We narrowed to 2,974 results for: rho
-
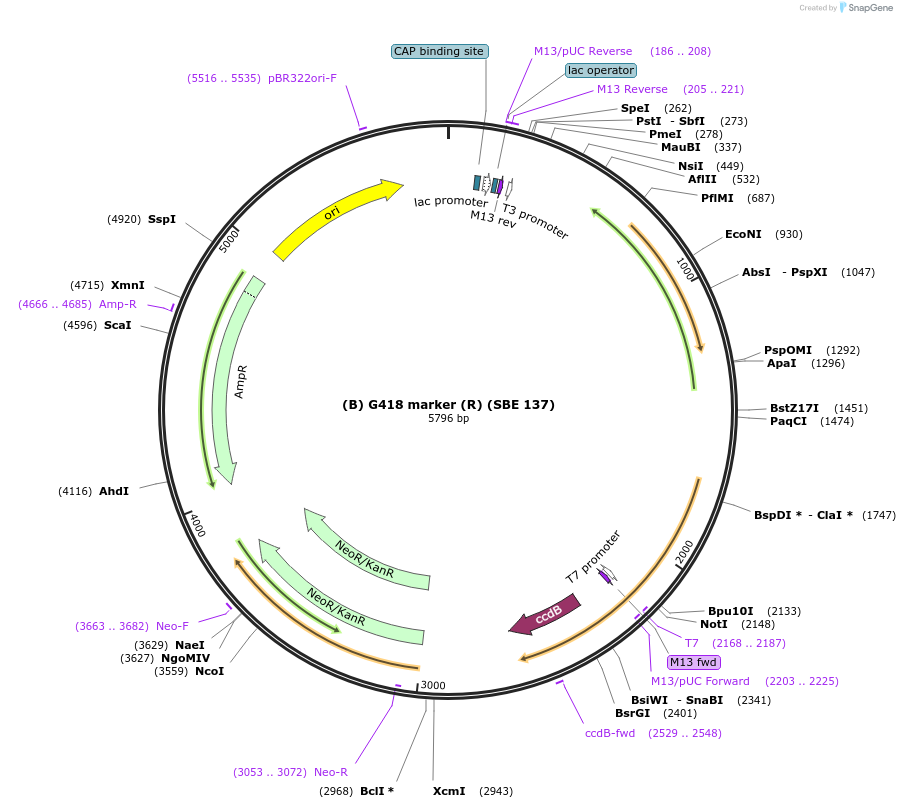
Plasmid#195011Purposegeneticin (G418) resistance genes codon-optimized for R. toruloides and expressed under pXYL and tNOSDepositorInsertpXYL+G418+tNOS
ExpressionBacterialPromoterpXYLAvailable SinceNov. 21, 2023AvailabilityAcademic Institutions and Nonprofits only -
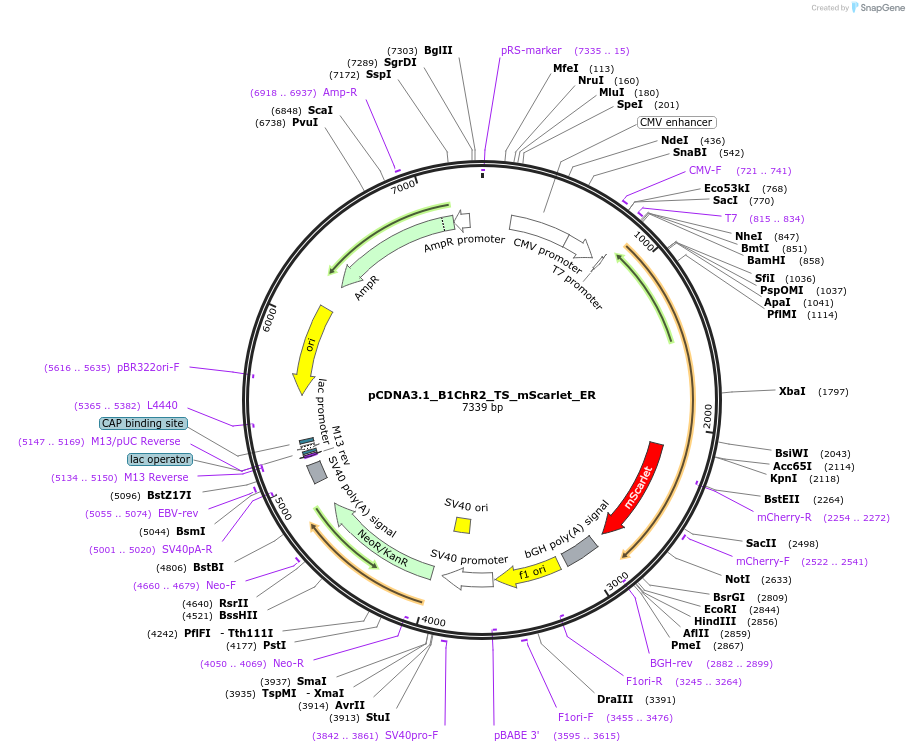
pCDNA3.1_B1ChR2_TS_mScarlet_ER
Plasmid#195191PurposeHighly K+ selective channelrhodopsinDepositorInsertBilabrum sp. K+ selective channelrhodopsin 2
TagsER-export sequence, Kir2.1 trafficking motif, and…ExpressionMammalianPromoterCMVAvailable SinceApril 14, 2023AvailabilityAcademic Institutions and Nonprofits only -
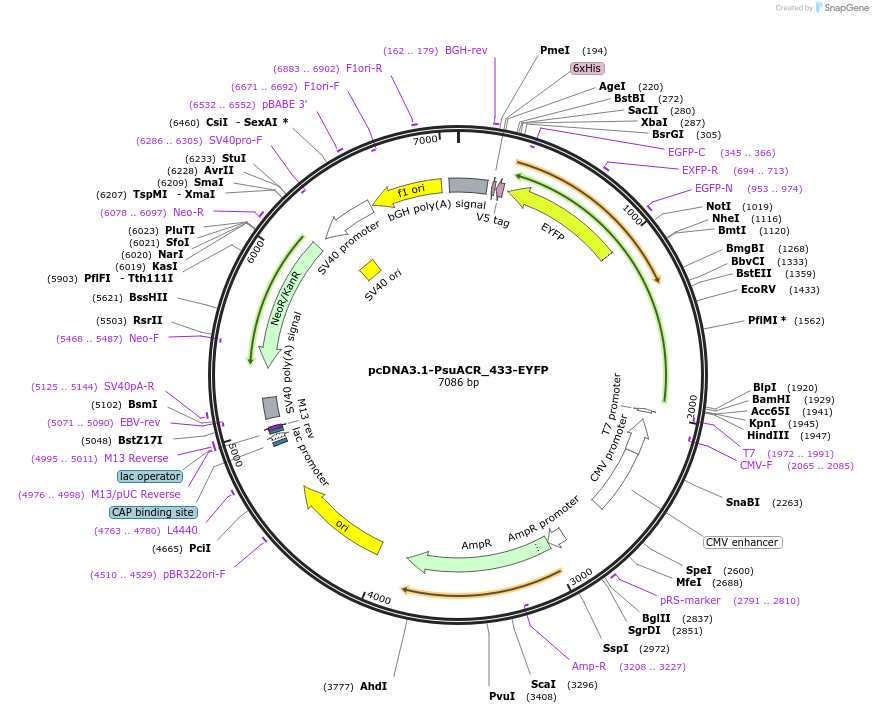
pcDNA3.1-PsuACR_433-EYFP
Plasmid#103529PurposeAnion channelrhodopsin from Proteomonas sulcata (CCMP704) for expression in mammalian cellsDepositorInsertAnion channelrhodopsin PsuACR_433
TagsEYFPExpressionMammalianMutationhuman codon optimizedPromoterCMVAvailable SinceNov. 29, 2017AvailabilityAcademic Institutions and Nonprofits only -
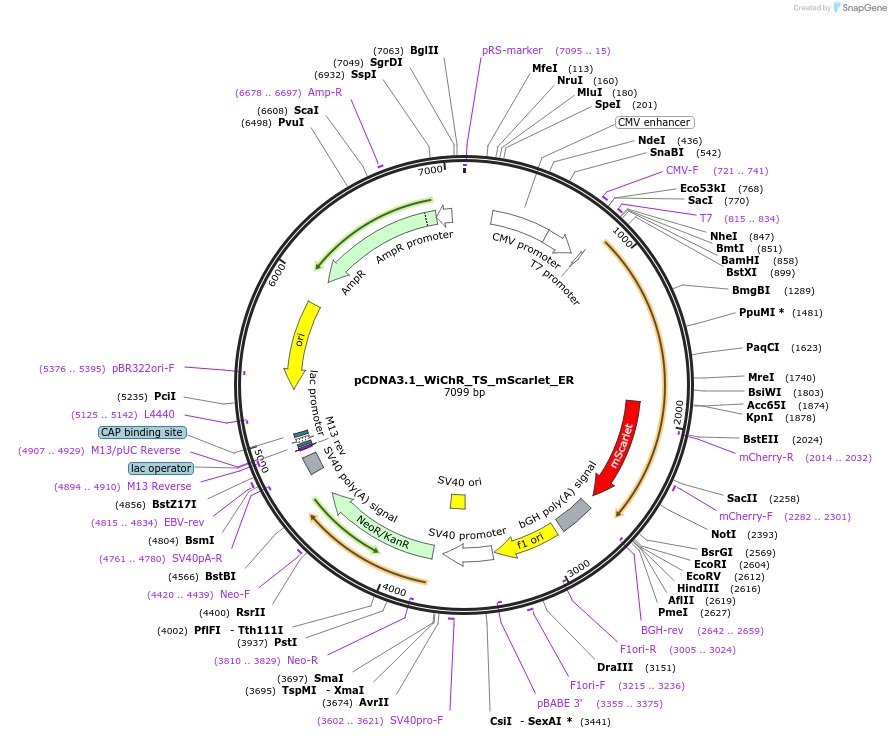
pCDNA3.1_WiChR_TS_mScarlet_ER
Plasmid#195190PurposeHigh efficient, K+ selective channelrhodopsin for the inhibition of excitable cellsDepositorInsertWobblia lunata inhibitory Channelrhodopsin
TagsER-export sequence, Kir2.1 trafficking motif, and…ExpressionMammalianPromoterCMVAvailable SinceApril 18, 2023AvailabilityAcademic Institutions and Nonprofits only -
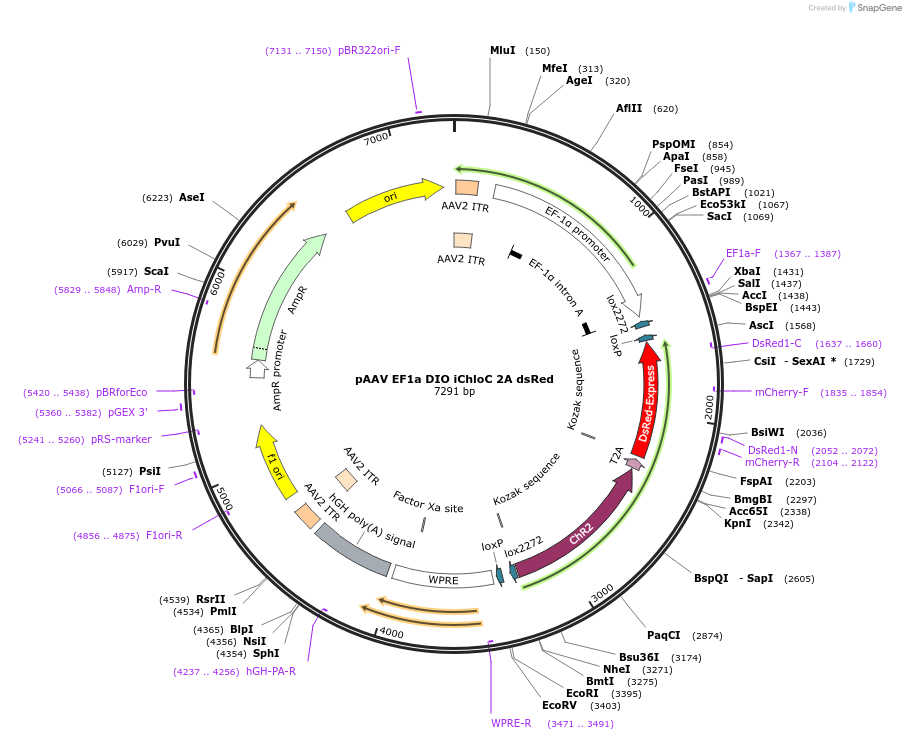
pAAV EF1a DIO iChloC 2A dsRed
Plasmid#70762PurposeAn improved chloride-conducting channelrhodopsin for light-induced inhibition of neuronal activity in vivo. Cre inducible expression (double floxed inversed ORF)DepositorInsertsChannelrhodopsin-2
red fluorescent protein
UseAAVExpressionMammalianMutationE83Q,E90R,E101S,D156N,T159CPromoterEF1aAvailable SinceJan. 7, 2016AvailabilityAcademic Institutions and Nonprofits only -
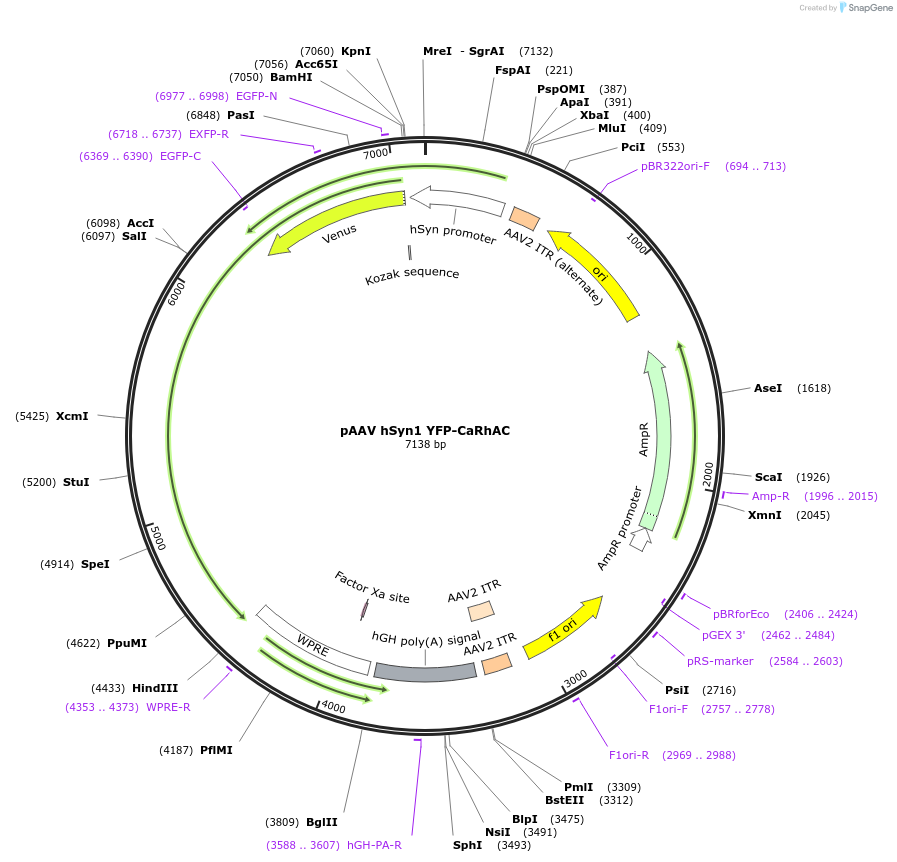
pAAV hSyn1 YFP-CaRhAC
Plasmid#101721PurposeYellow fluorescent protein (YFP) – tagged rhodopsin adenylyl cyclase created from the rhodopsin guanylyl cyclase of Catenaria anguillulae with a neuron-specific promoter. Useful for raising intracelluDepositorInsertYFP CaRhAC
UseAAVTagsYFPExpressionMammalianPromoterhSyn1Available SinceJan. 2, 2018AvailabilityAcademic Institutions and Nonprofits only -
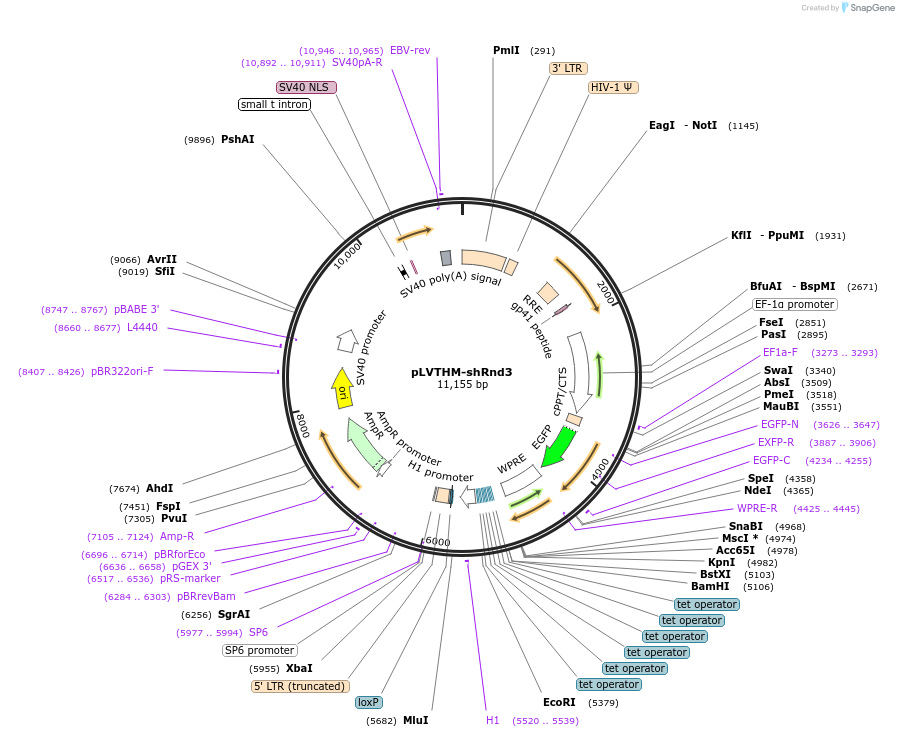
pLVTHM-shRnd3
Plasmid#86437Purpose2nd generation doxycycline-inducible lentiviral vector expressing Rnd3-targeting shRNA and GFPDepositorAvailable SinceFeb. 1, 2017AvailabilityAcademic Institutions and Nonprofits only -
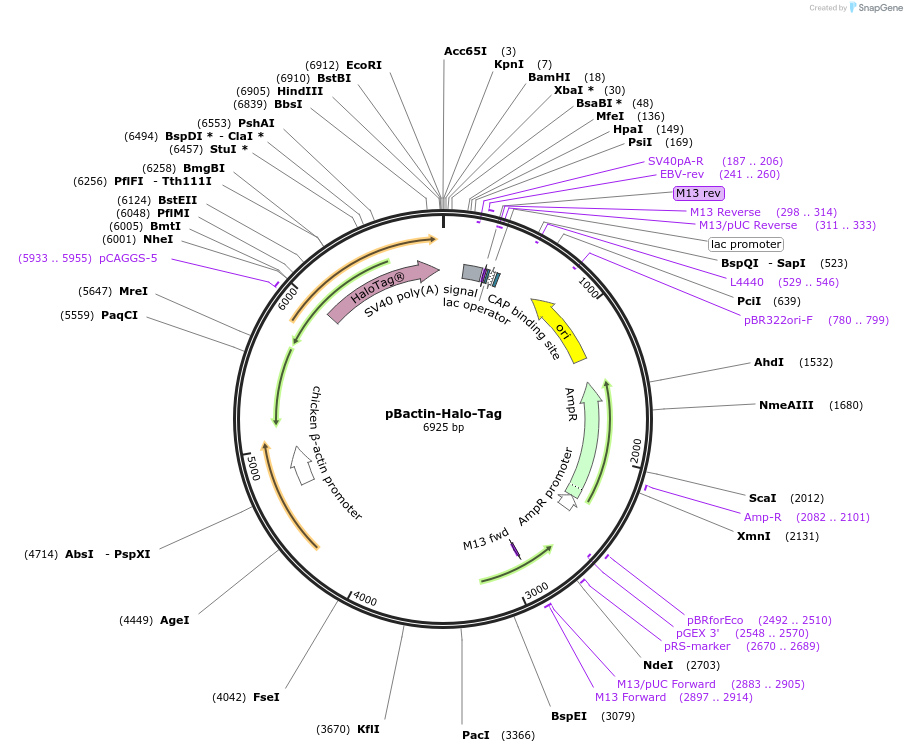
pBactin-Halo-Tag
Plasmid#196898PurposeExpression of Halo-Tag. Use as control for Halo-tag fusionsDepositorInsertHalo-Tag
ExpressionMammalianPromoterChicken bactin (plus Chicken bactin intron)Available SinceOct. 19, 2023AvailabilityAcademic Institutions and Nonprofits only -
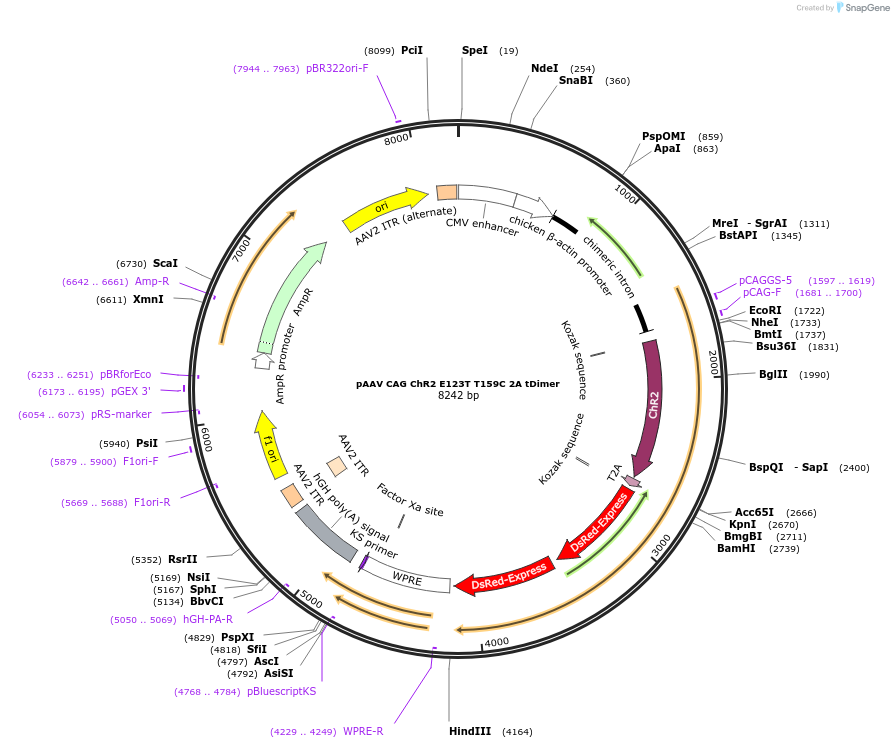
pAAV CAG ChR2 E123T T159C 2A tDimer
Plasmid#85399PurposeHigh efficiency channelrhodopsin with cytomegalovirus enhancer fused to chicken β-actin (CAG) promoterDepositorInsertsChannelrhodopsin-2
red fluorescent protein
UseAAVExpressionMammalianMutationE123T , T159CPromoterCAG and ribosomal skip sequence 2AAvailable SinceJan. 2, 2018AvailabilityAcademic Institutions and Nonprofits only -
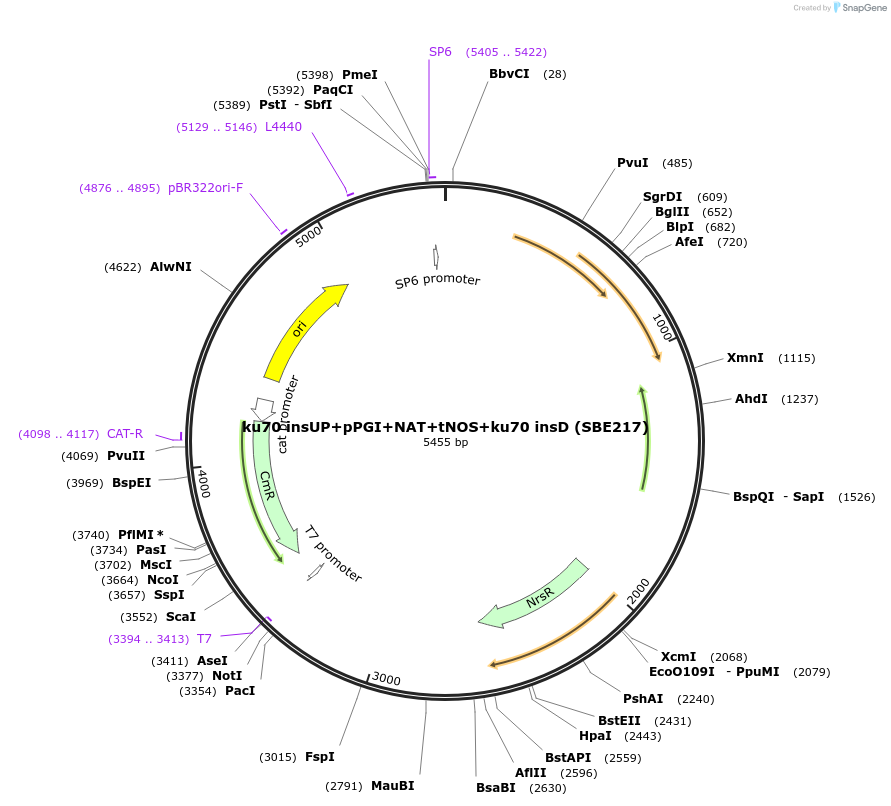
ku70 insUP+pPGI+NAT+tNOS+ku70 insD (SBE217)
Plasmid#195049Purposeintegration cassette for ku70 deletion and expression of NAT gene under pPGI for characterizationDepositorInsertNAT
ExpressionBacterialPromoterpPGIAvailable SinceNov. 21, 2023AvailabilityAcademic Institutions and Nonprofits only -
SL537
Plasmid#49945PurposeExpresses a TALE-TET1CD (catalytic domain of TET1) fusion protein engineered to bind a site in the human RHOXF2/2B homeobox (RHOXF2) gene and demethylate CpGs adjacent to the binding siteDepositorAvailable SinceDec. 20, 2013AvailabilityAcademic Institutions and Nonprofits only -
SL534
Plasmid#49940PurposeExpresses a TALE-TET1CD (catalytic domain of TET1) fusion protein engineered to bind a site in the human RHOXF2/2B homeobox (RHOXF2) gene and demethylate CpGs adjacent to the binding siteDepositorAvailable SinceDec. 10, 2013AvailabilityAcademic Institutions and Nonprofits only -
SL535
Plasmid#49941PurposeExpresses a TALE-TET1CD (catalytic domain of TET1) fusion protein engineered to bind a site in the human RHOXF2/2B homeobox (RHOXF2) gene and demethylate CpGs adjacent to the binding siteDepositorAvailable SinceDec. 10, 2013AvailabilityAcademic Institutions and Nonprofits only -
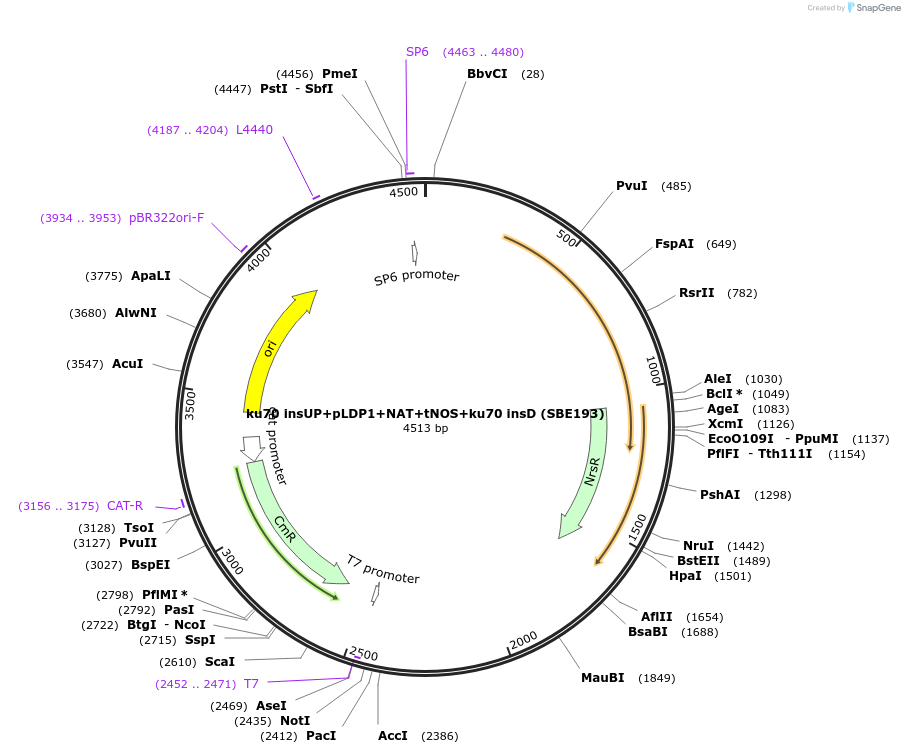
ku70 insUP+pLDP1+NAT+tNOS+ku70 insD (SBE193)
Plasmid#195054Purposeintegration cassette for ku70 deletion and expression of NAT gene under pLDP1 for characterizationDepositorInsertNAT
ExpressionBacterialPromoterpLDP1Available SinceNov. 21, 2023AvailabilityAcademic Institutions and Nonprofits only -
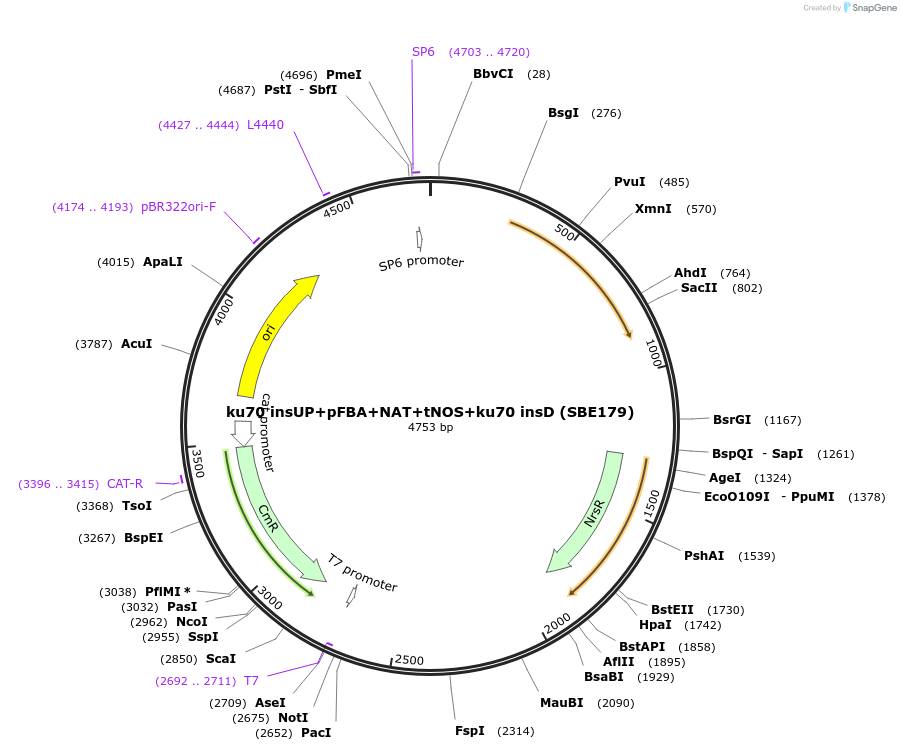
ku70 insUP+pFBA+NAT+tNOS+ku70 insD (SBE179)
Plasmid#195053Purposeintegration cassette for ku70 deletion and expression of NAT gene under pFBA for characterizationDepositorInsertNAT
ExpressionBacterialPromoterpFBAAvailable SinceNov. 21, 2023AvailabilityAcademic Institutions and Nonprofits only -
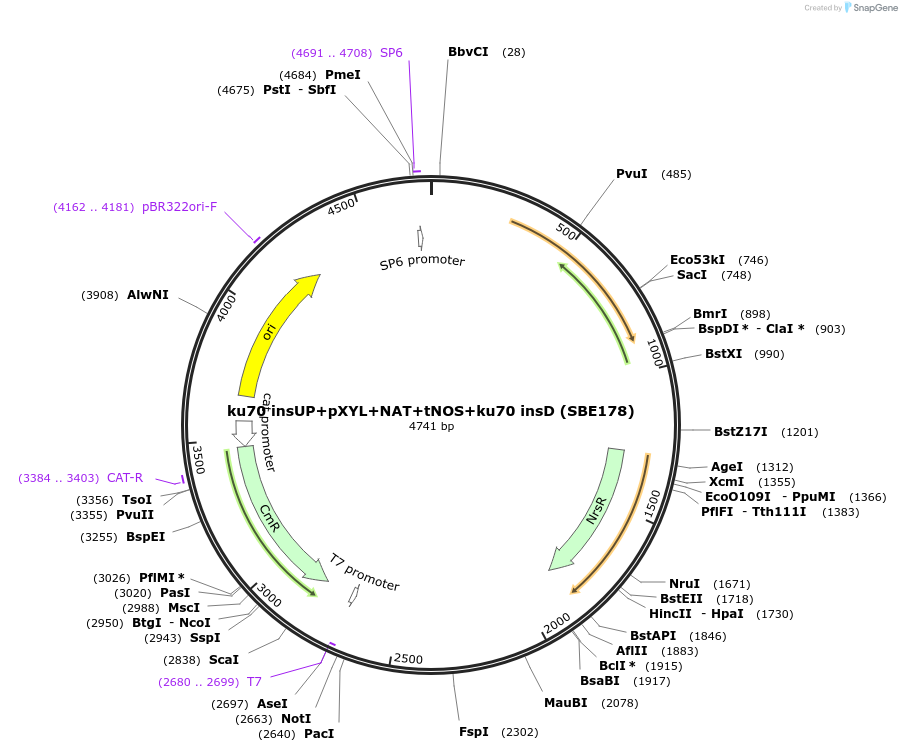
ku70 insUP+pXYL+NAT+tNOS+ku70 insD (SBE178)
Plasmid#195052Purposeintegration cassette for ku70 deletion and expression of NAT gene under pXYL for characterizationDepositorInsertNAT
ExpressionBacterialPromoterpXYLAvailable SinceNov. 21, 2023AvailabilityAcademic Institutions and Nonprofits only -
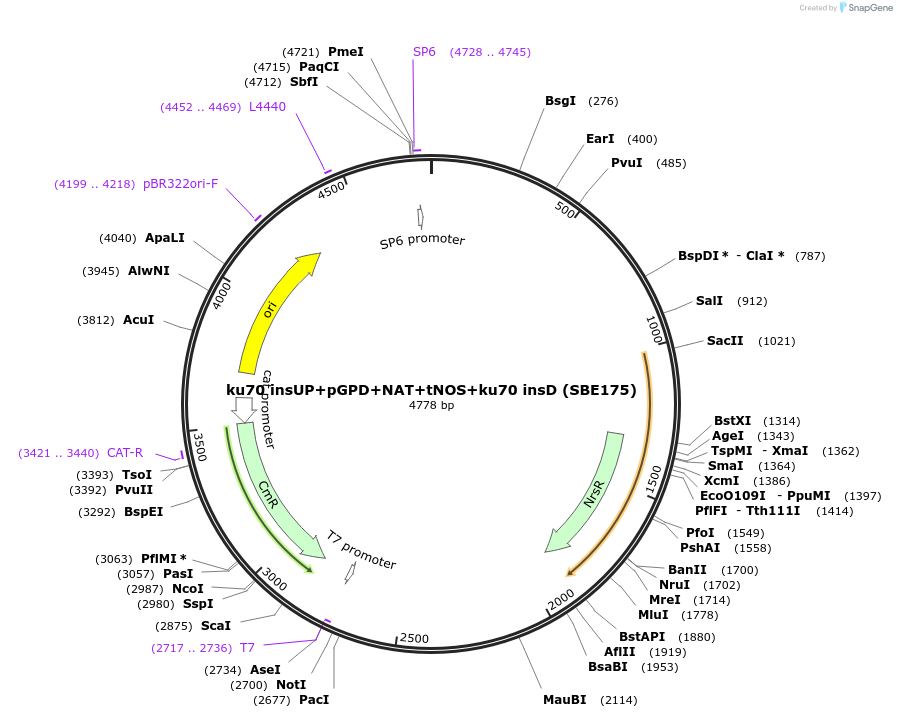
ku70 insUP+pGPD+NAT+tNOS+ku70 insD (SBE175)
Plasmid#195051Purposeintegration cassette for ku70 deletion and expression of NAT gene under pGPD for characterizationDepositorInsertNAT
ExpressionBacterialPromoterpGPDAvailable SinceNov. 21, 2023AvailabilityAcademic Institutions and Nonprofits only -
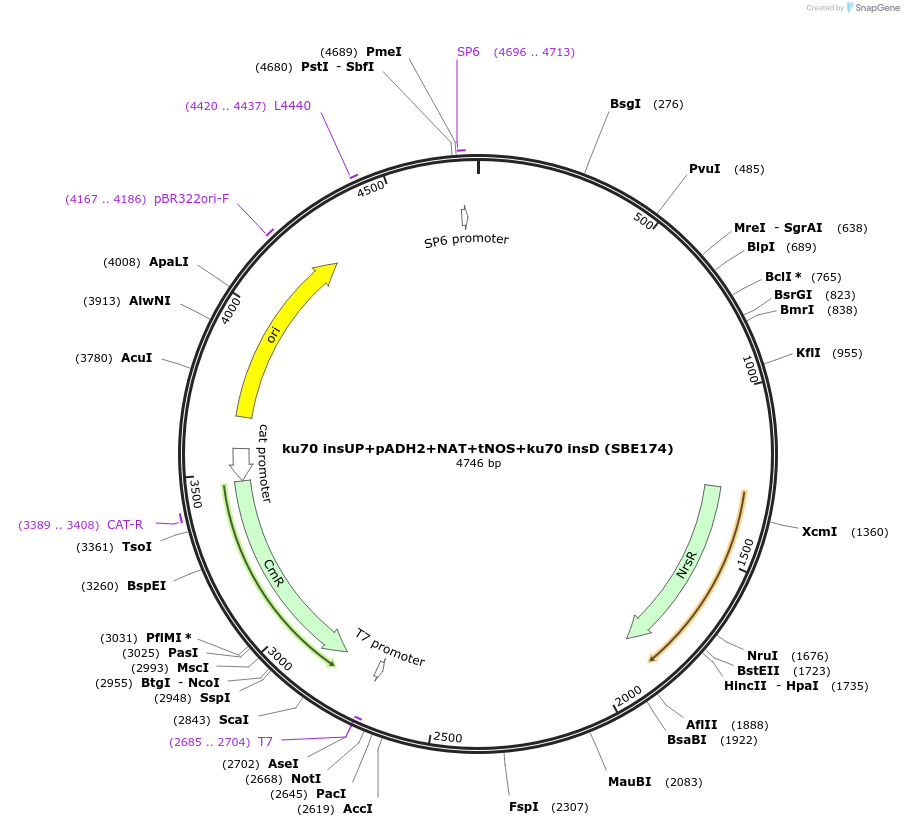
ku70 insUP+pADH2+NAT+tNOS+ku70 insD (SBE174)
Plasmid#195050Purposeintegration cassette for ku70 deletion and expression of NAT gene under pADH2 for characterizationDepositorInsertNAT
ExpressionBacterialPromoterpADH2Available SinceNov. 21, 2023AvailabilityAcademic Institutions and Nonprofits only -
SL542
Plasmid#49951PurposeExpresses a TALE-TET1CD (catalytic domain of TET1) fusion protein engineered to bind a site in the human RHOXF2/2B homeobox (RHOXF2) gene and demethylate CpGs adjacent to the binding siteDepositorAvailable SinceJan. 22, 2014AvailabilityAcademic Institutions and Nonprofits only -
SL544
Plasmid#49970PurposeExpresses a TALE-TET1CD (catalytic domain of TET1) fusion protein engineered to bind a site in the human RHOXF2/2B homeobox (RHOXF2) gene and demethylate CpGs adjacent to the binding siteDepositorAvailable SinceDec. 10, 2013AvailabilityAcademic Institutions and Nonprofits only -
SL545
Plasmid#49953PurposeExpresses a TALE-TET1CD (catalytic domain of TET1) fusion protein engineered to bind a site in the human RHOXF2/2B homeobox (RHOXF2) gene and demethylate CpGs adjacent to the binding siteDepositorAvailable SinceDec. 10, 2013AvailabilityAcademic Institutions and Nonprofits only -
SL543
Plasmid#49952PurposeExpresses a TALE-TET1CD (catalytic domain of TET1) fusion protein engineered to bind a site in the human RHOXF2/2B homeobox (RHOXF2) gene and demethylate CpGs adjacent to the binding siteDepositorAvailable SinceDec. 10, 2013AvailabilityAcademic Institutions and Nonprofits only -
SL541
Plasmid#49950PurposeExpresses a TALE-TET1CD (catalytic domain of TET1) fusion protein engineered to bind a site in the human RHOXF2/2B homeobox (RHOXF2) gene and demethylate CpGs adjacent to the binding siteDepositorAvailable SinceDec. 10, 2013AvailabilityAcademic Institutions and Nonprofits only -
SL540
Plasmid#49949PurposeExpresses a TALE-TET1CD (catalytic domain of TET1) fusion protein engineered to bind a site in the human RHOXF2/2B homeobox (RHOXF2) gene and demethylate CpGs adjacent to the binding siteDepositorAvailable SinceDec. 10, 2013AvailabilityAcademic Institutions and Nonprofits only -
MLM3720
Plasmid#49944PurposeExpresses a mutant TALE-TET1CD (inactive catalytic domain of TET1) fusion protein engineered to bind a site in the human RHOXF2/2B homeobox (RHOXF2) gene for use as a controlDepositorInsert3x FLAG Tet1CDmut RH-3 (RHOXF2 Human)
UseTALENTags3x FLAGMutationcatalytic domain of TET1 inactive (H1671Y and D16…PromoterEF1aAvailable SinceDec. 20, 2013AvailabilityAcademic Institutions and Nonprofits only -
MLM3709
Plasmid#49943PurposeExpresses a 3x Flag TALE-TET1CD (catalytic domain of TET1) fusion protein engineered to bind a site in the human RHOXF2/2B homeobox (RHOXF2) gene and demethylate CpGs adjacent to the binding siteDepositorAvailable SinceDec. 20, 2013AvailabilityAcademic Institutions and Nonprofits only -
MLM3713
Plasmid#49946PurposeExpresses a 3x Flag TALE-TET1CD (catalytic domain of TET1) fusion protein engineered to bind a site in the human RHOXF2/2B homeobox (RHOXF2) gene and demethylate CpGs adjacent to the binding siteDepositorAvailable SinceDec. 31, 2013AvailabilityAcademic Institutions and Nonprofits only -
MLM3762
Plasmid#49947PurposeExpresses a mutant TALE-TET1CD (inactive catalytic domain of TET1) fusion protein engineered to bind a site in the human RHOXF2/2B homeobox (RHOXF2) gene for use as a controlDepositorInsert3x FLAG Tet1CDmut RH-4 (RHOXF2 Human)
UseTALENTags3x FLAGMutationcatalytic domain of TET1 inactive (H1671Y and D16…PromoterEF1aAvailable SinceDec. 31, 2013AvailabilityAcademic Institutions and Nonprofits only -
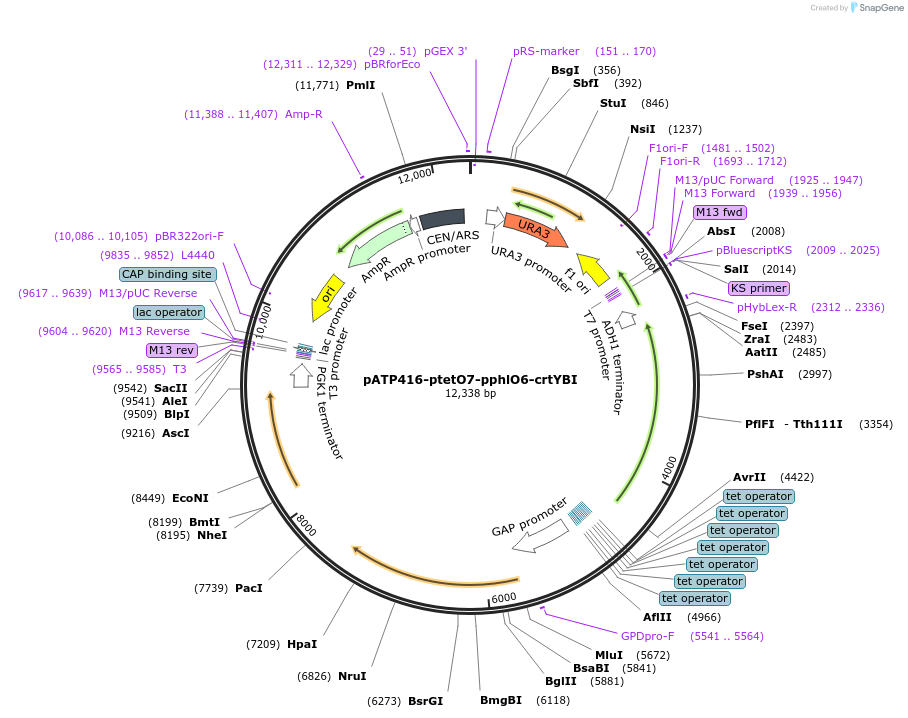
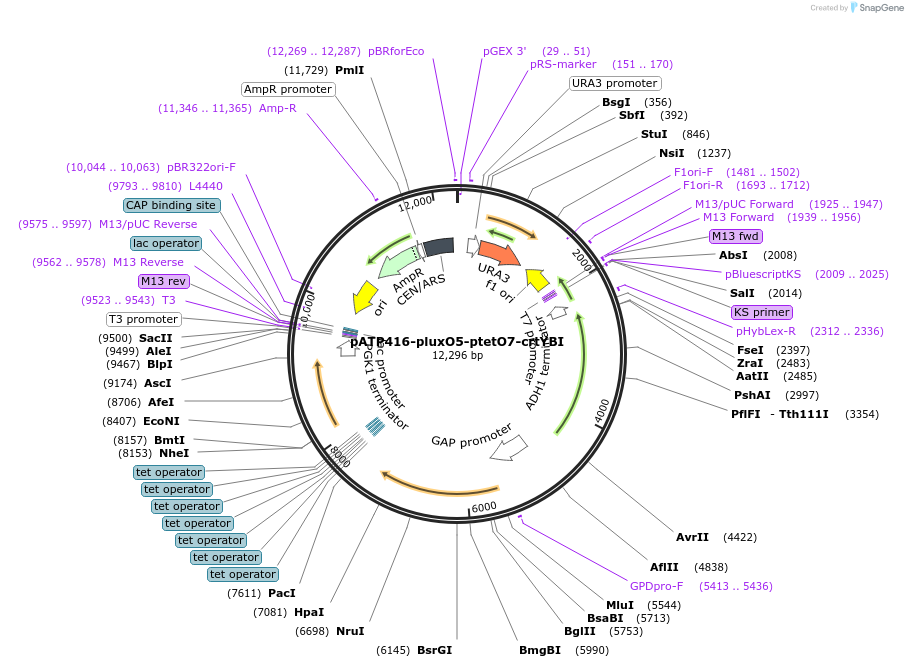
pATP416-ptetO7-pphlO6-crtYBI
Plasmid#165976PurposeExpresses carotenoid biosynthesis gene in response to 2,4-diacetylphloroglucinol and doxycycline in yeast expressing rtTA and PhlTADepositorInsertsXanthophyllomyces dendrorhous phytoene-beta carotene synthase (crtYB) mRNA, complete cds
Xanthophyllomyces dendrorhous phytoene desaturase mRNA, complete cds
BTS1 (BTS1 Budding Yeast)
UseSynthetic BiologyExpressionYeastMutationC1281A, G1698A and C66A, C1359TPromoterSynthetic promoter (pphlO6), Synthetic promoter (…Available SinceMarch 29, 2021AvailabilityAcademic Institutions and Nonprofits only -
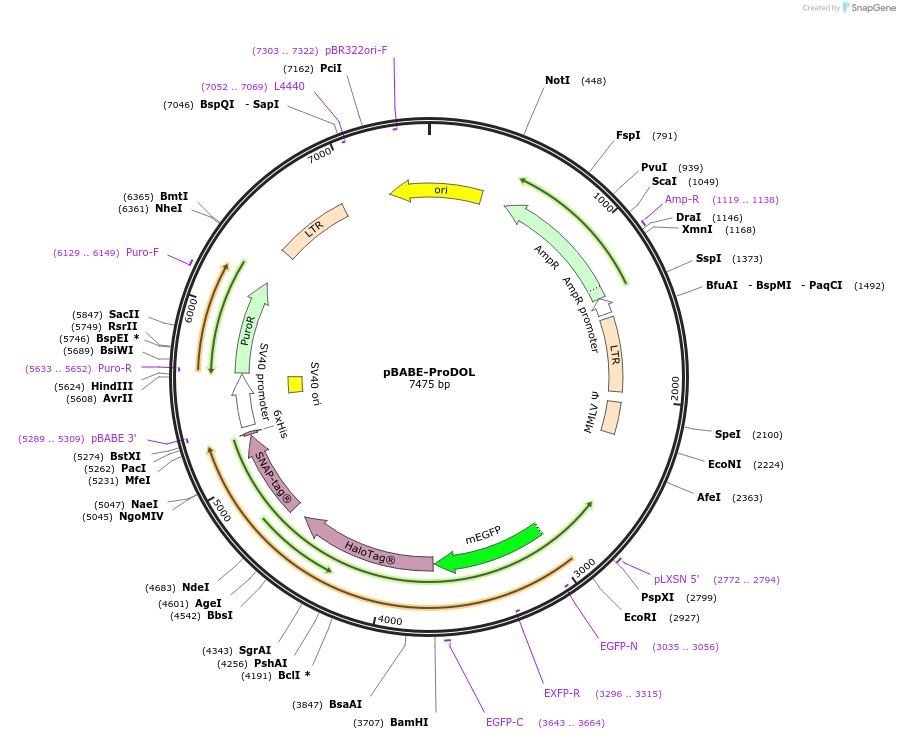
pBABE-ProDOL
Plasmid#206866PurposeExpression of ProDOL probe - LynAnchor-eGFP-SNAP-tag-HaloTagDepositorInsertProDOL
UseRetroviralTagsHaloTag, His-Tag, SNAP-tag, and eGFPExpressionMammalianMutationContains Lyn amino acids 1-16Available SinceAug. 8, 2024AvailabilityAcademic Institutions and Nonprofits only -
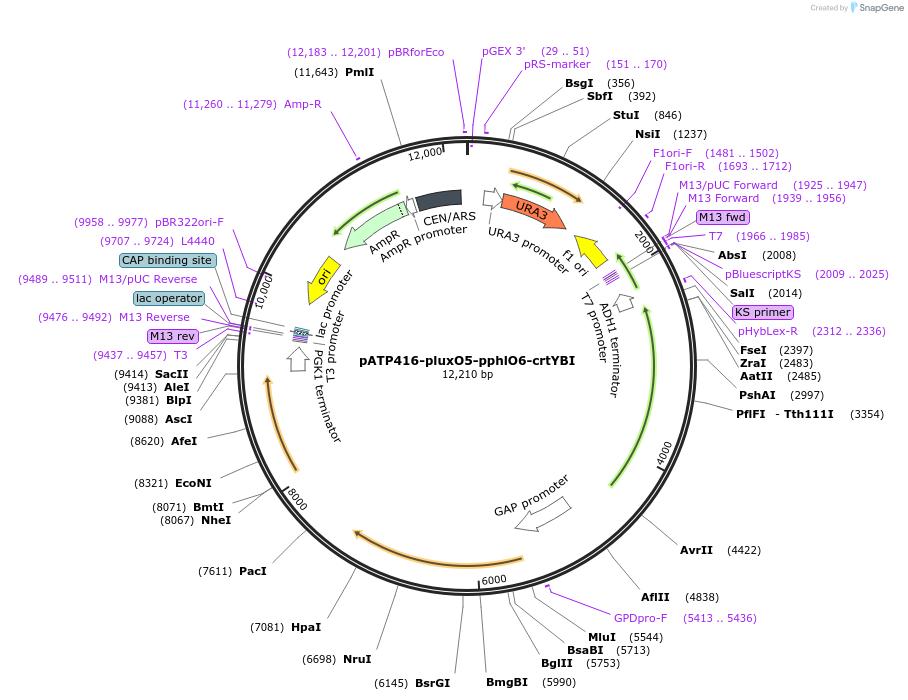
pATP416-pluxO5-pphlO6-crtYBI
Plasmid#165977PurposeExpresses carotenoid biosynthesis gene in response to homoserine lactone and 2,4-diacetylphloroglucinol in yeast expressing PhlTA and LuxTADepositorInsertsXanthophyllomyces dendrorhous phytoene-beta carotene synthase (crtYB) mRNA, complete cds
Xanthophyllomyces dendrorhous phytoene desaturase mRNA, complete cds
BTS1 (BTS1 Budding Yeast)
UseSynthetic BiologyExpressionYeastMutationC1281A, G1698A and C66A, C1359TPromoterSynthetic promoter (pluxO5), Synthetic promoter (…Available SinceMarch 29, 2021AvailabilityAcademic Institutions and Nonprofits only -
pATP416-pluxO5-ptetO7-crtYBI
Plasmid#165975PurposeExpresses carotenoid biosynthesis gene in response to homoserine lactone and doxycycline in yeast expressing LuxTA and rtTADepositorInsertsXanthophyllomyces dendrorhous phytoene-beta carotene synthase (crtYB) mRNA, complete cds
Xanthophyllomyces dendrorhous phytoene desaturase mRNA, complete cds
BTS1 (BTS1 Budding Yeast)
UseSynthetic BiologyExpressionYeastMutationC1281A, G1698A and C66A, C1359TPromoterSynthetic promoter (pluxO5), Synthetic promoter (…Available SinceMarch 29, 2021AvailabilityAcademic Institutions and Nonprofits only -
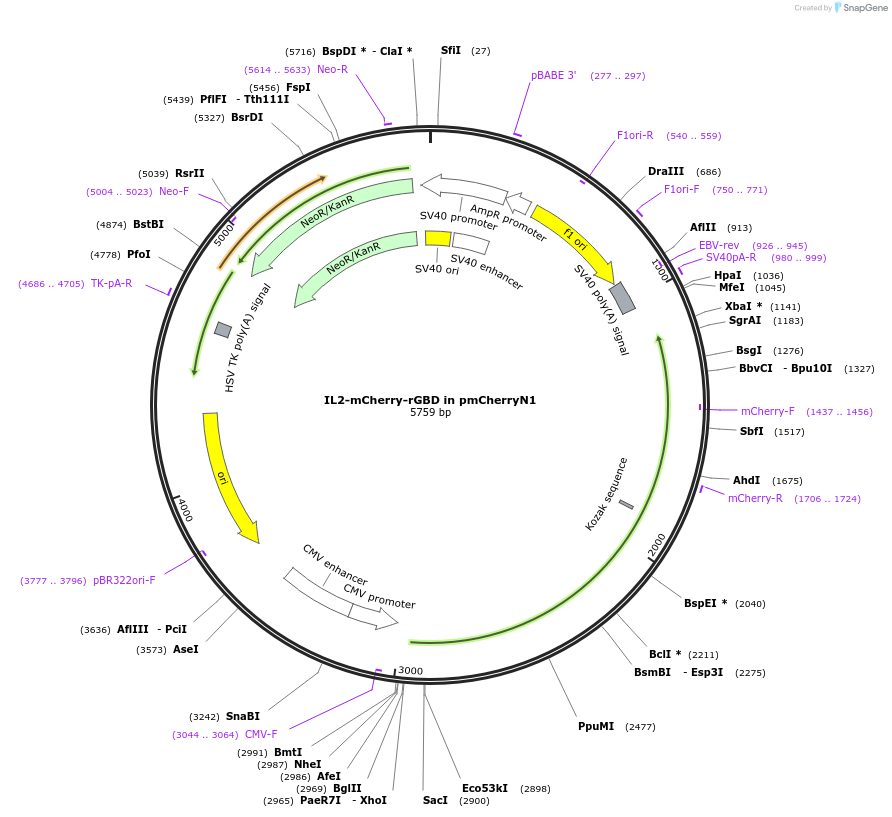
IL2-mCherry-rGBD in pmCherryN1
Plasmid#187286PurposeExpress IL2-Rhotekin GTPase-binding domain-mCherryDepositorTagsmCherryExpressionMammalianPromoterCMVAvailable SinceSept. 14, 2022AvailabilityAcademic Institutions and Nonprofits only -
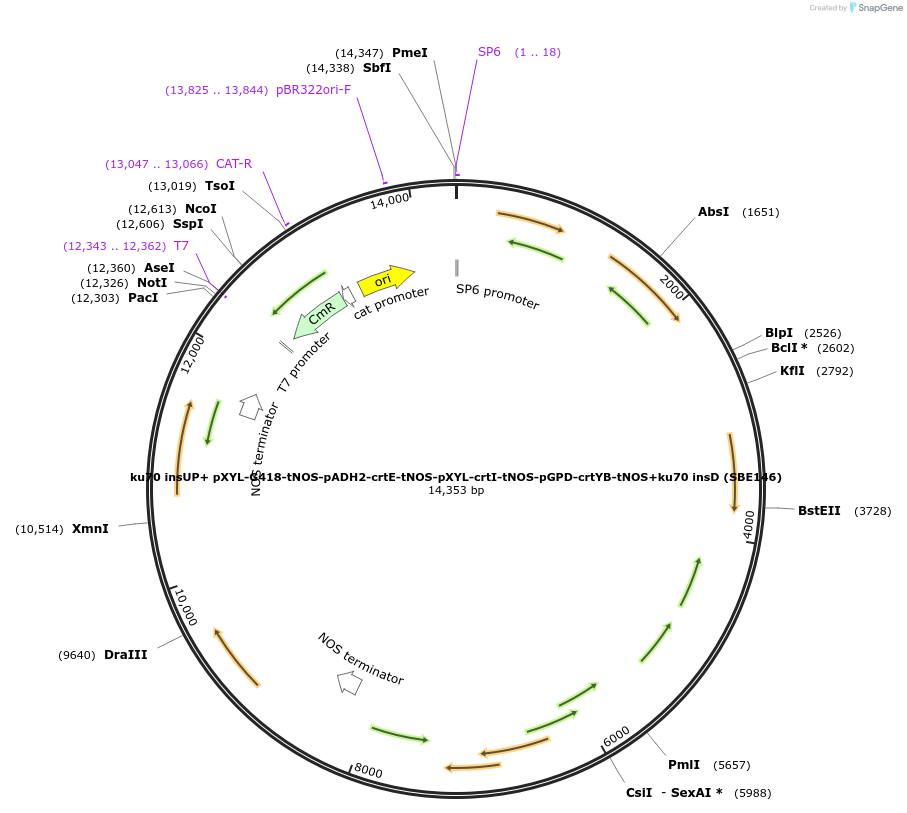
ku70 insUP+ pXYL-G418-tNOS-pADH2-crtE-tNOS-pXYL-crtI-tNOS-pGPD-crtYB-tNOS+ku70 insD (SBE146)
Plasmid#195048Purposecassette for the deletion of ku70 and overexpression of crtE, crtI, and crtYB under different combination of promotersDepositorInsertscrtE
crtI
crtYB
ExpressionYeastPromoterpADH2, pGPD, and pXYLAvailable SinceNov. 21, 2023AvailabilityAcademic Institutions and Nonprofits only -
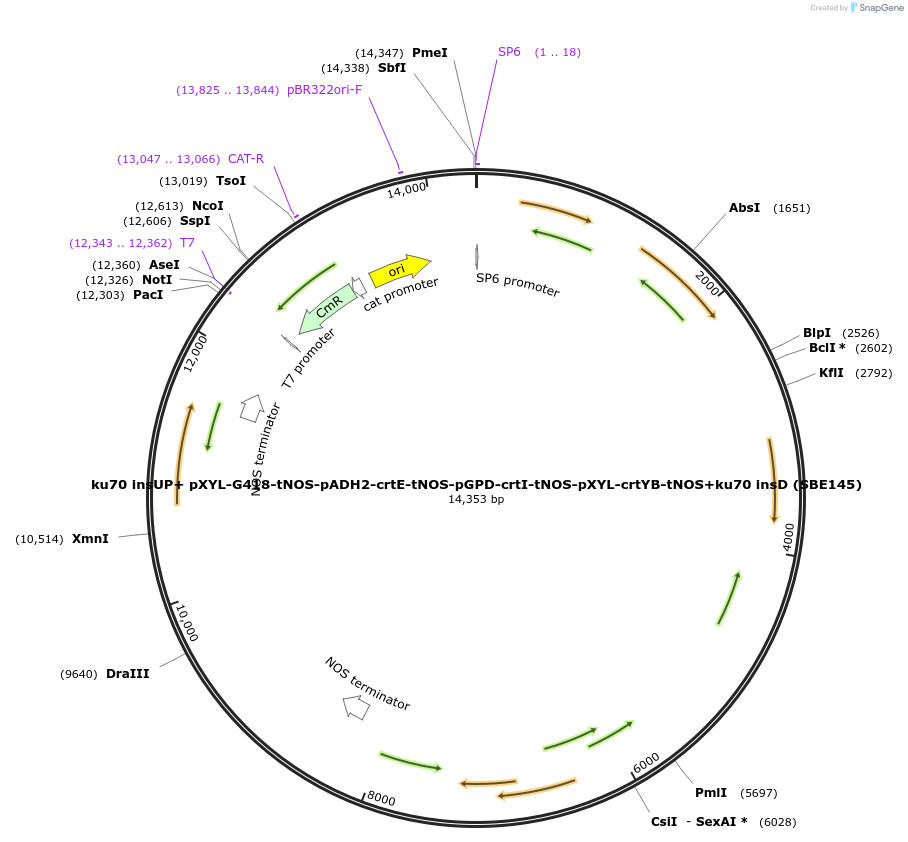
ku70 insUP+ pXYL-G418-tNOS-pADH2-crtE-tNOS-pGPD-crtI-tNOS-pXYL-crtYB-tNOS+ku70 insD (SBE145)
Plasmid#195047Purposecassette for the deletion of ku70 and overexpression of crtE, crtI, and crtYB under different combination of promotersDepositorInsertscrtE
crtI
crtYB
ExpressionYeastPromoterpADH2, pGPD, and pXYLAvailable SinceNov. 21, 2023AvailabilityAcademic Institutions and Nonprofits only -
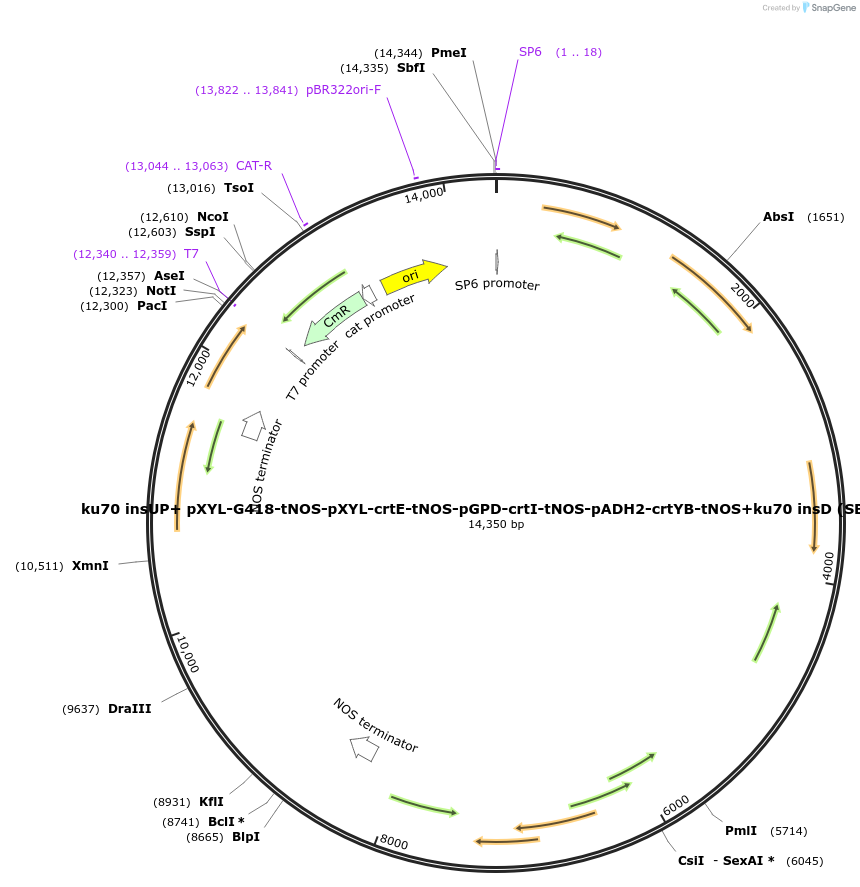
ku70 insUP+ pXYL-G418-tNOS-pXYL-crtE-tNOS-pGPD-crtI-tNOS-pADH2-crtYB-tNOS+ku70 insD (SBE144)
Plasmid#195046Purposecassette for the deletion of ku70 and overexpression of crtE, crtI, and crtYB under different combination of promotersDepositorInsertscrtE
crtI
crtYB
ExpressionYeastPromoterpADH2, pGPD, and pXYLAvailable SinceNov. 21, 2023AvailabilityAcademic Institutions and Nonprofits only -
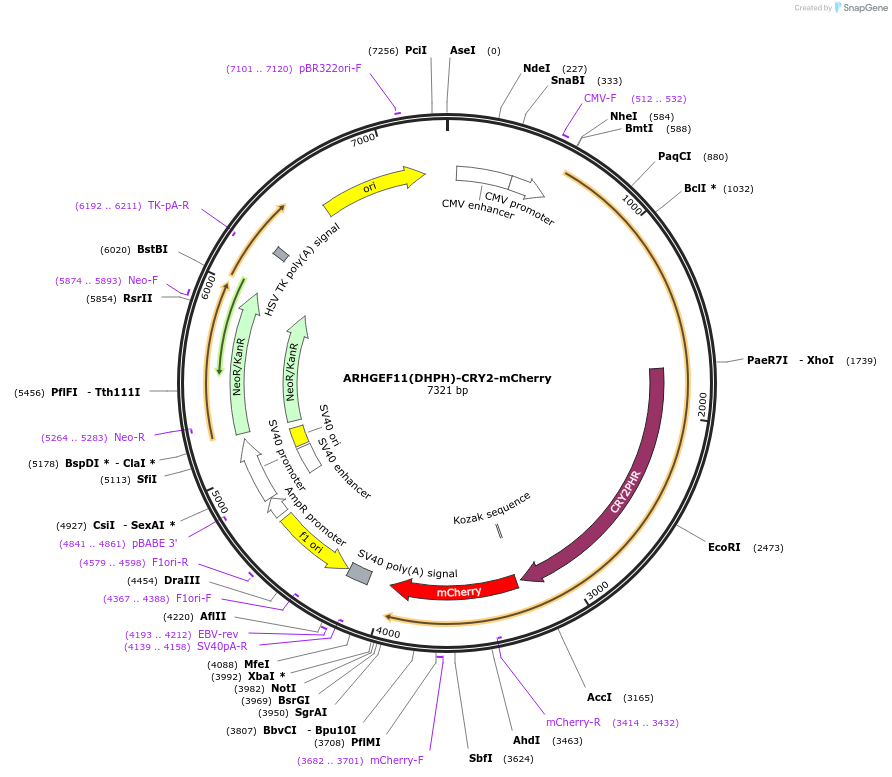
ARHGEF11(DHPH)-CRY2-mCherry
Plasmid#89481PurposeAlso known as optoGEF-RhoA, encodes for a fusion between CRY2(1-498)-mCherry and the DHPH domain of ARHGEF11DepositorTagsmCherryExpressionMammalianPromoterCMVAvailable SinceJan. 9, 2019AvailabilityAcademic Institutions and Nonprofits only -
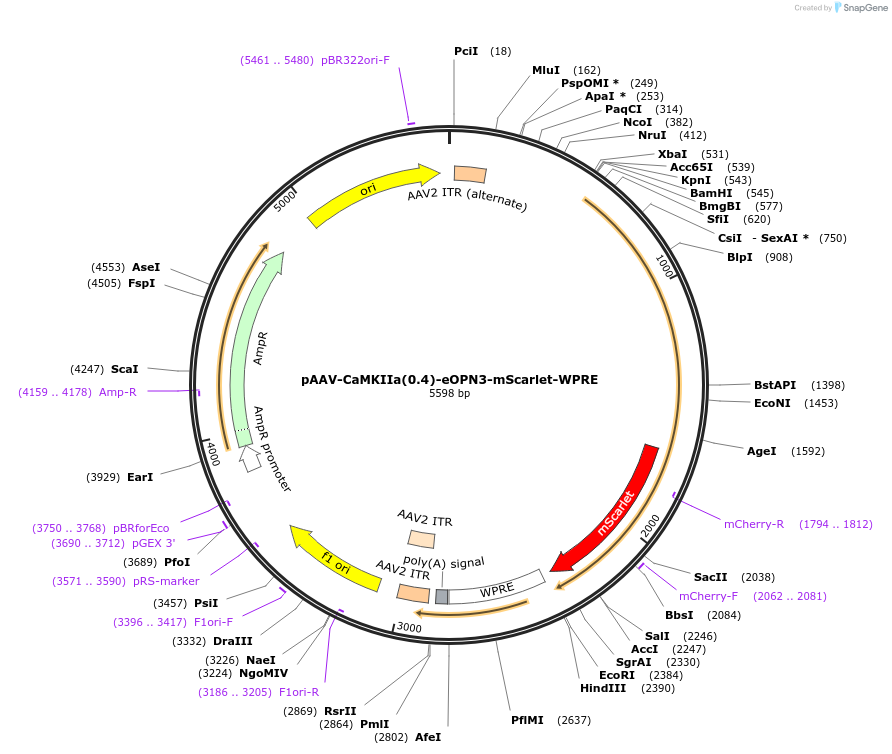
pAAV-CaMKIIa(0.4)-eOPN3-mScarlet-WPRE
Plasmid#125712PurposeExpresses an optimized mosquito OPN3 rhodopsin in-frame with mScarletDepositorHas ServiceAAV Retrograde, AAV1, and AAV5InserteOPN3
UseAAVTagsRho1D4 and mScarletExpressionMammalianMutationdeleted amino acids 331-429Promotershort version of αCamKinase II promoter (0.4kb)Available SinceFeb. 19, 2021AvailabilityAcademic Institutions and Nonprofits only -
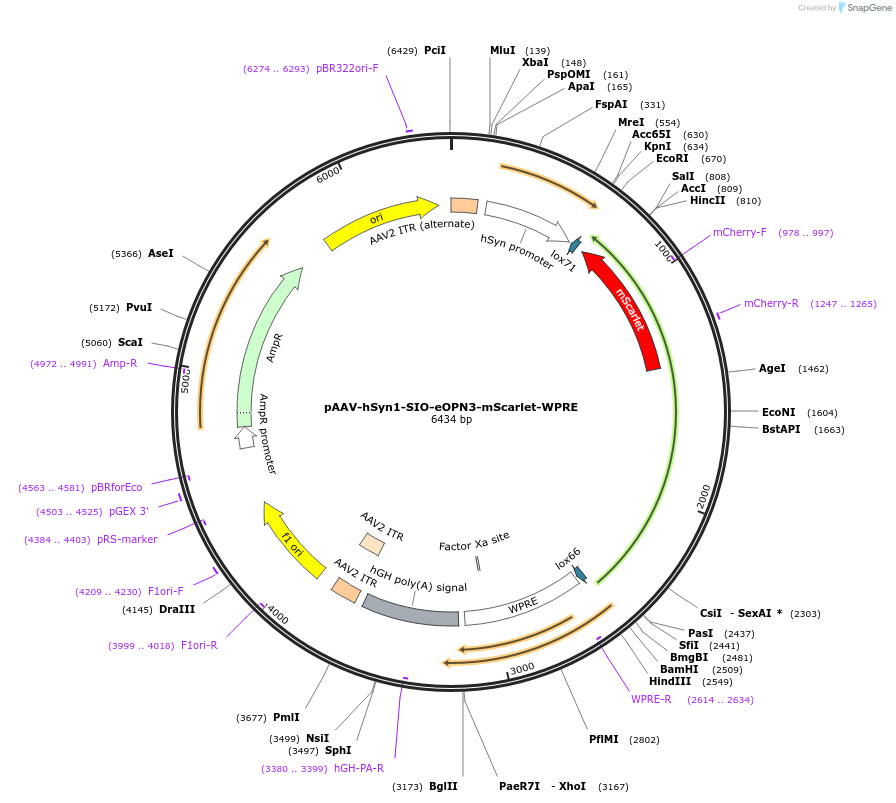
pAAV-hSyn1-SIO-eOPN3-mScarlet-WPRE
Plasmid#125713PurposeCre-dependent vector expressing an optimized mosquito OPN3 rhodopsin in-frame with mScarletDepositorHas ServiceAAV1 and AAV5InserteOPN3
UseAAV and Cre/LoxTagsRho1D4 and mScarletExpressionMammalianMutationdeleted amino acids 331-429PromoterhSyn1Available SinceSept. 20, 2019AvailabilityAcademic Institutions and Nonprofits only -
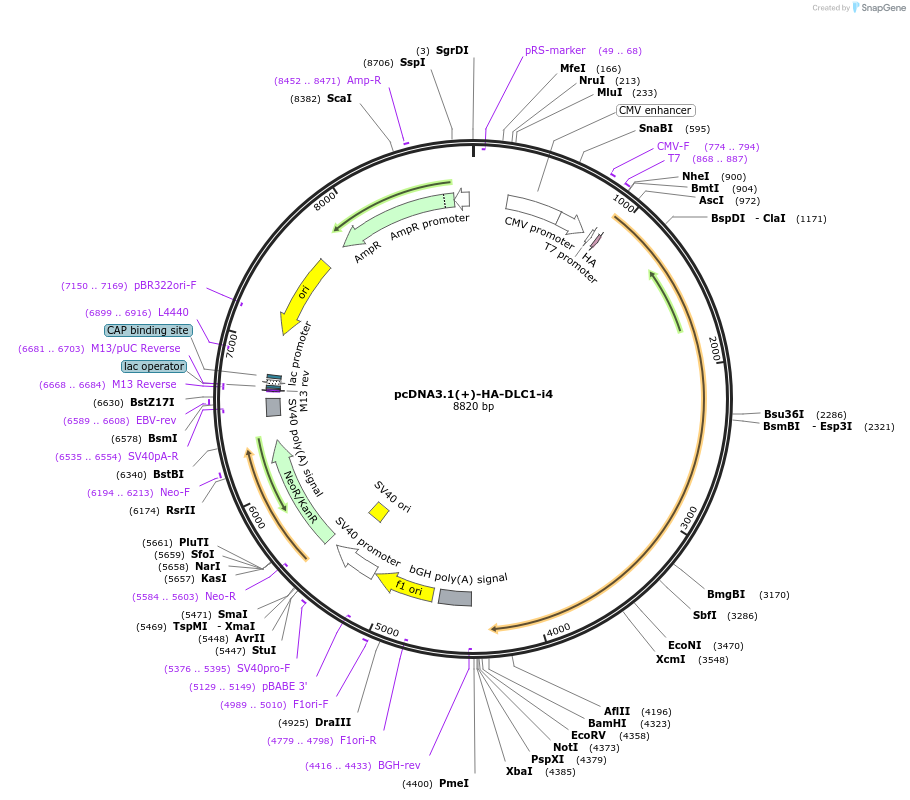
pcDNA3.1(+)-HA-DLC1-i4
Plasmid#99534PurposeDLC1-isoform 4 expression plasmidDepositorAvailable SinceNov. 14, 2017AvailabilityAcademic Institutions and Nonprofits only -
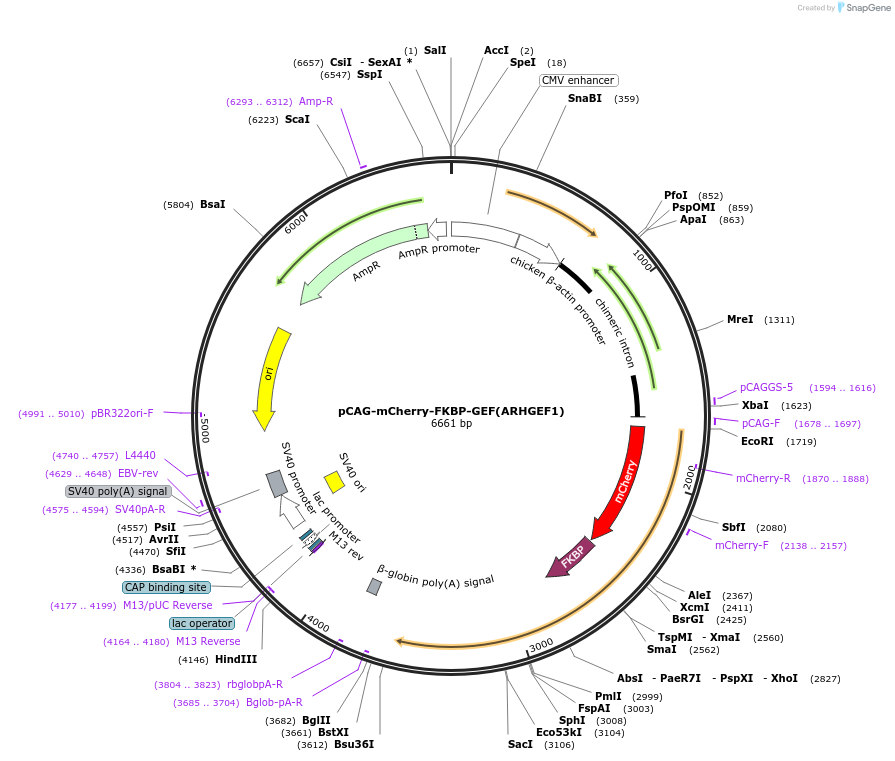
pCAG-mCherry-FKBP-GEF(ARHGEF1)
Plasmid#85152Purposeacute activation of Rho through rapamycin-induced translocation to membrane-localized FRBDepositorInsertFusion of mCherry, FKBP, and the DH-domain of ARHGEF1 (amino acids 380-630) (ARHGEF1 Human)
TagsmCherry-FKBPExpressionMammalianPromoterCAGAvailable SinceFeb. 13, 2017AvailabilityAcademic Institutions and Nonprofits only -
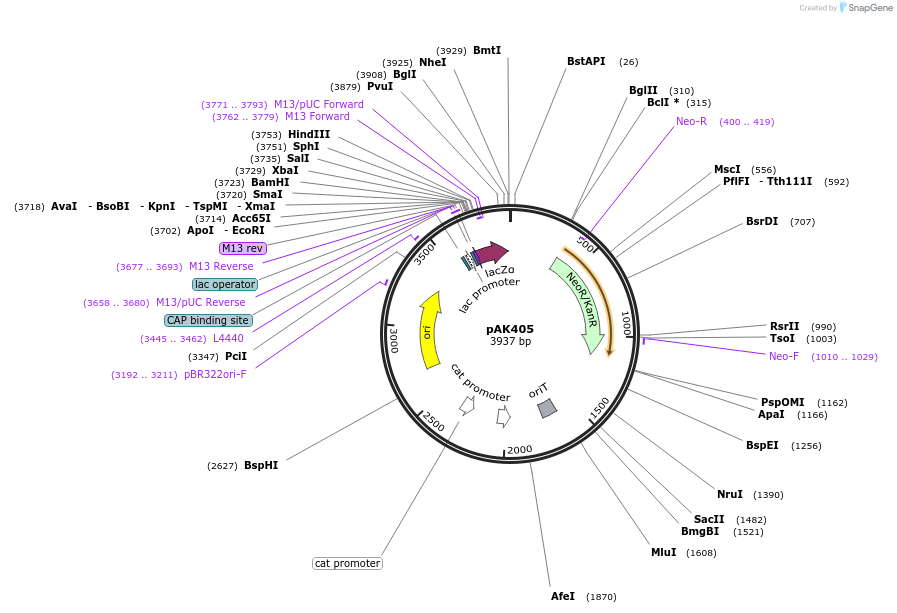
pAK405
Plasmid#37114DepositorTypeEmpty backboneExpressionBacterialAvailable SinceAug. 8, 2012AvailabilityAcademic Institutions and Nonprofits only -
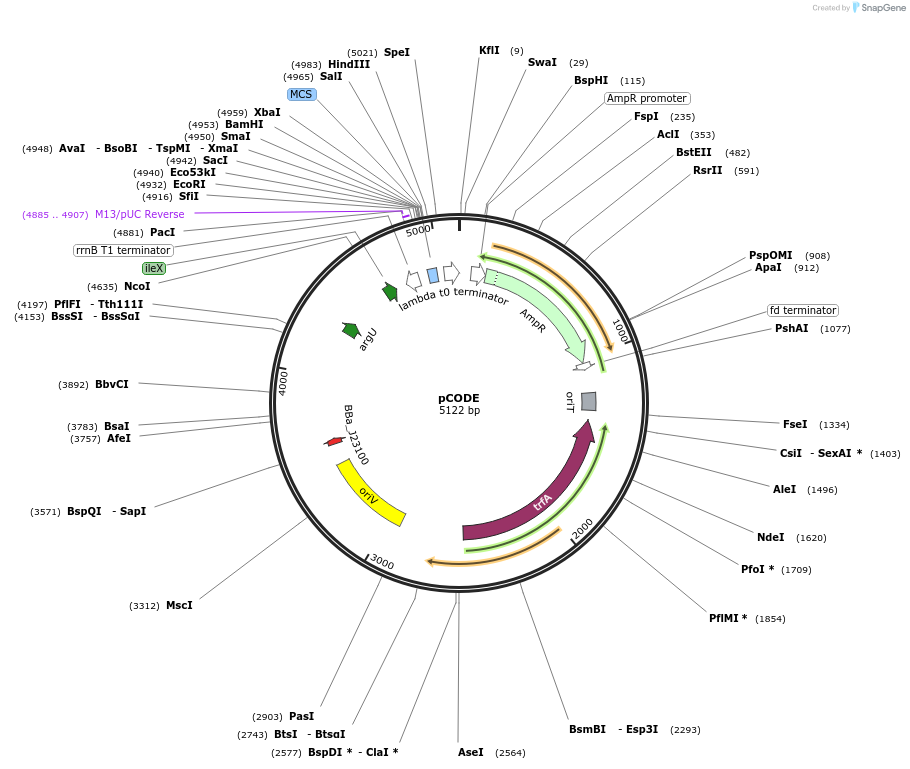
pCODE
Plasmid#247473PurposePlasmid for constitutive expression of six tRNA encoding genes (argX, glyT, leuW, proL, argU, and ileX) in Escherichia coli, based on pSEVA121 backboneDepositorInsertargX, glyT, leuW, proL, argU, and ileX
ExpressionBacterialMutationRearrangement of tRNA operonPromoterJ23100Available SinceFeb. 23, 2026AvailabilityAcademic Institutions and Nonprofits only -
pCODE-3
Plasmid#247475PurposePlasmid for inducible expression of six tRNA encoding genes (argX, glyT, leuW, proL, argU, and ileX) in Escherichia coli, based on pSEVA121 backboneDepositorInsertargX, glyT, leuW, proL, argU, and ileX
ExpressionBacterialMutationRearrangement of tRNA operon, tRNA flanking regio…PromoterpTetAvailable SinceNov. 4, 2025AvailabilityAcademic Institutions and Nonprofits only -
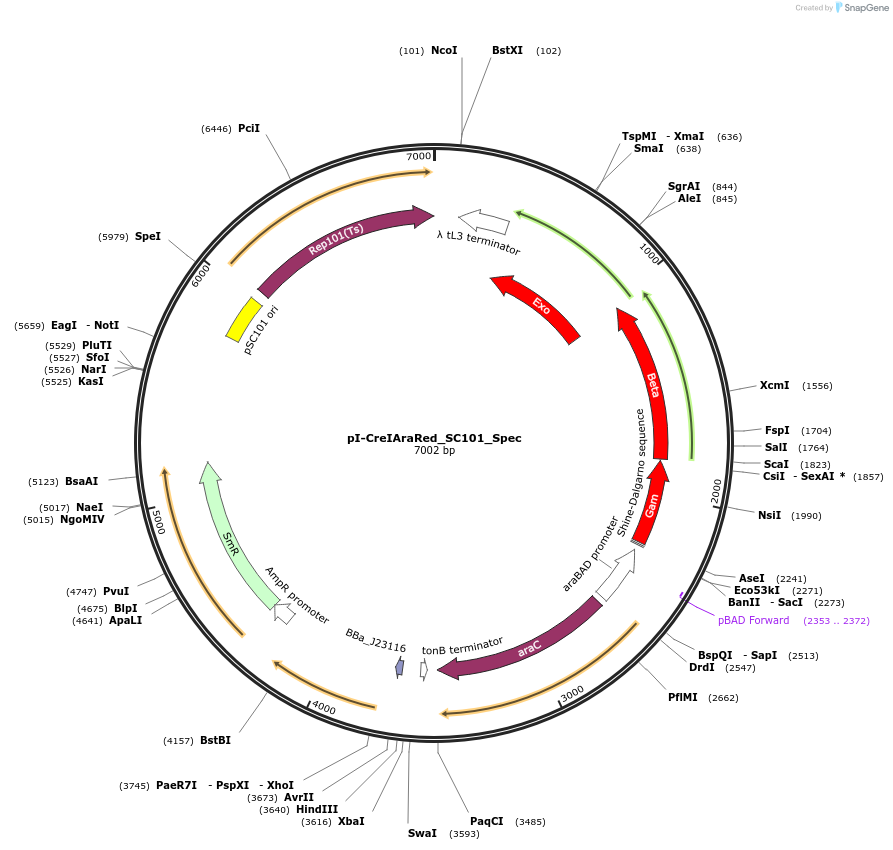
pI-CreIAraRed_SC101_Spec
Plasmid#240155PurposeExpresses lambda red machinery and i-creIDepositorInsertsgam
beta
exo
i-creI
ExpressionBacterialAvailable SinceOct. 22, 2025AvailabilityAcademic Institutions and Nonprofits only -
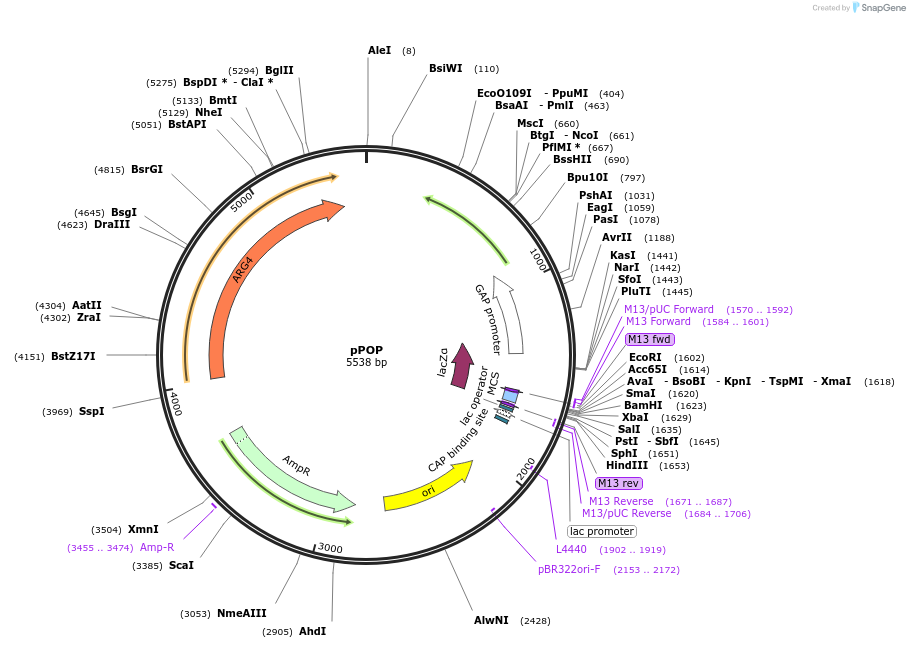
pPOP
Plasmid#59526PurposeVector for pop-in/pop-out gene replacement in Pichia pastoris.DepositorTypeEmpty backboneExpressionYeastPromoterGAPAvailable SinceOct. 1, 2014AvailabilityAcademic Institutions and Nonprofits only -
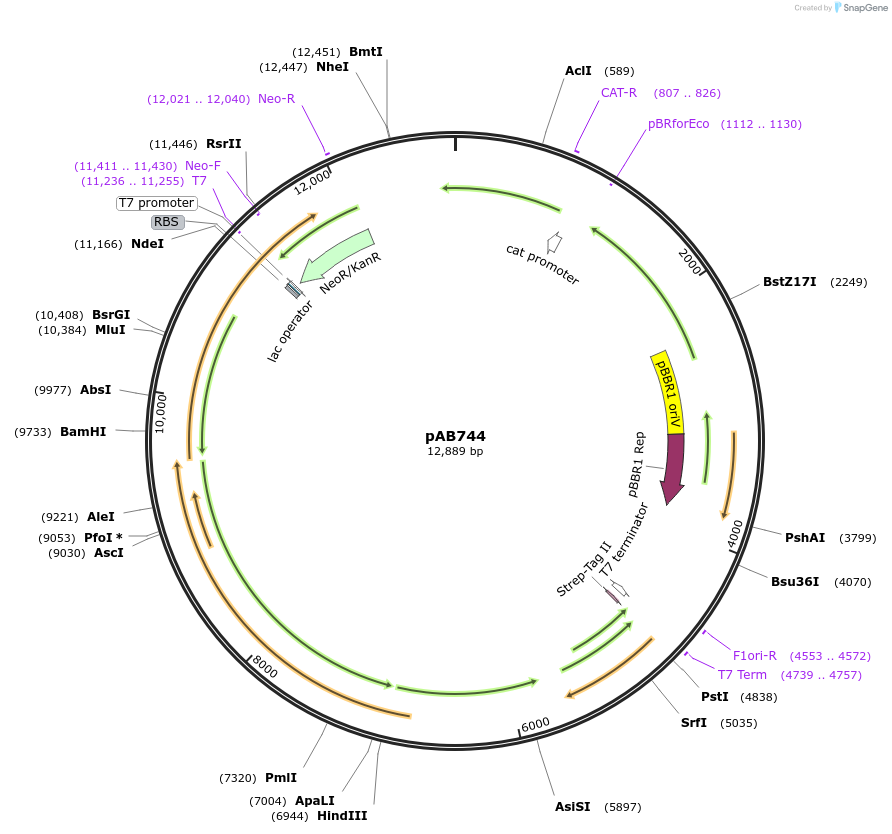
pAB744
Plasmid#176579PurposeExpressing the whole n-butanol pathway.DepositorInsertphaJ, ter, adhE2, phaA, phaB
ExpressionBacterialMutationNoneAvailable SinceNov. 16, 2021AvailabilityAcademic Institutions and Nonprofits only -
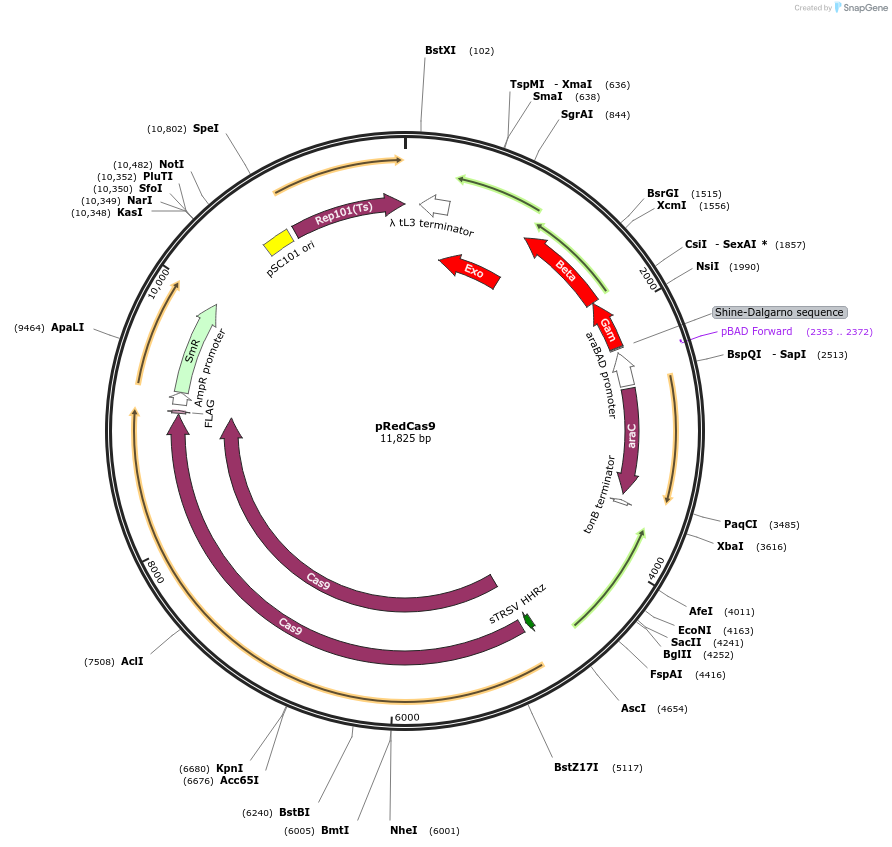
pRedCas9
Plasmid#240153PurposeExpresses lambda red machinery and cas9DepositorInsertsgam
beta
exo
cas9
UseCRISPRExpressionBacterialAvailable SinceOct. 22, 2025AvailabilityAcademic Institutions and Nonprofits only -
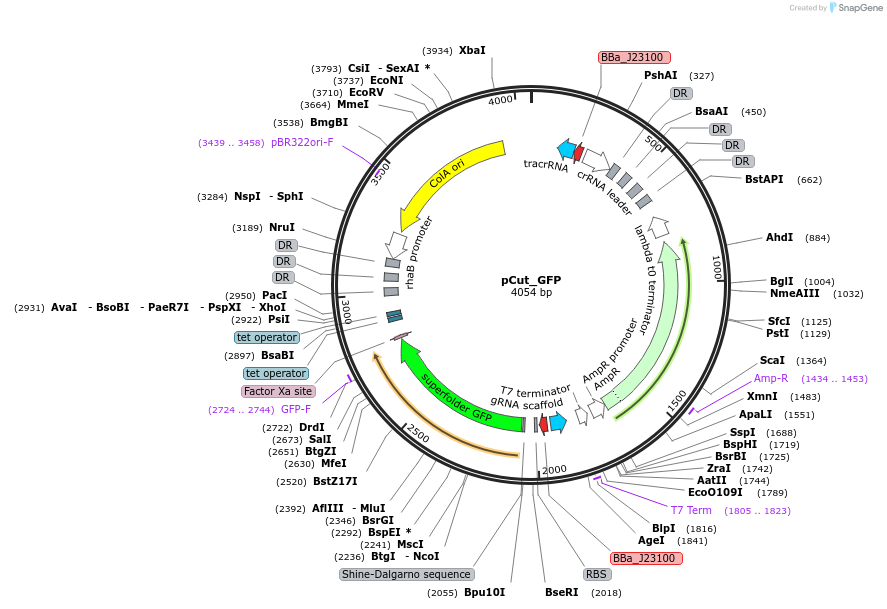
pCut_GFP
Plasmid#240154PurposeExpresses CRISPR guidesDepositorInsertsconstant cut sites CRISPR guide array
"self-curing" CRISPR guide array
GFP
UseCRISPRExpressionBacterialAvailable SinceOct. 22, 2025AvailabilityAcademic Institutions and Nonprofits only -
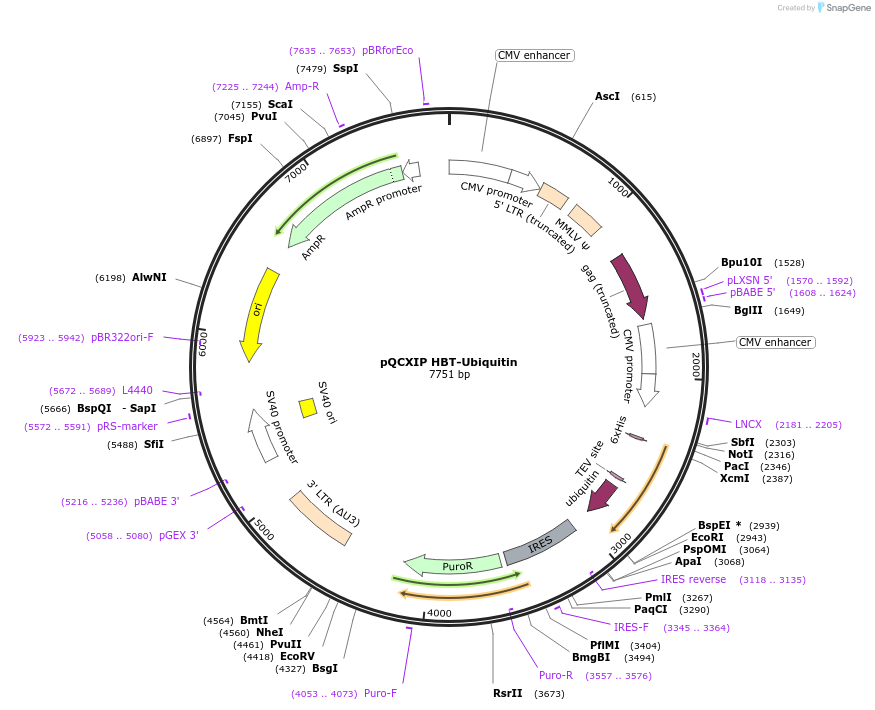
pQCXIP HBT-Ubiquitin
Plasmid#26865DepositorInsertubiquitin
UseRetroviralTagsHBTExpressionMammalianAvailable SinceDec. 15, 2010AvailabilityAcademic Institutions and Nonprofits only