We narrowed to 31,572 results for: REP
-
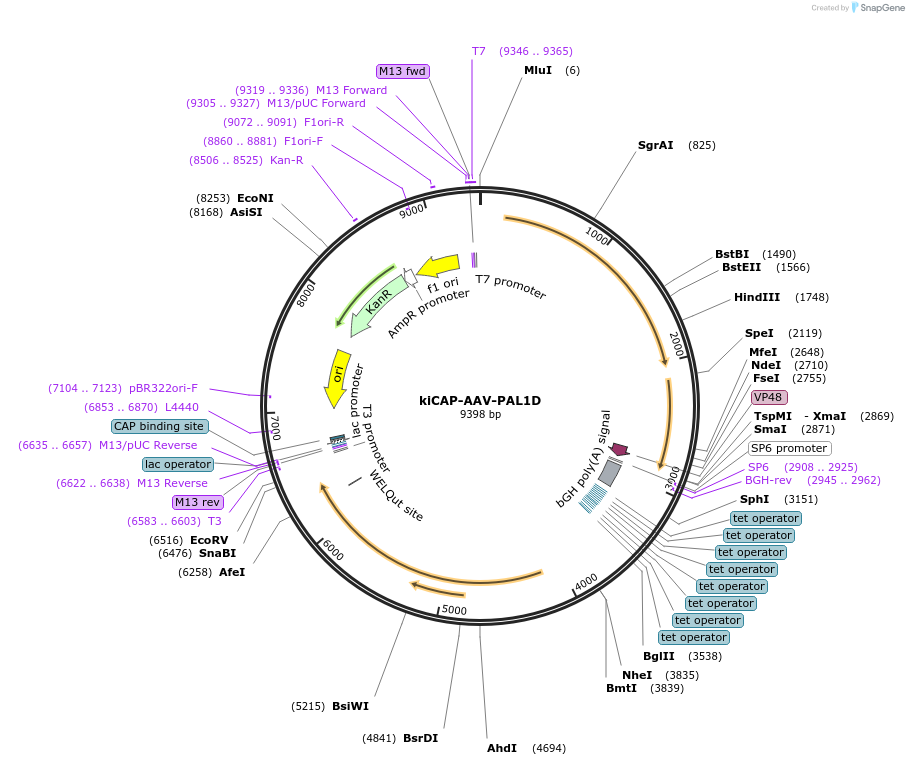
Plasmid#196686PurposeRep/Cap plasmid for the production of PAL1D, an AAV capsid with CNS tropism in macaques.DepositorInsertAAV9 VP1 modified with 7mer insertion between amino acids 588 and 589
UseAAVMutationPTQGTFR insert between amino acids 588 and 589 of…Promoterp41Available SinceApril 7, 2023AvailabilityAcademic Institutions and Nonprofits only -
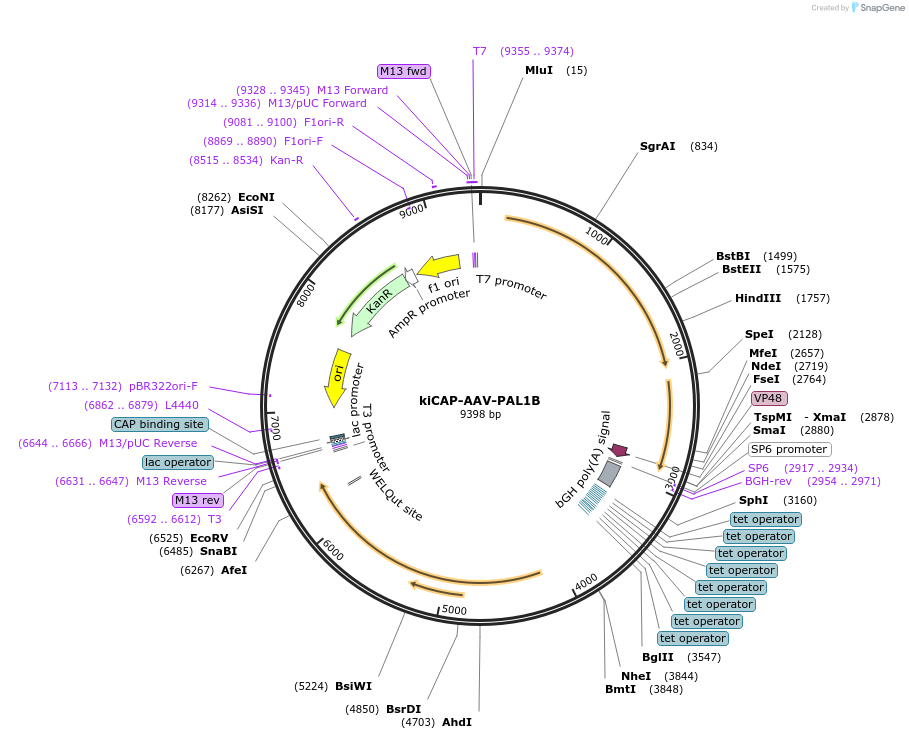
kiCAP-AAV-PAL1B
Plasmid#196684PurposeRep/Cap plasmid for the production of PAL1B, an AAV capsid with CNS tropism in macaques.DepositorInsertAAV9 VP1 modified with 7mer insertion between amino acids 588 and 589
UseAAVMutationPSQGTLR insert between amino acids 588 and 589 of…Promoterp41Available SinceApril 7, 2023AvailabilityAcademic Institutions and Nonprofits only -
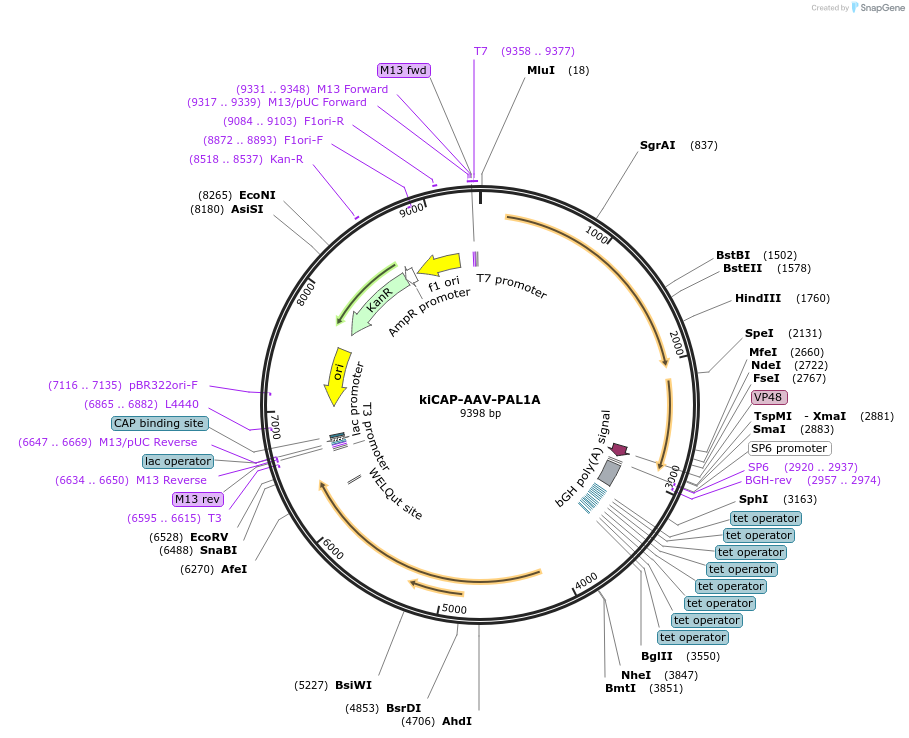
kiCAP-AAV-PAL1A
Plasmid#196683PurposeRep/Cap plasmid for the production of PAL1A, an AAV capsid with CNS tropism in macaques.DepositorInsertAAV9 VP1 modified with 7mer insertion between amino acids 588 and 589
UseAAVMutationPTQGTVR insert between amino acids 588 and 589 of…Promoterp41Available SinceApril 7, 2023AvailabilityAcademic Institutions and Nonprofits only -
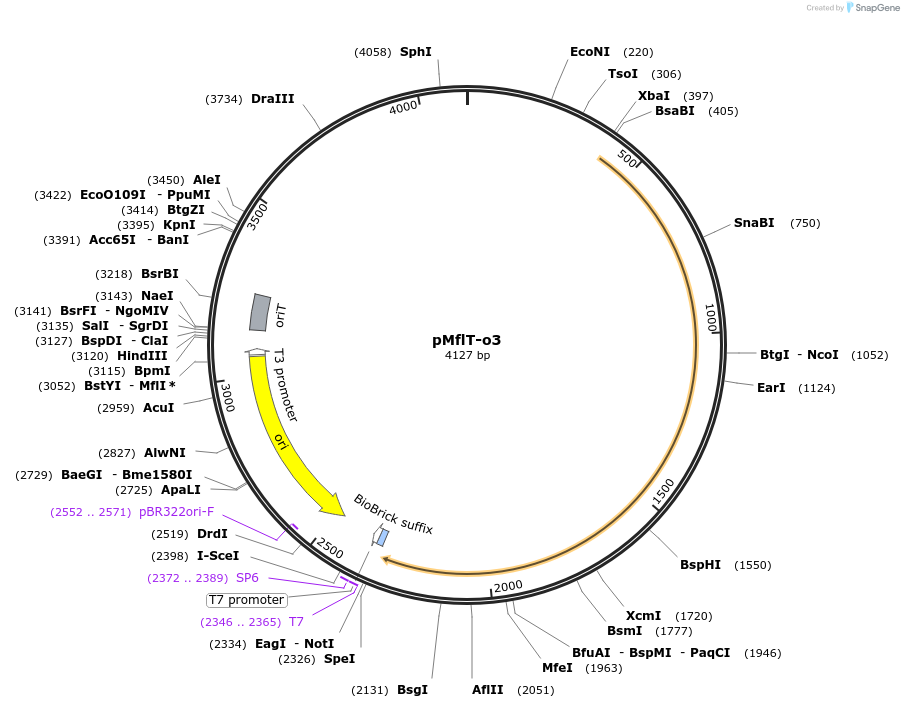
pMflT-o3
Plasmid#101311PurposeContains: oriC3 fragment of M. florum L1 replication origin (rpmH/dnaA and dnaA/dnaN intergenic regions, w/o dnaA), colE1 rep origin, recoded tetM resistance gene, origin of transfer of RP4 plasmid.DepositorInsertoriC3 of M. florum strain L1 (rpmH/dnaA and dnaA/dnaN intergenic regions, w/o dnaA)
UseSynthetic BiologyExpressionBacterialAvailable SinceNov. 7, 2017AvailabilityAcademic Institutions and Nonprofits only -
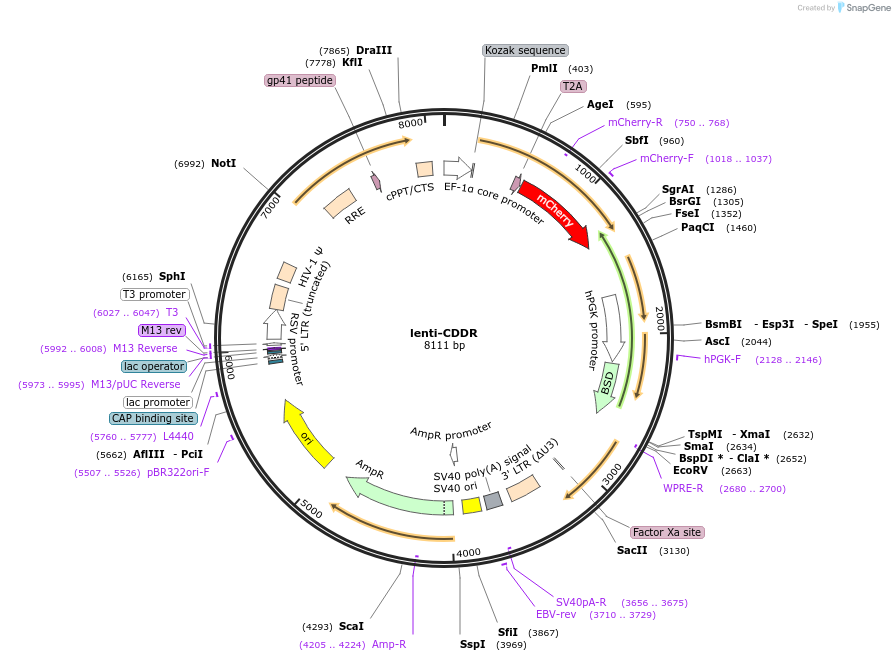
lenti-CDDR
Plasmid#149355PurposeCRISPR/Cas9-based dual-fluorescent DSB repair reporterDepositorInsertsCDDR Reporter
Blasticidin resistance
UseLentiviralExpressionMammalianPromoterEF-1a and hPGKAvailable SinceOct. 7, 2020AvailabilityAcademic Institutions and Nonprofits only -
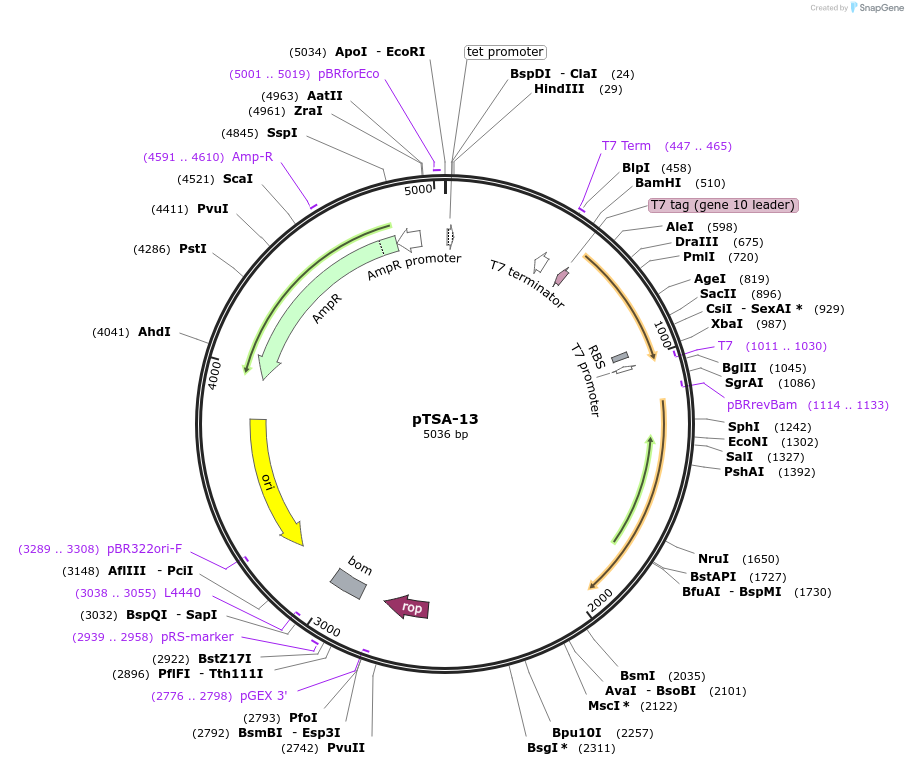
pTSA-13
Plasmid#17327DepositorInsertstreptavidin
ExpressionBacterialMutationTruncated to include only amino acids 16-133 of m…Available SinceFeb. 25, 2008AvailabilityAcademic Institutions and Nonprofits only -
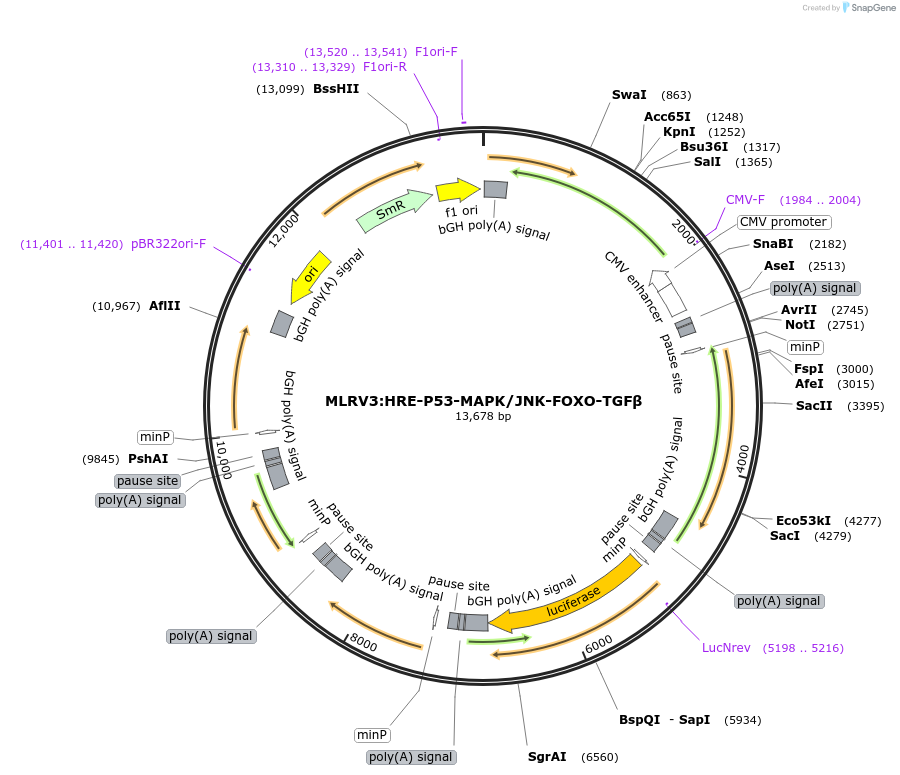
MLRV3:HRE-P53-MAPK/JNK-FOXO-TGFβ
Plasmid#178320PurposeMultiplex luciferase reporter vector, with luciferase reporters for the pathways HRE, P53, AP-1, FOXO and SMAD.DepositorInsertsHRE RedFirefly reporter
P53 FLuc reporter
AP-1 Renilla reporter
FOXO NLuc Reporter
SMAD GrRenilla reporter
UseLuciferase and Synthetic BiologyExpressionMammalianAvailable SinceJuly 20, 2022AvailabilityAcademic Institutions and Nonprofits only -
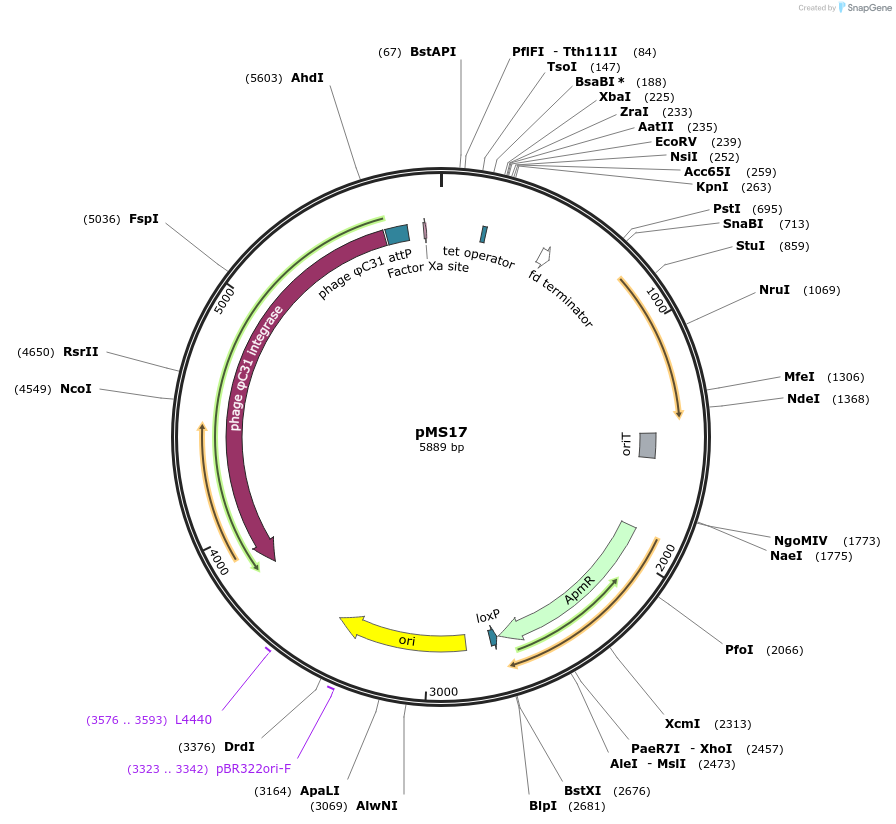
pMS17
Plasmid#127088PurposeIntegrating vector containing a tetracycline inducible promoter for use in Streptomyces sp.DepositorInsertstcp830
TetRiS
terminator mmrt
Terminator fdt
UseStreptomyce integration vector derived from phic3…Available SinceJuly 10, 2019AvailabilityAcademic Institutions and Nonprofits only -
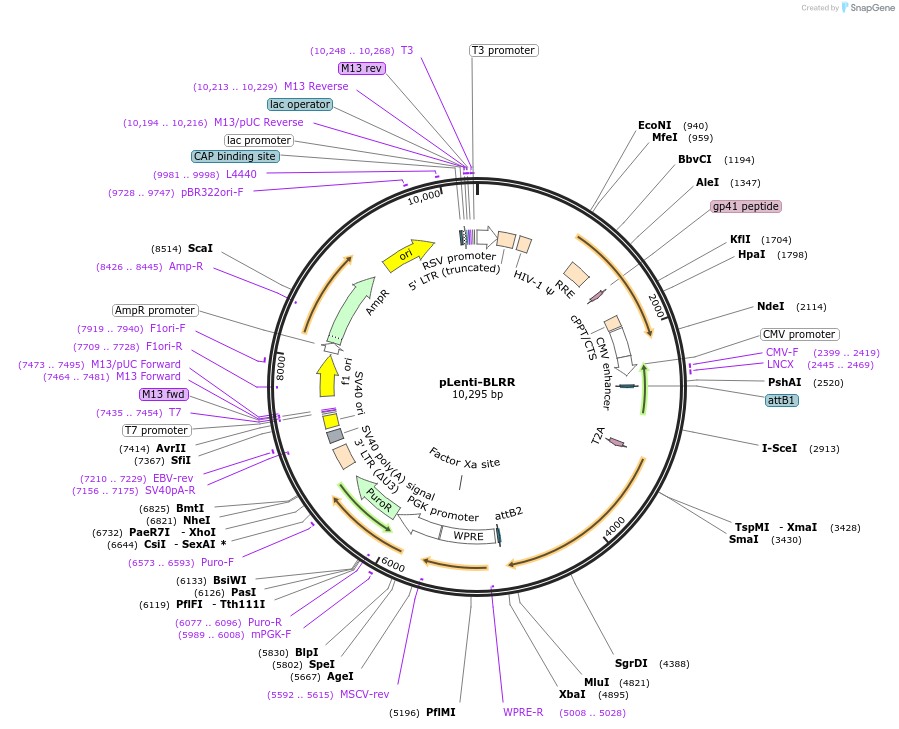
pLenti-BLRR
Plasmid#158958PurposeBioluminescent repair reporter (BLRR) for DNA double strand break (DSB) repairs: homology-directed repair (HDR) and non-homologous end joining (NHEJ).DepositorInsertBLRR
UseLentiviralExpressionMammalianPromoterCMVAvailable SinceNov. 10, 2020AvailabilityAcademic Institutions and Nonprofits only -
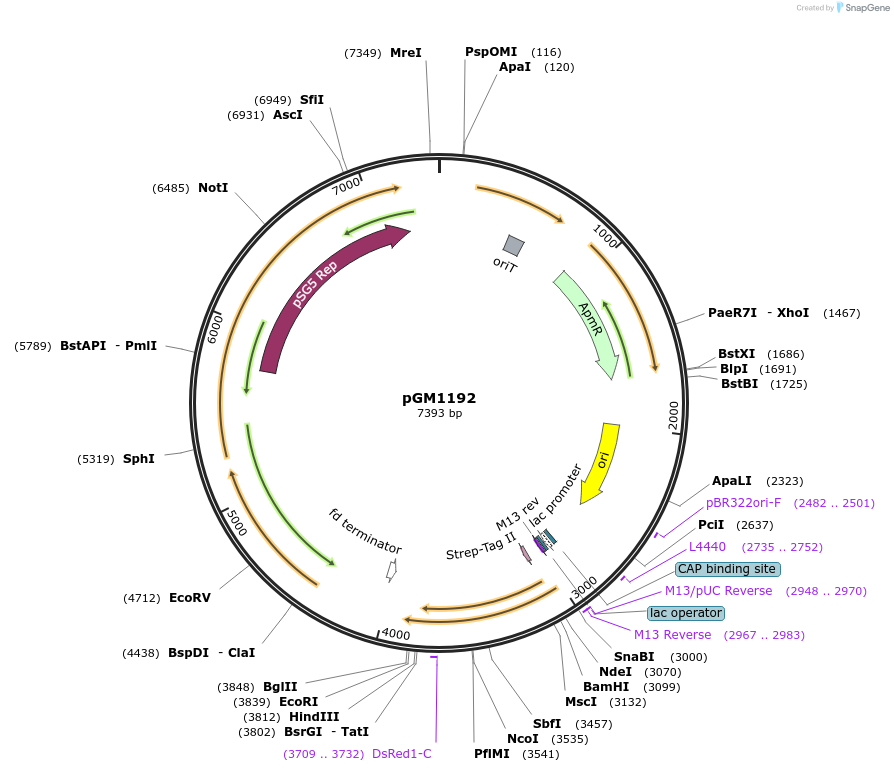
pGM1192
Plasmid#115683PurposeE. coli-Streptomyces shuttle vector for gene expressionDepositorInsertmCherry, aac(3)IV, tsr
TagsStrep-tagII on mCherry, codon optimizedExpressionBacterialMutationmCherry codon optimized for StreptomycesPromoterSF14*Available SinceFeb. 27, 2019AvailabilityAcademic Institutions and Nonprofits only -
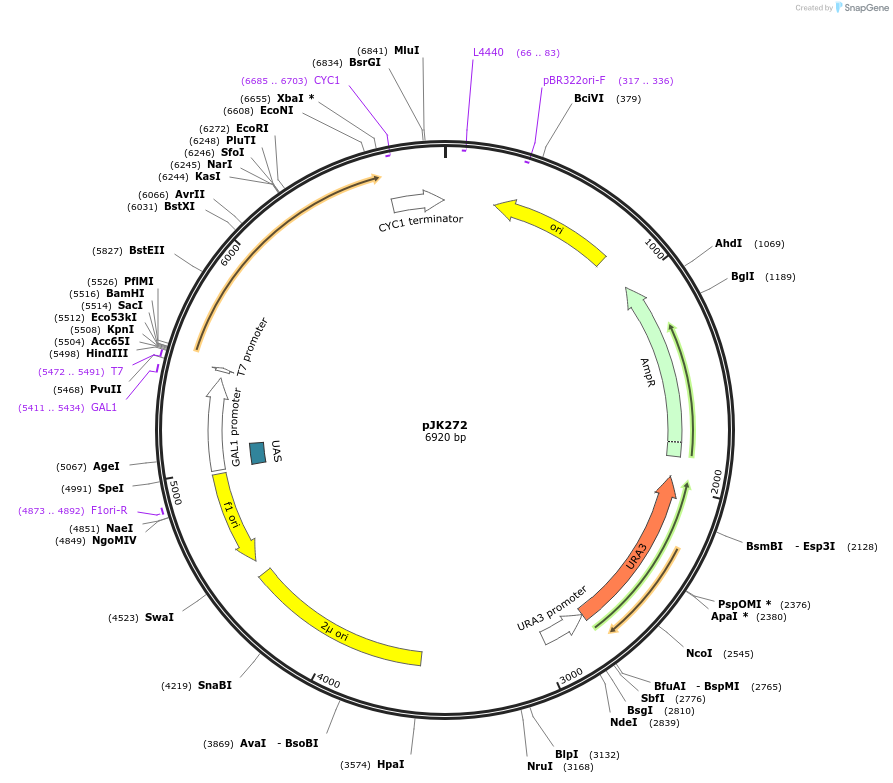
pJK272
Plasmid#71562PurposeProduces Crepis alpina delta12 fatty acid acetylenase (vFAD2) in yeastDepositorInsertdelta12 fatty acid acetylenase
ExpressionBacterial and YeastPromoterGAL1Available SinceJan. 5, 2016AvailabilityAcademic Institutions and Nonprofits only -
pTSA-38
Plasmid#17328DepositorInsertstreptavidin
ExpressionBacterialMutationTruncated to include only amino acids 16-133 of m…Available SinceFeb. 25, 2008AvailabilityAcademic Institutions and Nonprofits only -
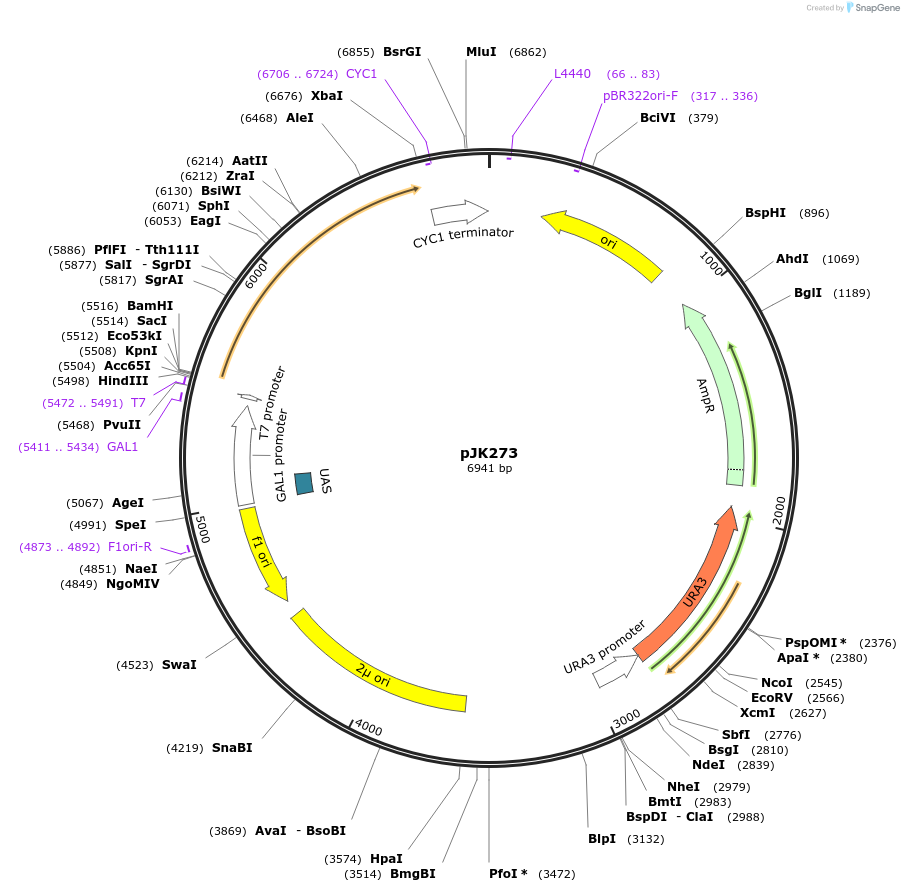
pJK273
Plasmid#71563PurposeProduces Crepis alpina delta12 fatty acid desaturase (FAD2-2) in yeastDepositorInsertdelta12 fatty acid desaturase (form 2)
ExpressionBacterial and YeastPromoterGAL1Available SinceJan. 5, 2016AvailabilityAcademic Institutions and Nonprofits only -
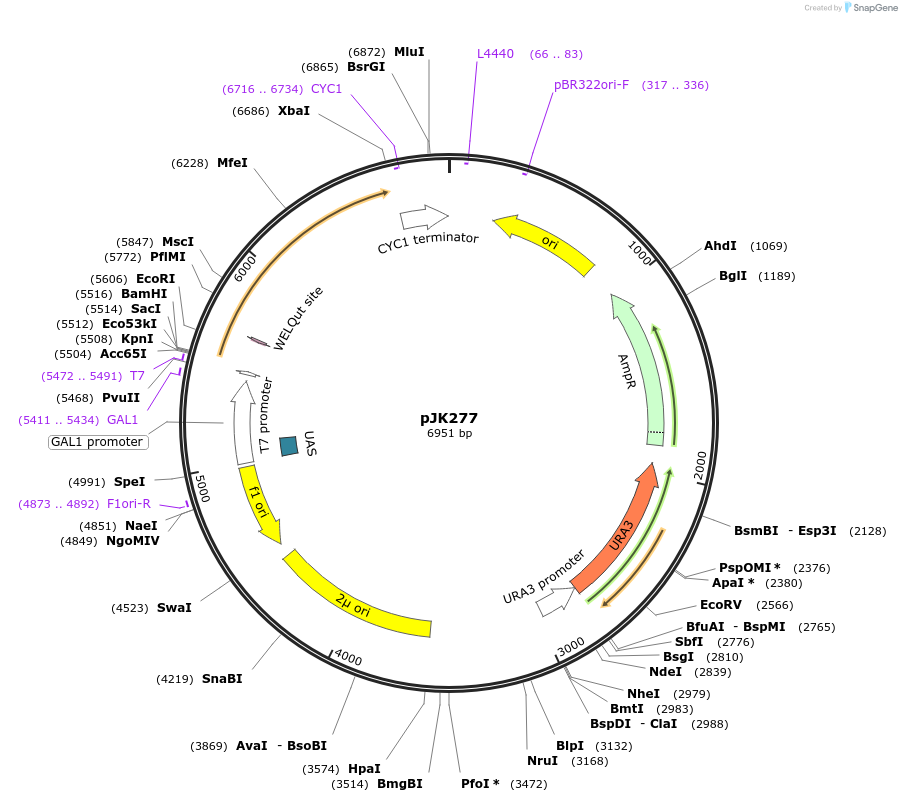
pJK277
Plasmid#71565PurposeProduces Crepis alpina delta15 fatty acid desaturase (FAD3) in yeastDepositorInsertdelta15 fatty acid desaturase
ExpressionBacterial and YeastPromoterGAL1Available SinceJan. 14, 2016AvailabilityAcademic Institutions and Nonprofits only -
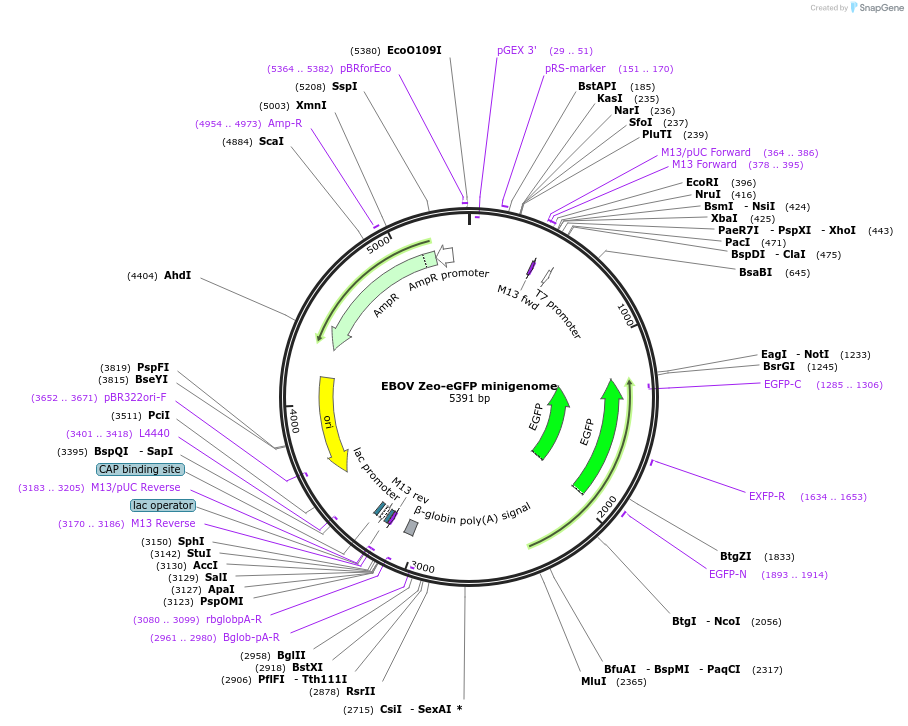
EBOV Zeo-eGFP minigenome
Plasmid#249393PurposeT7-driven EBOV minigenome vRNA with Zeo-eGFP reporter selection cassetteDepositorInsertZeocin-eGFP fusion gene flanked by EBOV 5'UTR and 3'UTR
ExpressionBacterialPromoterT7Available SinceJan. 15, 2026AvailabilityAcademic Institutions and Nonprofits only -
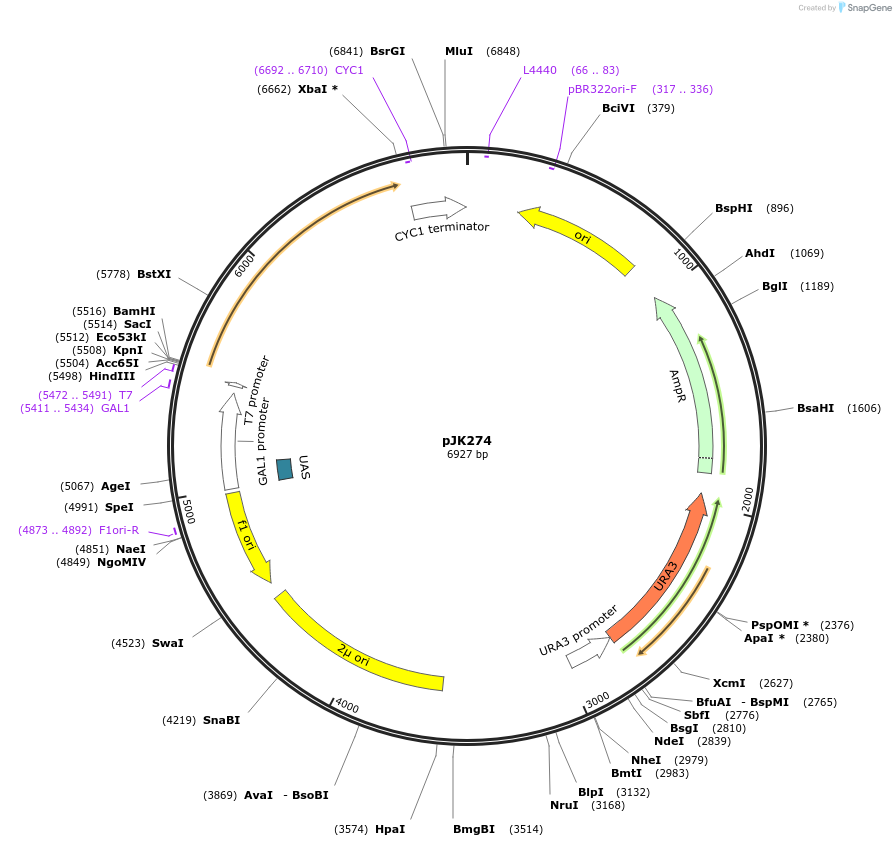
pJK274
Plasmid#71564PurposeProduces Crepis alpina delta12 fatty acid desaturase (FAD2-3) in yeastDepositorInsertdelta12 fatty acid desaturase (form 3)
ExpressionBacterial and YeastPromoterGAL1 promoterAvailable SinceJan. 5, 2016AvailabilityAcademic Institutions and Nonprofits only -
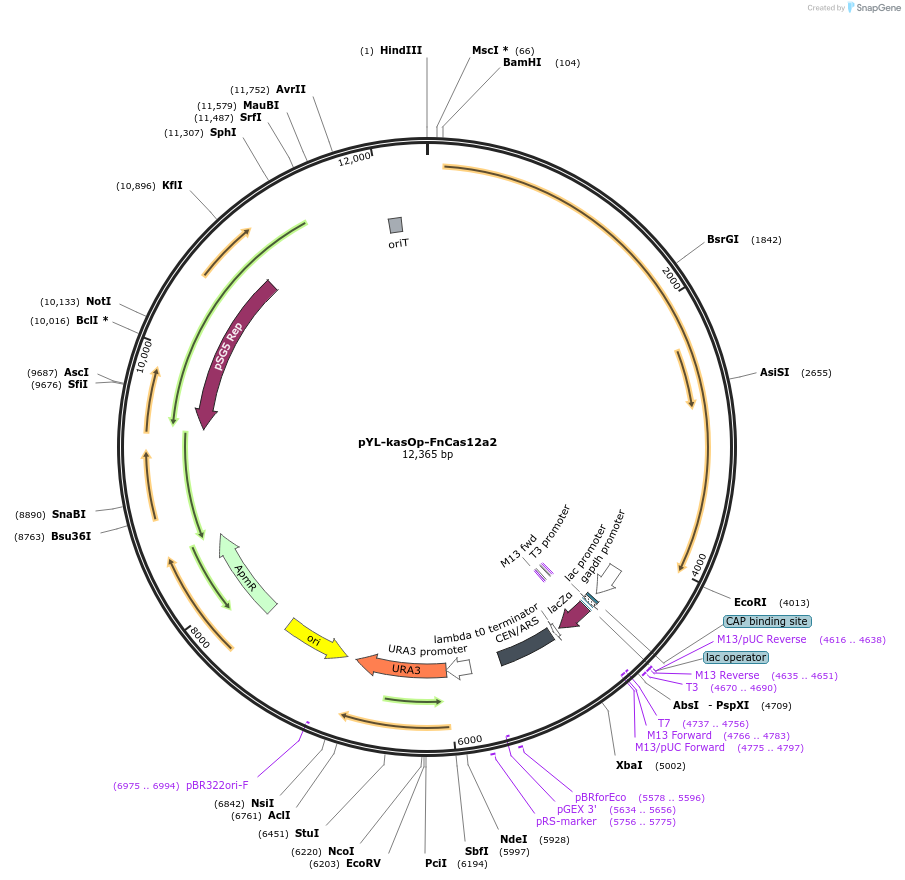
pYL-kasOp-FnCas12a2
Plasmid#177874PurposeExpresses Streptomyces codon optimized FnCas12a. Used to modify genome of Streptomyces.DepositorInsertStreptomyces codon optimized FnCas12a
UseCRISPRPromoterkasOp*Available SinceJan. 13, 2022AvailabilityAcademic Institutions and Nonprofits only -
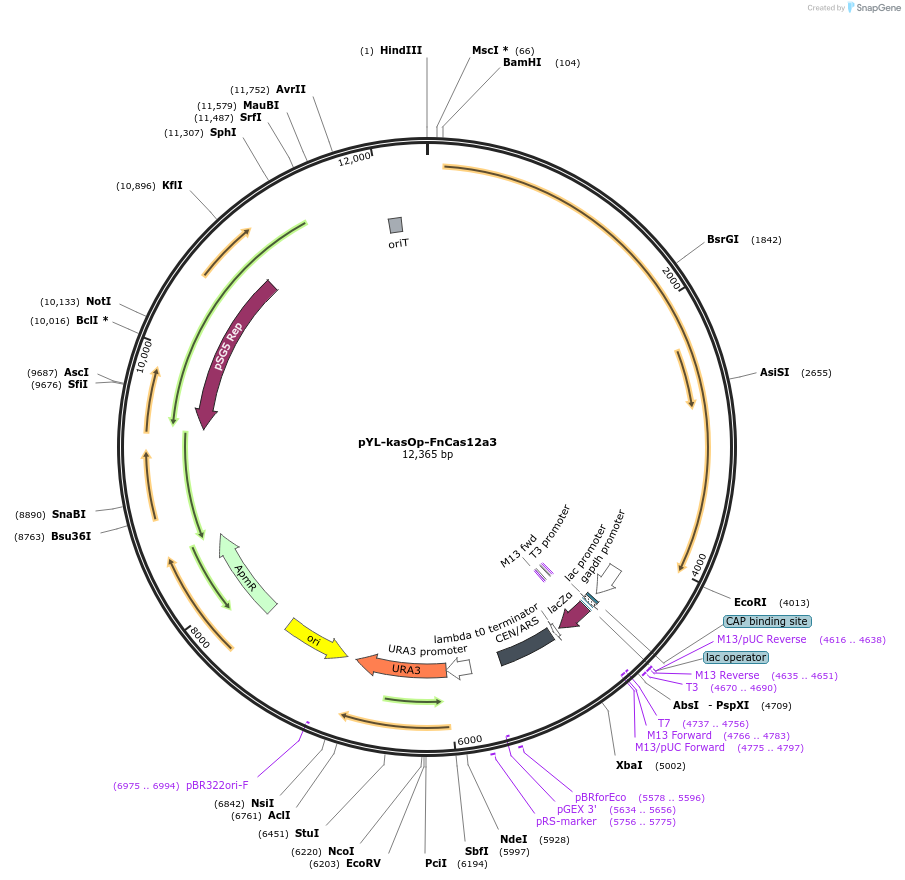
pYL-kasOp-FnCas12a3
Plasmid#177875PurposeExpresses Streptomyces codon optimized FnCas12a. Used to modify genome of Streptomyces.DepositorInsertStreptomyces codon optimized the FnCas12a mutant EP16
UseCRISPRPromoterkasOp*Available SinceJan. 13, 2022AvailabilityAcademic Institutions and Nonprofits only -
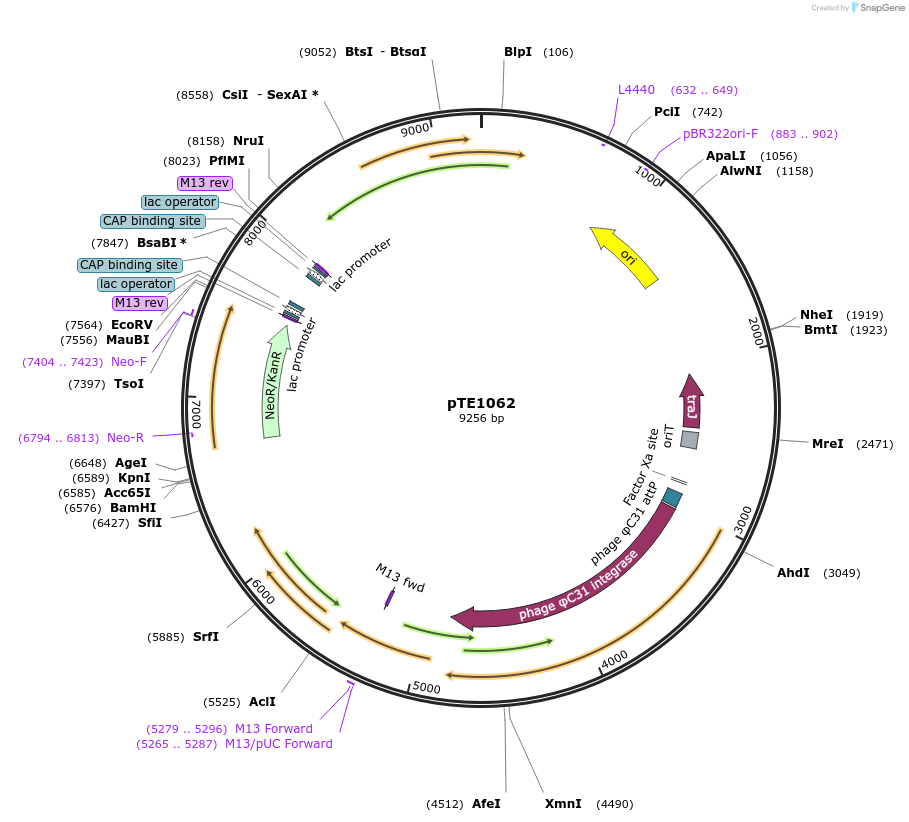
pTE1062
Plasmid#172356PurposeDetection of Streptomyces coelicolor butyrolactones (SCB) by kanamycin bioassay. Streptomyces genomic integrative vector.DepositorInsertsUseSynthetic Biology; Integrative into streptomyces …ExpressionBacterialAvailable SinceDec. 17, 2021AvailabilityAcademic Institutions and Nonprofits only -
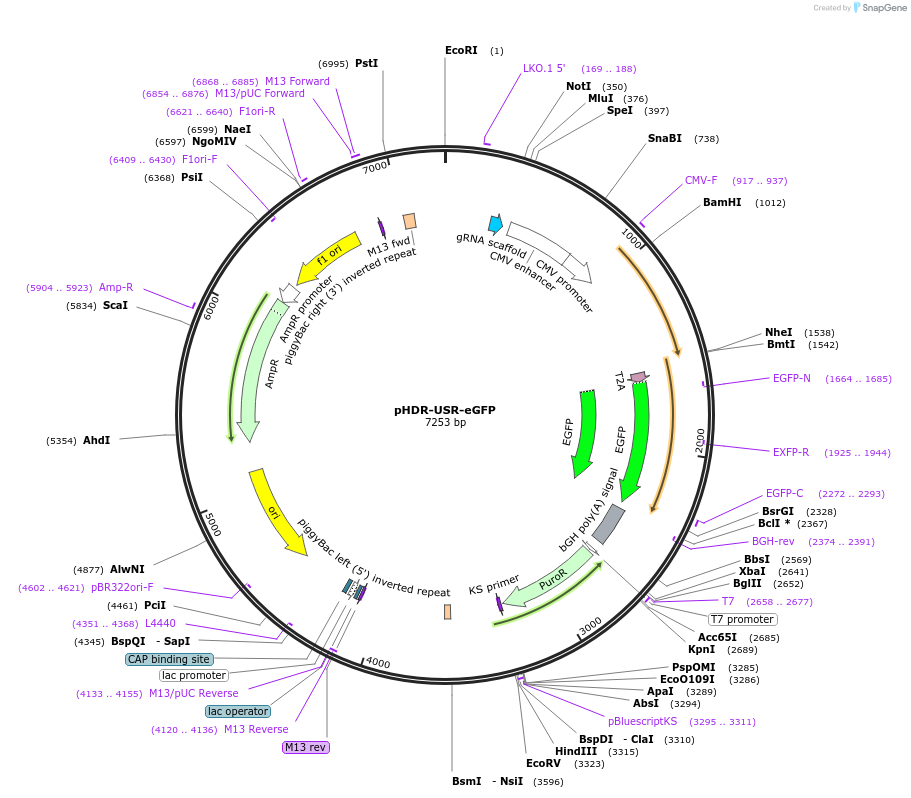
pHDR-USR-eGFP
Plasmid#208830PurposeHDR Universal Surrogate Reporter with a sgRNA and consistently expressed eGFP but without Cas9. The selective PuroR reporter gene was designed to be repaired only by the sgRNA/Cas9-triggered HDR.DepositorTypeEmpty backboneExpressionMammalianPromoterCMV, U6Available SinceNov. 16, 2023AvailabilityAcademic Institutions and Nonprofits only