Rinehart Lab Phosphoprotein Reagents
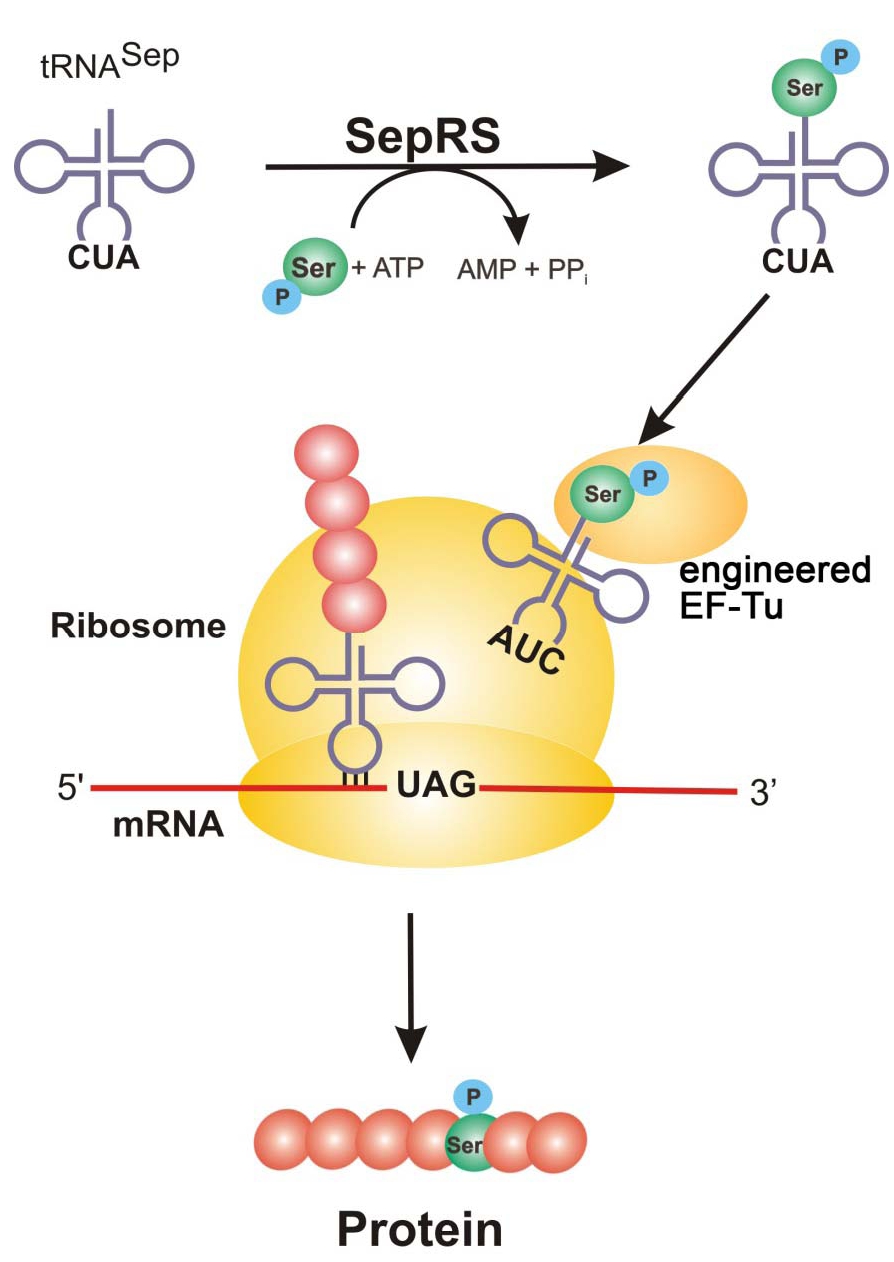
Figure 1: SepRS adds a phosphoserine to tRNA^Sep with anticodon, CUA. Engineered EF-Tu binds the phosphoserine-charged tRNA (AUC) and is brought to the ribosome where it recognizes the UAG amber codon and incorporates phosphoserine into the growing polypeptide chain to generate the new phosphorylated protein.
Protein Phosphorylation
Protein phosphorylation is one of the most abundant forms of posttranslational modification in cells and research into its many roles in protein function and signaling networks continues to expand. The Rinehart and Söll labs at Yale University changed the way researchers can explore important questions surrounding serine phosphorylation by adding this phosphorylated amino acid to the genetic code of E. coli (Park et al., 2011). Follow up work from the Rinehart and Issacs labs at Yale have made improvements to this system by genomically recoding E.coli (Lajoie et al., 2013) and improving the orthogonal translation system to make phosphorylated proteins with one or more phosphorylation sites (Pirman et al., 2015).
Improved phosphoprotein production with rEcoli XpS and a new pSerOTS
The Rinehart lab developed a new strain of recoded E.coli, rEcoli XpS (Bacterial strain #192872), and an optimized phosphoserine orthogonal translation system, pSerOTS-C1* (V70) (Plasmid #188537). These reagents are described in detail in Mohler et al. (2023) and help deliver improved phosphoprotein yield, rEcoli growth characteristics, and an overall improved ease of use when expressing recombinant phosphoproteins. These attributes make rEcoli XpS (Bacterial Strain #192872) and pSerOTS-C1* (V70) (Plasmid #188537) a better choice in place of the popular SepOTSλ (Plasmid #68292).
An important technical note: pSerOTS-C1* (V70) is a ROP minus plasmid and is not compatible with other plasmids containing high copy ColE1/PUC-like origins. Therefore, when choosing plasmids to express the phosphoprotein of interest, a medium copy RSF1030 origins or a low copy p15a origins should be used. Find further details in Mohler et al. (2023) and find general tips on phosphoprotein expression in rEcoli XpS in the Methods for iSPI, rEcoli XPS, pSerOTS-C1* (V70) (DOCX, 551 KB).
The synthetic human phosphoproteome
The Rinehart lab synthesized 110,139 unique DNA sequences corresponding to every previously observed instance of human serine phosphorylation (Rinehart Human Serine Phosphopeptide Library). The phosphorylation sites (“phosphosites”) consist of a central phosphoserine site flanked by 15 amino acids on either side occurring within the parent protein. Using the recoded strain of E. coli, C321.ΔA (Bacterial strain #68306), with the optimized phosphoserine orthogonal translation system, SepOTSλ (Plasmid #68292), they were able to demonstrate unambiguous site-specific incorporation of phosphoserine in >36,000 phosphosites by tandem mass spectrometry (Barber et al., 2018). This phosphosite library was generated in a single mixed pool by expressing and purifying the phosphosite library as GST-fusion peptides (“Mode #1” expression). The Mode #1 phosphosite library (Pooled Library #111704) is anticipated to be useful for laboratories performing phosphoproteomic experiments in need of reference spectra or phosphopeptide standards for quantitative assessments. The Mode #1 phosphosite library can be generated with either phosphoserine using SepOTSλ (Plasmid #68292) or serine using supD tRNA (SupD (Plasmid #68307)) at the central position.
Introducing Hi-P for screening phosphorylation-dependent protein-protein interactions

Figure 2: Mode #1 and Mode #2 phosphosite library configuration for overexpression in E. coli.
The phosphosite DNA library can also be introduced into a second expression vector encoding a split mCherry fluorescent reporter, enabling the detection of phosphorylation-dependent protein-peptide interactions (“Mode #2” expression, Barber et al., 2018). The phosphosite library is fused to the N-terminal portion of split mCherry, while a phosphobinding domain is fused to the C-terminal mCherry fragment. Upon interaction between the phosphobinding domain and the phosphosite, mCherry fluorescence is restored, so interactions can be discovered by fluorescence-activated cell sorting (FACS). The Mode #2 phosphosite library contains four different phosphobinding domains: 14-3-3β, 14-3-3σ, the WW2 domain of NEDD4, and the WW2 domain of NEDD4-2 (Pooled Libraries #111705-8).
Screening kinases
The Mode #1 phosphosite library can also be synthesized using supD tRNA (SupD (Plasmid #68307)) to generate the phosphosites with serine instead of phosphoserine (i.e., a “phosphorylatable” phosphosite library). This serine phosphosite library can then be used to identify candidate human substrates of kinases of interest by mass spectrometry in a technique called serine-oriented library/kinase-library reactions (SERIOHL-KILR, Barber et al., 2018).
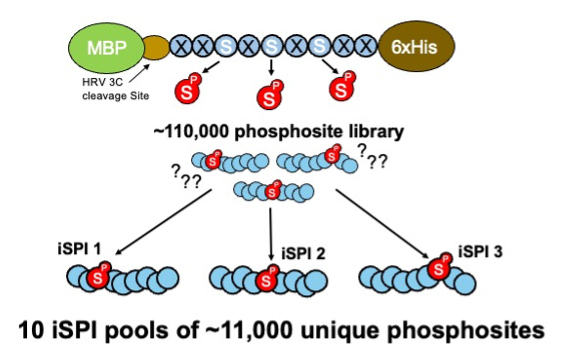
Figure 3: The library of ~110,000 phosphosites is subdivided into 10 iSPI subpools of ~11,000 unique phosphosites.
Iterative Synthetically Phosphorylated Isomers (iSPI)
The iSPI resource described in Galloway et al. (2022) is adapted from the synthetic human phosphoproteome. It subdivides the 110,139 human phosphoserine site library (Pooled Library #188536) into 10 subpools of ~11,000 phosphosites (Pooled Libraries #188526; #188527; #188528; #188529; #188530; #188531; #188532; #188533; #188534; #188535). Importantly, phosphorylation positional isomers are allocated to different subpools, allowing precise knowledge of phosphorylated positions for testing phosphoproteomics methods, proteomics search algorithms, proteomics phosphosite localization algorithms, or any method confounded by phosphosite ambiguity.
Rinehart Lab Improved Phosphoprotein Synthesis Reagents
| Type | ID | Item |
|---|---|---|
| Strain | 192872 | rEcoli XpS (MG1655-C321 mutS+, λ-, Δ(ybhB-bioAB)::zeoR, ΔprfA, ΔserB) |
| Plasmid | 188537 | pSerOTS-C1* (V70) |
| Pooled Library | 188536 | iSPI_pSer_Full_library |
| Pooled Library | 188535 | iSPI_pSer_Subpool#10 |
| Pooled Library | 188534 | iSPI_pSer_Subpool#9 |
| Pooled Library | 188533 | iSPI_pSer_Subpool#8 |
| Pooled Library | 188532 | iSPI_pSer_Subpool#7 |
| Pooled Library | 188531 | iSPI_pSer_Subpool#6 |
| Pooled Library | 188530 | iSPI_pSer_Subpool#5 |
| Pooled Library | 188529 | iSPI_pSer_Subpool#4 |
| Pooled Library | 188528 | iSPI_pSer_Subpool#3 |
| Pooled Library | 188527 | iSPI_pSer_Subpool#2 |
| Pooled Library | 188526 | iSPI_pSer_Subpool#1 |
| Plasmid | 112031 | pNAS1b split mCherry 14-3-3β ctrl |
| Plasmid | 112030 | pNAS1b split mCherry NEDD4 WW2 neg ctrl |
| Plasmid | 112029 | pNAS1b split mCherry NEDD4 WW2 pos ctrl |
| Plasmid | 111883 | pNAS1b split mCherry NEDD4-2 WW2 |
| Plasmid | 111882 | pNAS1b split mCherry NEDD4 WW2 |
| Plasmid | 111881 | pNAS1b split mCherry 14-3-3σ |
| Plasmid | 111880 | pNAS1b split mCherry 14-3-3β |
| Pooled Library | 111708 | Mode #2 Library (NEDD4-2 WW2 domain) |
| Pooled Library | 111707 | Mode #2 Library (NEDD4 WW2 domain) |
| Pooled Library | 111706 | Mode #2 Library (14-3-3σ) |
| Pooled Library | 111705 | Mode #2 Library (14-3-3β) |
| Pooled Library | 111704 | Mode #1 Library |
| Plasmid | 69118 | E17TAG GFP zeo resistance |
| Plasmid | 68307 | SupD |
| Strain | 68306 | C321.ΔA.Δserb.Amp |
| Plasmid | 68305 | Beta lactamase S68TAG |
| Plasmid | 68302 | MBP-MEK1 S222TAG |
| Plasmid | 68301 | MBP-MEK1 S218TAG |
| Plasmid | 68300 | MBP-MEK1 |
| Plasmid | 68299 | GFP E17TAG/Q157TAG |
| Plasmid | 68298 | GFP S2TAG/E17TAG |
| Plasmid | 68297 | GFP Q157TAG |
| Plasmid | 68296 | GFP S2TAG |
| Plasmid | 68295 | GFP E17TAG |
| Plasmid | 68294 | SepOTSν |
| Plasmid | 68292 | SepOTSλ |
| Plasmid | 68291 | SepOTSκ |
| Plasmid | 68290 | SepOTSι |
| Plasmid | 68289 | SepOTSθ |
| Plasmid | 68288 | SepOTSη |
| Plasmid | 68287 | SepOTSζ |
| Plasmid | 68286 | SepOTSϵ |
| Plasmid | 68285 | SepOTSδ |
| Plasmid | 68284 | SepOTSγ |
| Plasmid | 68283 | SepOTSβ |
| Plasmid | 53225 | MBP-MEK1 S218TAG/S222TAG (aka. PCRT7 tetR pLtetO MBP-MEK1 XX Amp) |
| Strain | 52055 | EcAR7 |
| Plasmid | 52054 | SepOTSµ (aka. B40 OTS) |
| Strain | 34929 | BL21ΔserB |
| Plasmid | 34624 | SepOTSα tRNA-Sep (aka. pCAT112TAG-SepT) |
| Plasmid | 34623 | SepOTSα SepRS/EF-Sep (aka. pKD-SepRS-EFSep) |
References & Protocols
Iterative Synthetically Phosphorylated Isomers (iSPI) described in:Gassaway, B. M., Li, J., Rad, R., Mintseris, J., Mohler, K., Levy, T., Aguiar, M., Beausoleil, S. A., Paulo, J. A., Rinehart, J., Huttlin, E. L., & Gygi, S. P. (2022). A multi-purpose, regenerable, proteome-scale, human phosphoserine resource for phosphoproteomics. Nature Methods, 19(11), 1371–1375. https://doi.org/10.1038/s41592-022-01638-5. PMID 36280721 (Link opens in a new window)
rEcoli XpS and a new pSerOTS described in:Mohler, K., Moen, J. M., Rogulina, S., & Rinehart, J. (2023). System‐wide optimization of an orthogonal translation system with enhanced biological tolerance. Molecular Systems Biology, 19(8). https://doi.org/10.15252/msb.202110591. PMID 37477096 (Link opens in a new window)
Protocols for using iSPI, rEcoli XPS, and pSerOTS-C1* (V70) (ca. 2022) from the Rinehart lab:Methods for iSPI, rEcoli XPS, pSerOTS-C1* (V70) (DOCX, 551 KB)
Human Serine Phosphopeptide Library described in:Barber, K. W., Muir, P., Szeligowski, R. V., Rogulina, S., Gerstein, M., Sampson, J. R., Isaacs, F. J., & Rinehart, J. (2018). Encoding human serine phosphopeptides in bacteria for proteome-wide identification of phosphorylation-dependent interactions. Nature Biotechnology, 36(7), 638–644. https://doi.org/10.1038/nbt.4150. PMID 29889213 (Link opens in a new window)
Barber, K. W., Miller, C. J., Jun, J. W., Lou, H. J., Turk, B. E., & Rinehart, J. (2018). Kinase substrate profiling using a proteome-wide Serine-Oriented human Peptide library. Biochemistry, 57(31), 4717–4725. https://doi.org/10.1021/acs.biochem.8b00410. PMID 29920078 (Link opens in a new window)
Improved Reagents (ca. 2015) described in:Pirman, N. L., Barber, K. W., Aerni, H. R., J, N., MA, Haimovich, A. D., Rogulina, S., Isaacs, F. J., & Rinehart, J. (2015). A flexible codon in genomically recoded Escherichia coli permits programmable protein phosphorylation. Nature Communications, 6(1). https://doi.org/10.1038/ncomms9130. PMID 26350500 (Link opens in a new window)
Oza, J. P., Aerni, H. R., Pirman, N. L., Barber, K. W., Ter Haar, C. M., Rogulina, S., Amrofell, M. B., Isaacs, F. J., Rinehart, J., & Jewett, M. C. (2015). Robust production of recombinant phosphoproteins using cell-free protein synthesis. Nature Communications, 6(1). https://doi.org/10.1038/ncomms9168. PMID 26350765 (Link opens in a new window)
Heo, J., Ordureau, A., Paulo, J. A., Rinehart, J., & Harper, J. W. (2015). The PINK1-PARKIN mitochondrial ubiquitylation pathway drives a program of OPTN/NDP52 recruitment and TBK1 activation to promote mitophagy. Molecular Cell, 60(1), 7–20. https://doi.org/10.1016/j.molcel.2015.08.016. PMID 26365381 (Link opens in a new window)
Protocol for using the Improved Reagents (ca. 2015) from the Rinehart lab:Users Guide for Improved 2015 Phosphoprotein Reagents (DOCX, 525.9 KB)
Reagents (ca. 2014) described in:Heinemann, I. U., Rovner, A. J., Aerni, H. R., Rogulina, S., Cheng, L., Olds, W., Fischer, J. T., Söll, D., Isaacs, F. J., & Rinehart, J. (2012). Enhanced phosphoserine insertion during Escherichia coli protein synthesis via partial UAG codon reassignment and release factor 1 deletion. FEBS Letters, 586(20), 3716–3722. https://doi.org/10.1016/j.febslet.2012.08.031. PMID 22982858 (Link opens in a new window)
Protocol for Reagents (ca. 2014) from the Rinehart lab:Users Guide for 2014 Phosphoprotein Reagents (PDF, 691.3 KB)
Original Kit (ca. 2011) described in:Park, H., Hohn, M. J., Umehara, T., Guo, L., Osborne, E. M., Benner, J., Noren, C. J., Rinehart, J., & Söll, D. (2011). Expanding the Genetic Code of Escherichia coli with Phosphoserine. Science, 333(6046), 1151–1154. https://doi.org/10.1126/science.1207203. PMID 21868676 (Link opens in a new window)
Additional Resources
- See our dedicated Genetic Code Expansion collection page
- Read about Using Phosphoserine to Study Protein Phosphorylation
- Read about GCE4All: Making Genetic Code Expansion Accessible


