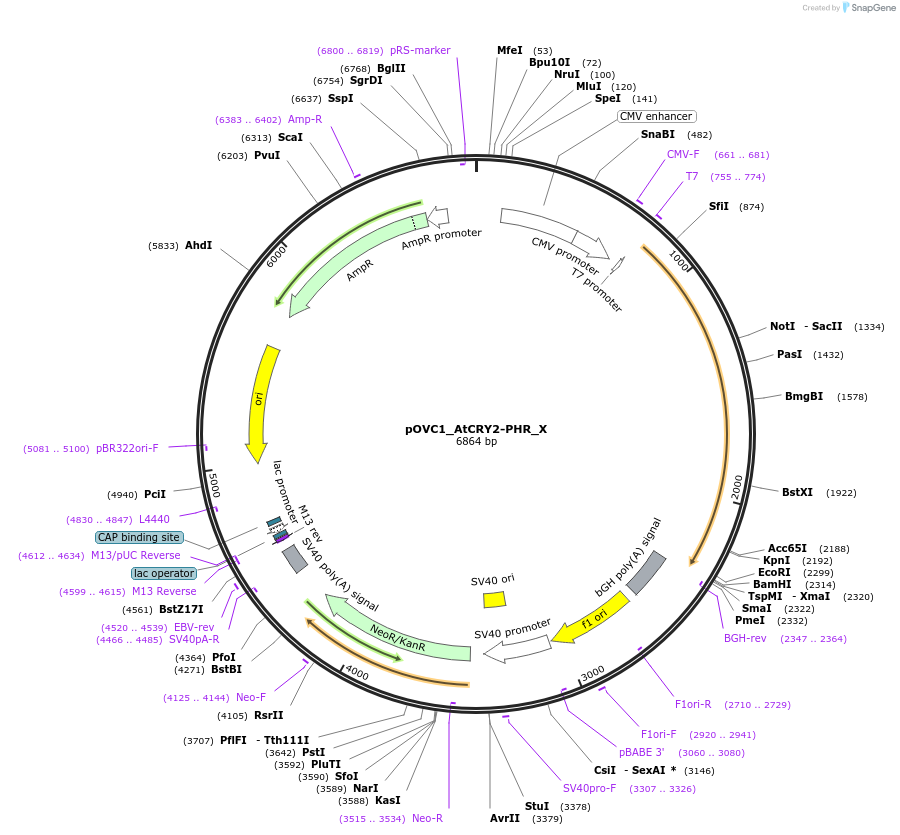
pOVC1_AtCRY2-PHR_X
Plasmid#131180PurposeExpresses AtCRY2-PHR (Arabidopsis thaliana) followed by restriction sitesDepositorInsertAtCRY2-PHR
ExpressionMammalianPromoterCMVAvailable SinceNov. 1, 2019AvailabilityAcademic Institutions and Nonprofits only -
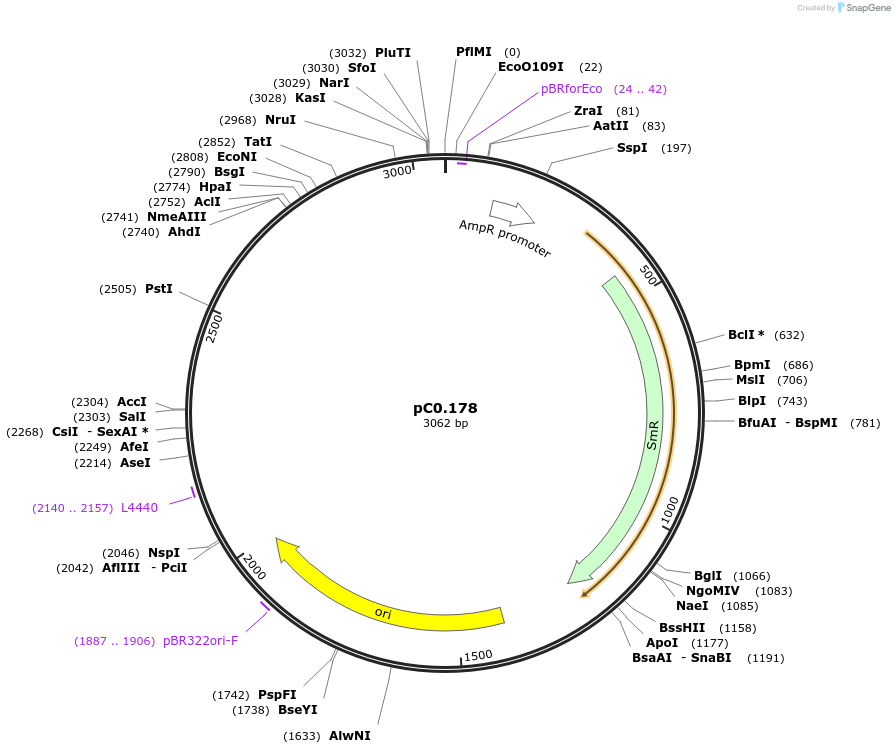
pC0.178
Plasmid#119645PurposeLevel 0 Part. Insertion siteDepositorInsert7942 NS3 Down Flank
UseSynthetic BiologyMutation4 nucleotide insertion between nt 65 and 66 in NS…Available SinceOct. 2, 2019AvailabilityAcademic Institutions and Nonprofits only -
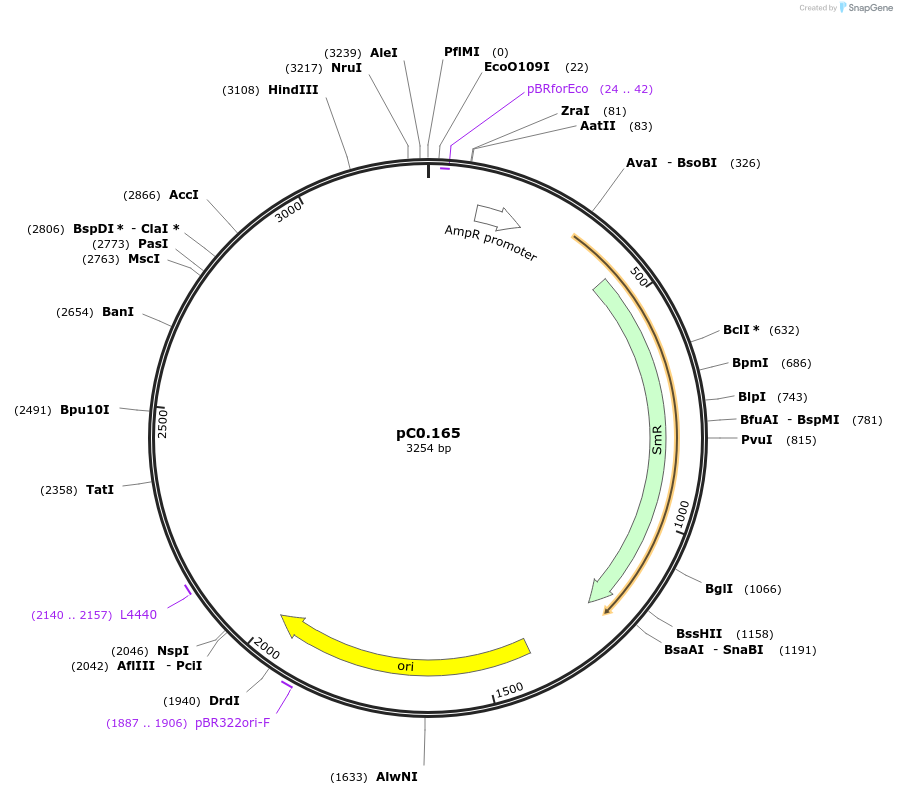
pC0.165
Plasmid#119634PurposeLevel 0 Part. Insertion siteDepositorInsert6803 NS1 Up Flank
UseSynthetic BiologyMutation1 nucleotide insertion between nt 21 and 22 in NS…Available SinceOct. 2, 2019AvailabilityAcademic Institutions and Nonprofits only -
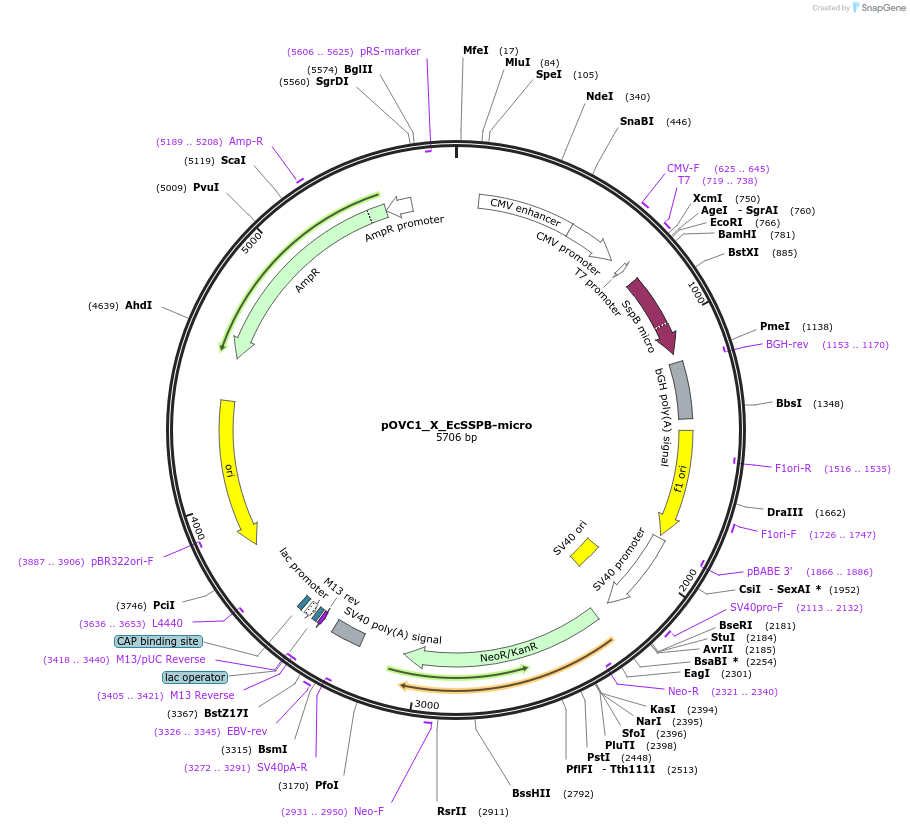
pOVC1_X_EcSSPB-micro
Plasmid#131218PurposeExpresses EcSSPB-nano (Escherichia coli) preceded by restriction sitesDepositorInsertEcSSPB-micro
ExpressionMammalianPromoterCMVAvailable SinceSept. 18, 2019AvailabilityAcademic Institutions and Nonprofits only -
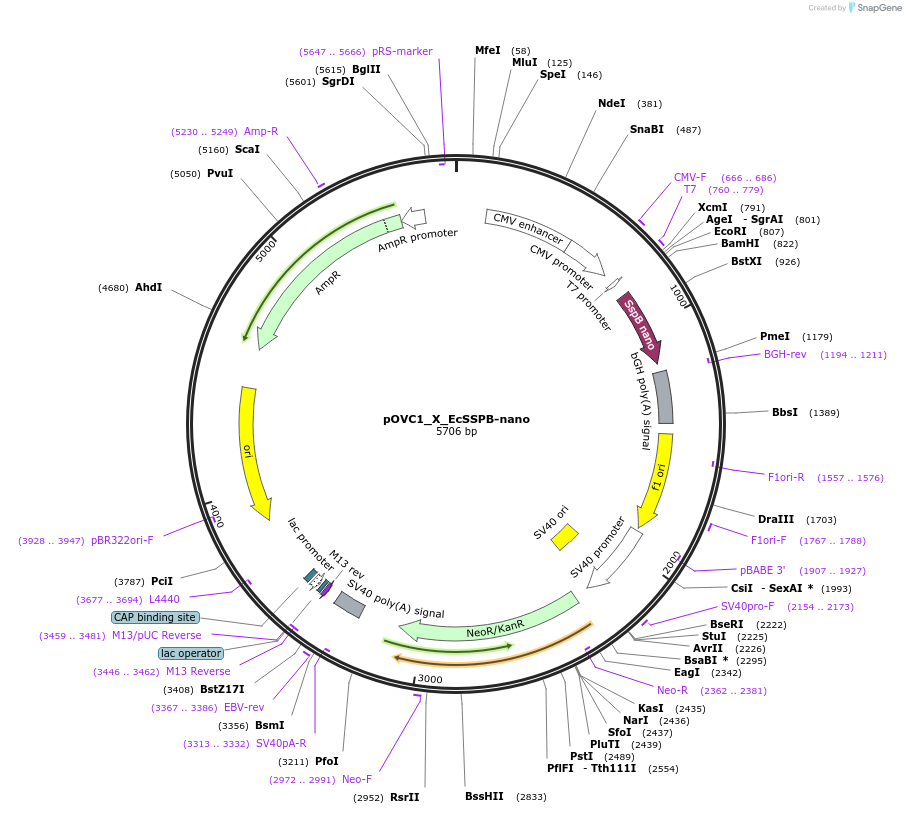
pOVC1_X_EcSSPB-nano
Plasmid#131219PurposeExpresses EcSSPB-micro (Escherichia coli) preceded by restriction sitesDepositorInsertEcSSPB-nano
ExpressionMammalianPromoterCMVAvailable SinceSept. 18, 2019AvailabilityAcademic Institutions and Nonprofits only -
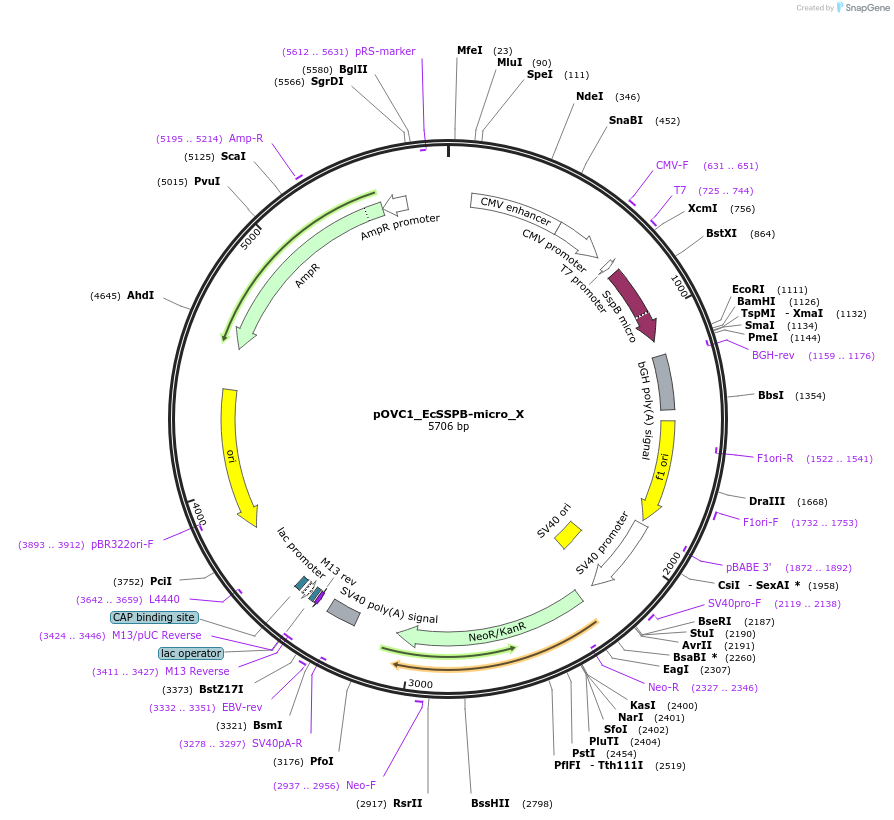
pOVC1_EcSSPB-micro_X
Plasmid#131213PurposeExpresses EcSSPB-micro (Escherichia coli) followed by restriction sitesDepositorInsertEcSSPB-micro
ExpressionMammalianPromoterCMVAvailable SinceSept. 18, 2019AvailabilityAcademic Institutions and Nonprofits only -
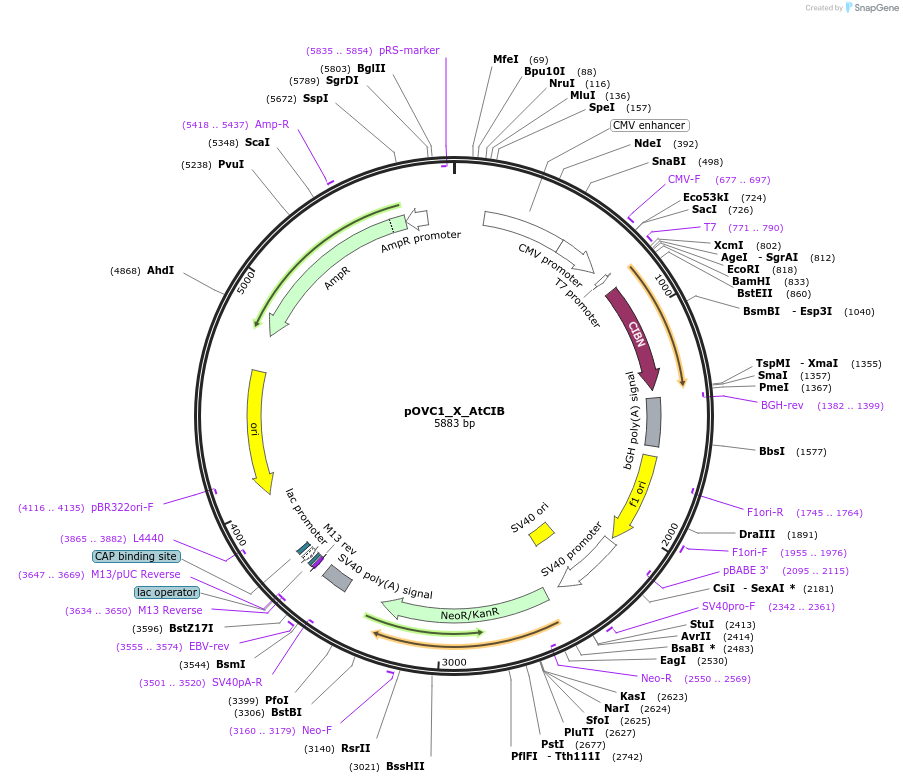
pOVC1_X_AtCIB
Plasmid#131194PurposeExpresses AtCIB (Arabidopsis thaliana) preceded by restriction sitesDepositorInsertAtCIB
ExpressionMammalianPromoterCMVAvailable SinceSept. 17, 2019AvailabilityAcademic Institutions and Nonprofits only -
pOVC1_X_AtPIF6
Plasmid#131192PurposeExpresses AtPIF6 (Arabidopsis thaliana) preceded by restriction sitesDepositorInsertAtPIF6
ExpressionMammalianPromoterCMVAvailable SinceSept. 17, 2019AvailabilityAcademic Institutions and Nonprofits only