We narrowed to 20,480 results for: Cre
-
Plasmid#12742PurposeThis plasmid was designed for screening purposes and may contain discrepancies relative to the current canonical GenBank version of the gene.DepositorInsertVV.Orf54
ExpressionMammalianAvailable SinceDec. 1, 2006AvailabilityAcademic Institutions and Nonprofits only -
DCT.VV.Orf57
Plasmid#12745PurposeThis plasmid was designed for screening purposes and may contain discrepancies relative to the current canonical GenBank version of the gene.DepositorInsertVV.Orf57
ExpressionMammalianAvailable SinceDec. 1, 2006AvailabilityAcademic Institutions and Nonprofits only -
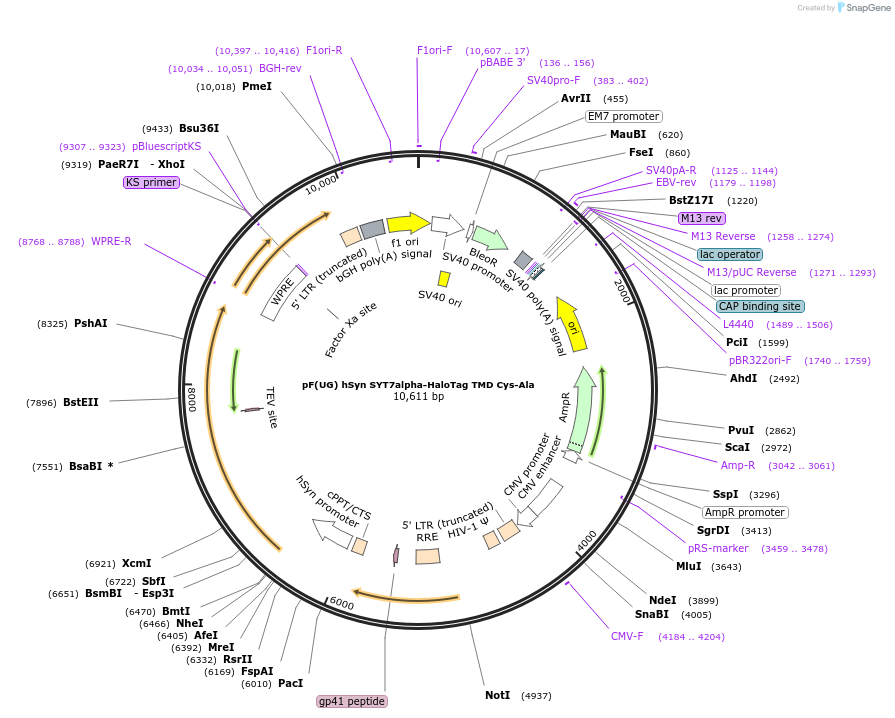
pF(UG) hSyn SYT7alpha-HaloTag TMD Cys-Ala
Plasmid#179716PurposeLentiviral expression of mouse Syt7 palmitoylation site mutant with C terminal HaloTagDepositorInsertSYT7 (Syt7 Mouse)
UseLentiviralAvailabilityAcademic Institutions and Nonprofits only -
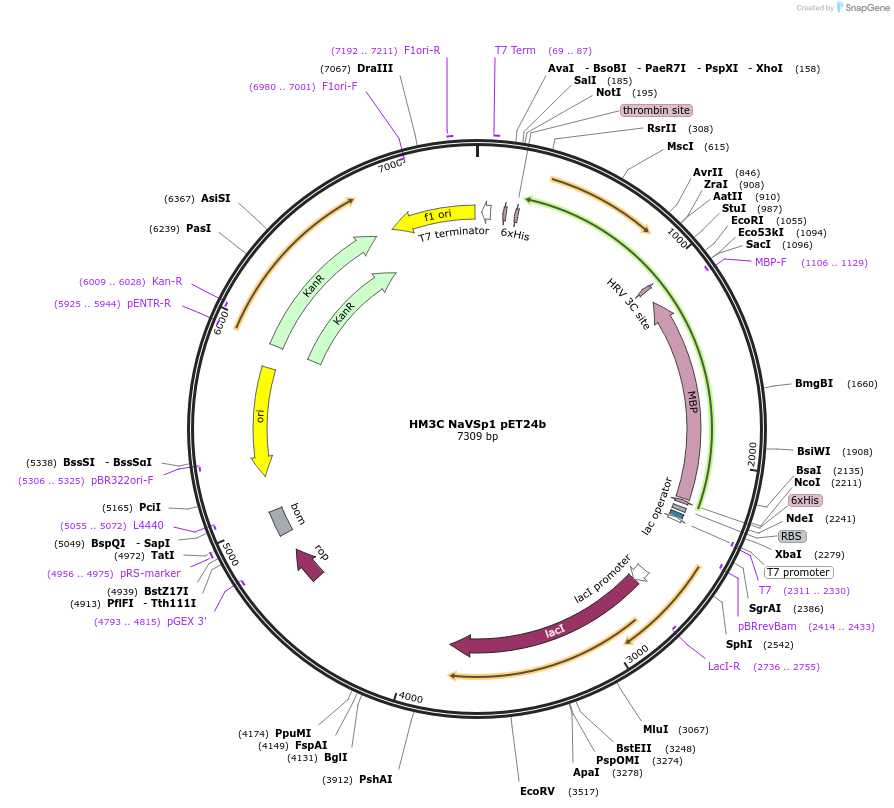
HM3C NaVSp1 pET24b
Plasmid#128088PurposeBacterial expression vector for NaVSp1 bacterial sodium channel. It expresses for HIStag-MBP-3C-channel proteinDepositorInsertNaVSp1p
Tags3C Cleavage, His Tag, and MBPExpressionBacterialAvailabilityAcademic Institutions and Nonprofits only -
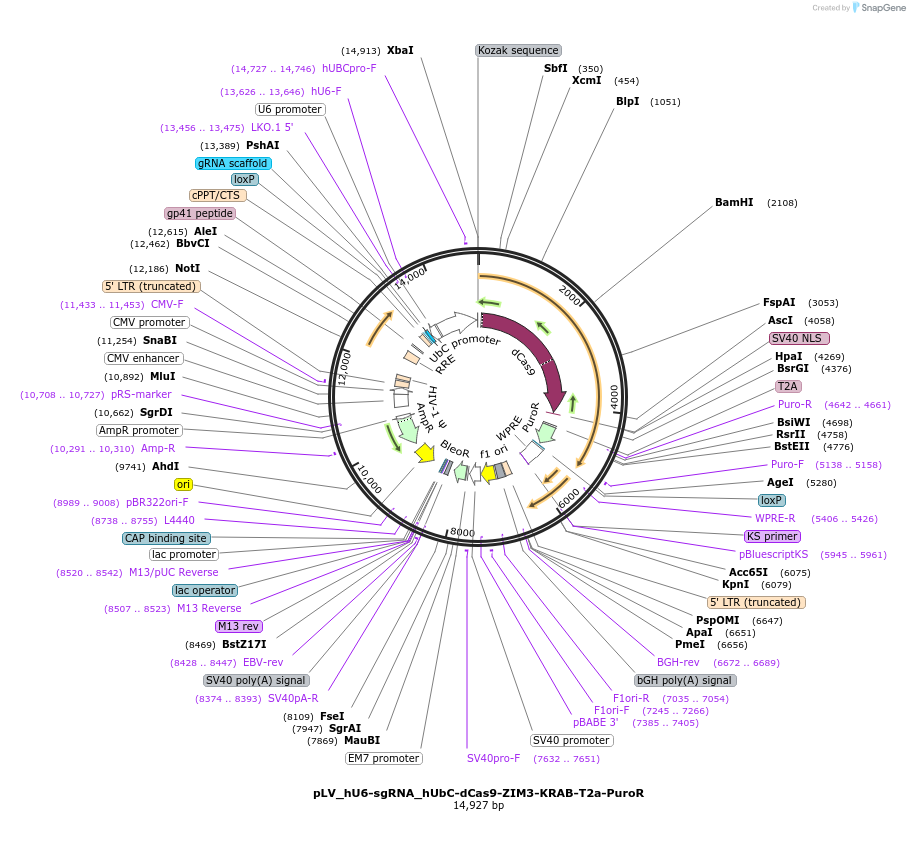
pLV_hU6-sgRNA_hUbC-dCas9-ZIM3-KRAB-T2a-PuroR
Plasmid#172982Purposecontrol & cloning vector for CRISPRi expressing dead Cas9-ZIM3-KRAB fusionDepositorInsertcontrol sgRNA
UseCRISPR and LentiviralAvailable SinceOct. 11, 2023AvailabilityAcademic Institutions and Nonprofits only -
ABE8.20-m
Plasmid#136300Purposeexpresses ABE8.20-m in mammalian cellsDepositorInsertABE8.20-m
TagsC-terminal BPNLSExpressionMammalianMutationD10A in S. pyogenes Cas9, TadA mutations describe…Available SinceMarch 16, 2020AvailabilityAcademic Institutions and Nonprofits only -
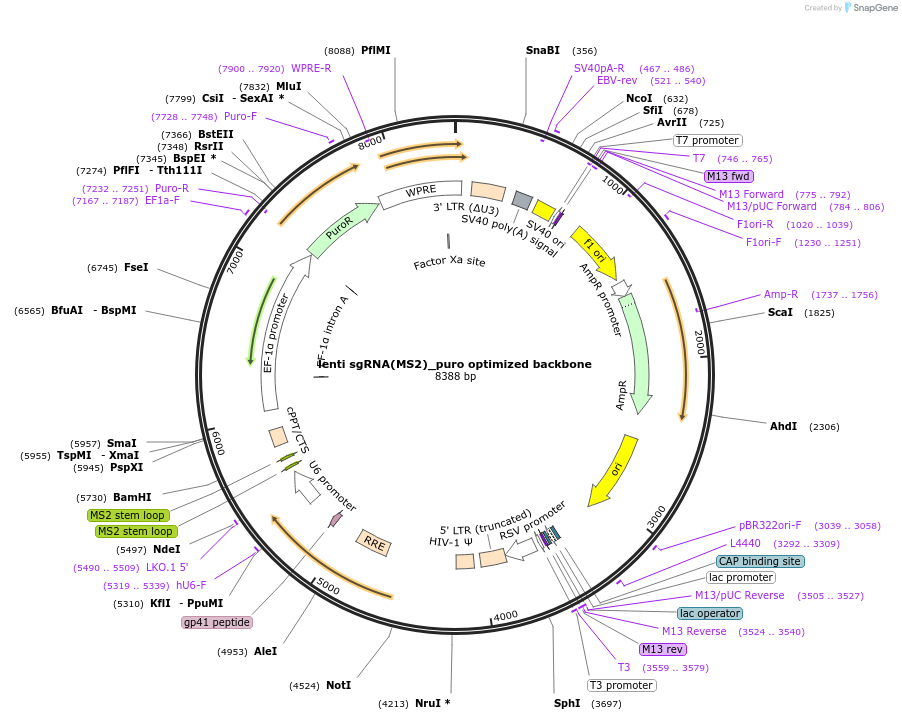
lenti sgRNA(MS2)_puro optimized backbone
Plasmid#73797Purposeoptimized lenti sgRNA cloning backbone with MS2 loops at tetraloop and stemloop 2 and EF1a-puro resistance marker. Contains BsmBI sites for insertion of spacer sequences.DepositorHas ServiceCloning Grade DNATypeEmpty backboneUseCRISPR and LentiviralExpressionMammalianPromoterU6 and EF1AAvailable SinceMarch 15, 2016AvailabilityAcademic Institutions and Nonprofits only -
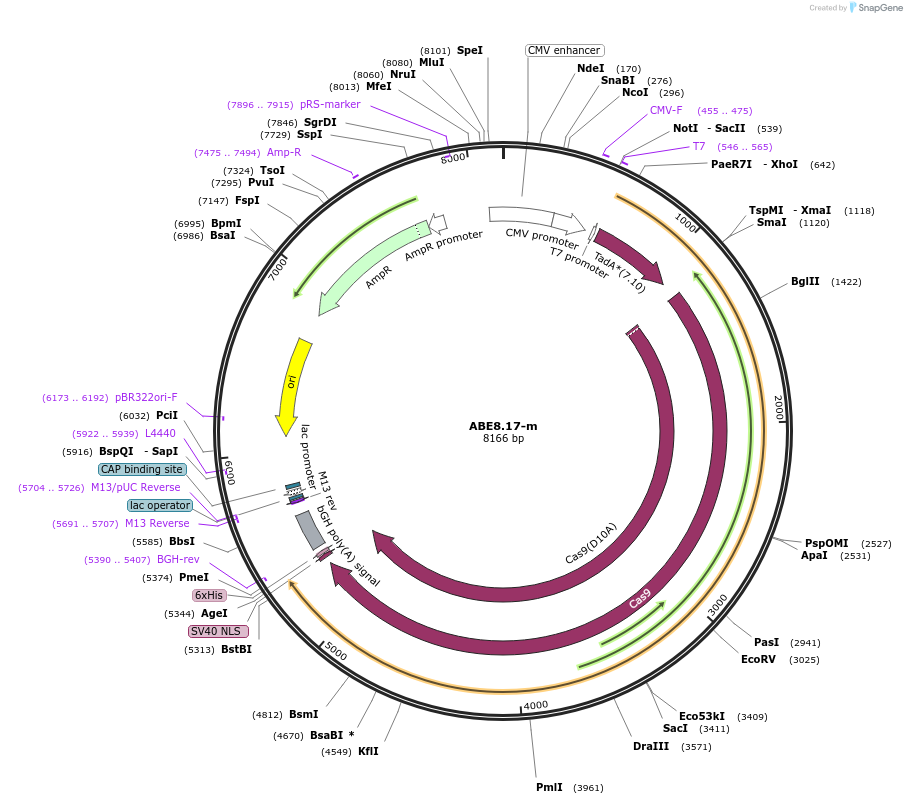
ABE8.17-m
Plasmid#136298Purposeexpresses ABE8.17-m in mammalian cellsDepositorInsertABE8.17-m
TagsC-terminal BPNLSExpressionMammalianMutationD10A in S. pyogenes Cas9, TadA mutations describe…Available SinceMarch 16, 2020AvailabilityAcademic Institutions and Nonprofits only -
eeBxb1
Plasmid#222339PurposePlasmid for expressing evolved and engineered Bxb1 in the mammalian cellsDepositorInsertCodon-optimized evolved and engineered Bxb1
UseCRISPRExpressionMammalianPromoterCMVAvailable SinceJune 27, 2024AvailabilityAcademic Institutions and Nonprofits only -
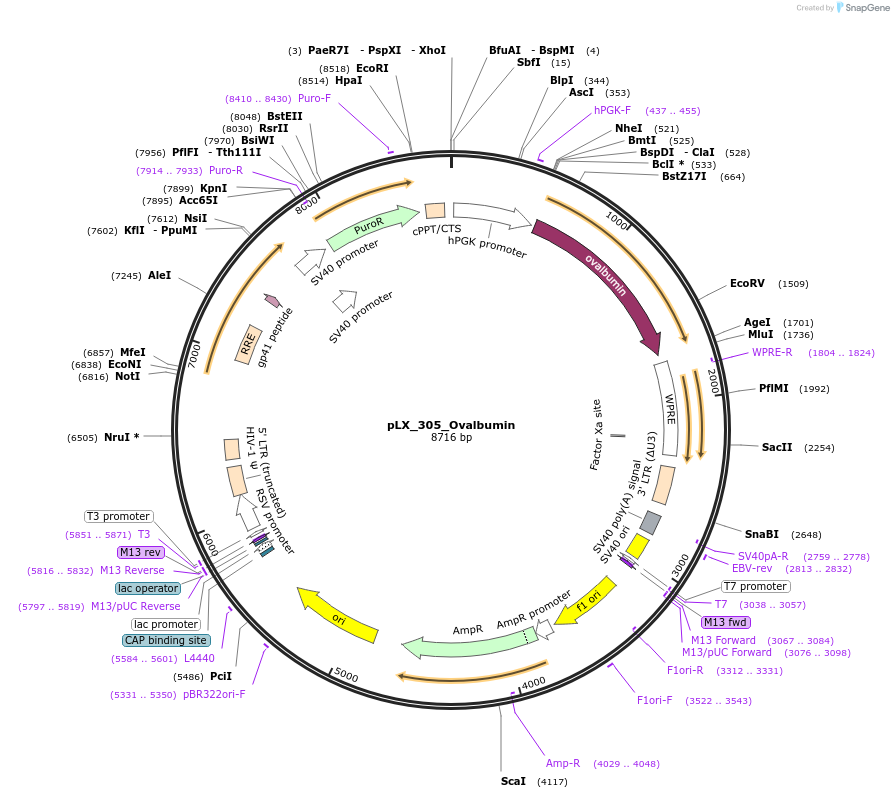
pLX_305_Ovalbumin
Plasmid#184924PurposeLentiviral vector for expressing Ovalbumin. Also expresses the puromycin resistance gene as a selectable marker.DepositorInsertOvalbumin (OVAL Chicken)
UseLentiviralExpressionMammalianMutationOur vector has an A188T mutation that does not im…PromoterPGKAvailable SinceFeb. 6, 2024AvailabilityAcademic Institutions and Nonprofits only -
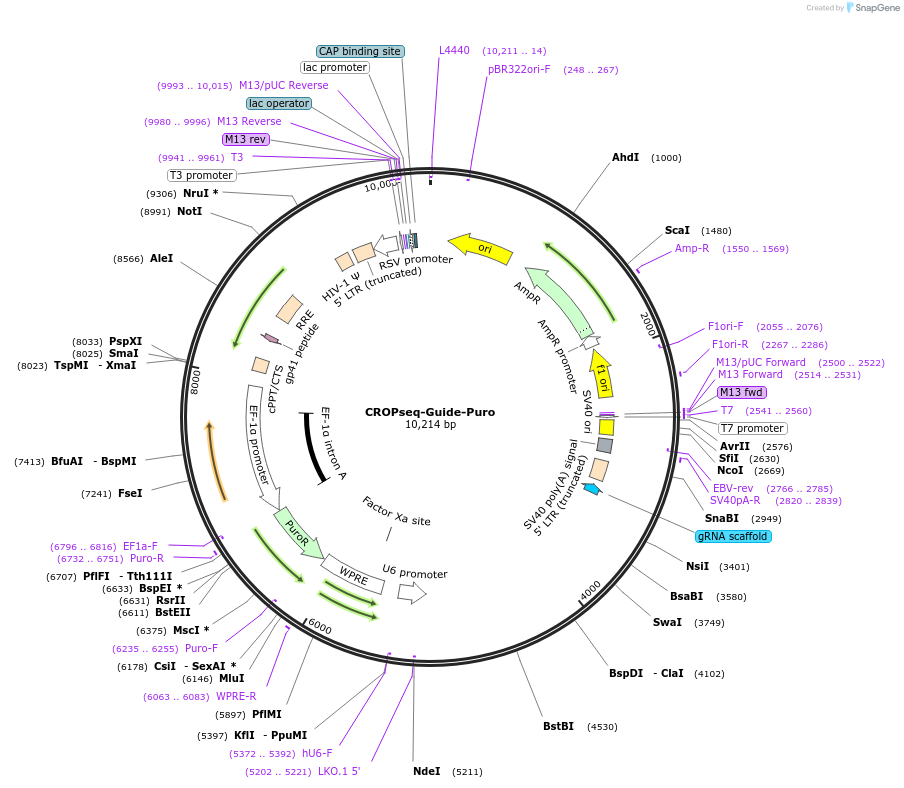
CROPseq-Guide-Puro
Plasmid#86708PurposeMakes CRISPR gRNAs detectable by single-cell RNA-seq. The gRNA is expressed as part of the puromycin resistance mRNA transcribed by RNA Pol II and in its functional form from the hU6 promoter.DepositorInsertEF1a-Puro-WPRE-hU6-gRNA
UseCRISPR and LentiviralExpressionMammalianPromoterhU6Available SinceFeb. 9, 2017AvailabilityAcademic Institutions and Nonprofits only -
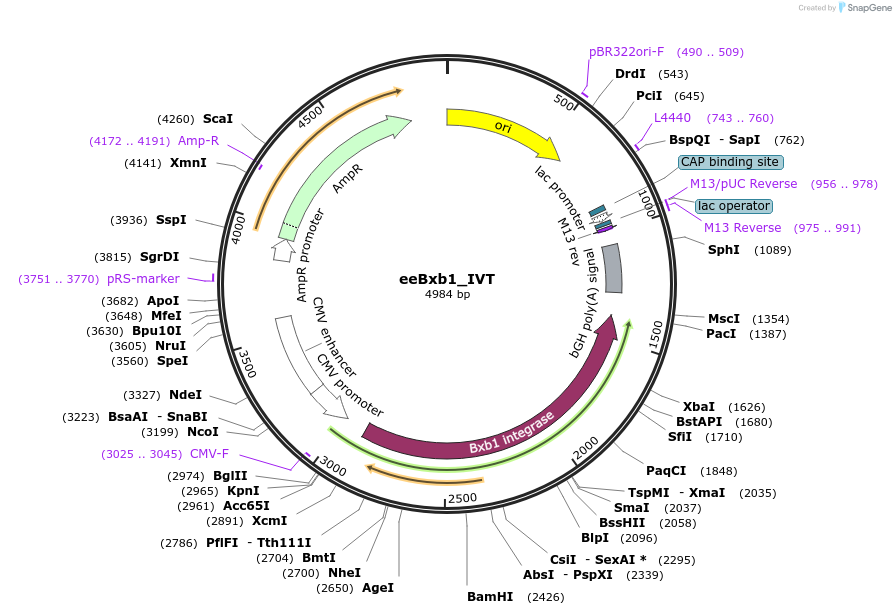
eeBxb1_IVT
Plasmid#222342PurposePlasmid for evolved and engineered Bxb1 mRNA in vitro transcription templateDepositorInsertCodon-optimized evolved and engineered Bxb1
UseTemplate for in vitro transcriptionPromoterT7 (inactivated)Available SinceJuly 25, 2024AvailabilityAcademic Institutions and Nonprofits only -
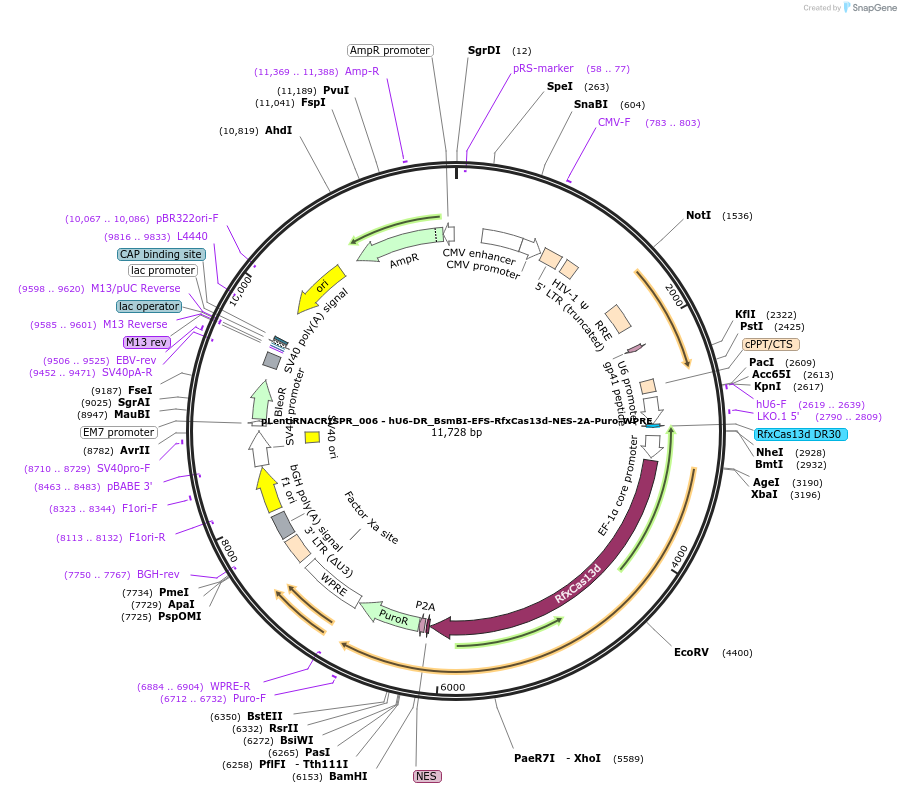
pLentiRNACRISPR_006 - hU6-DR_BsmBI-EFS-RfxCas13d-NES-2A-Puro-WPRE
Plasmid#138148PurposeExpresses RfxCas13d in mammalian cells, localized in the cytoplasm. For cloning of guide RNAs compatible with RfxCas13d. Contains a 5' direct repeat. Clone using BsmBI. F overhang aaac. R overhang aaaaDepositorInsertCasRx
UseLentiviralTagsNESAvailable SinceDec. 1, 2020AvailabilityAcademic Institutions and Nonprofits only -
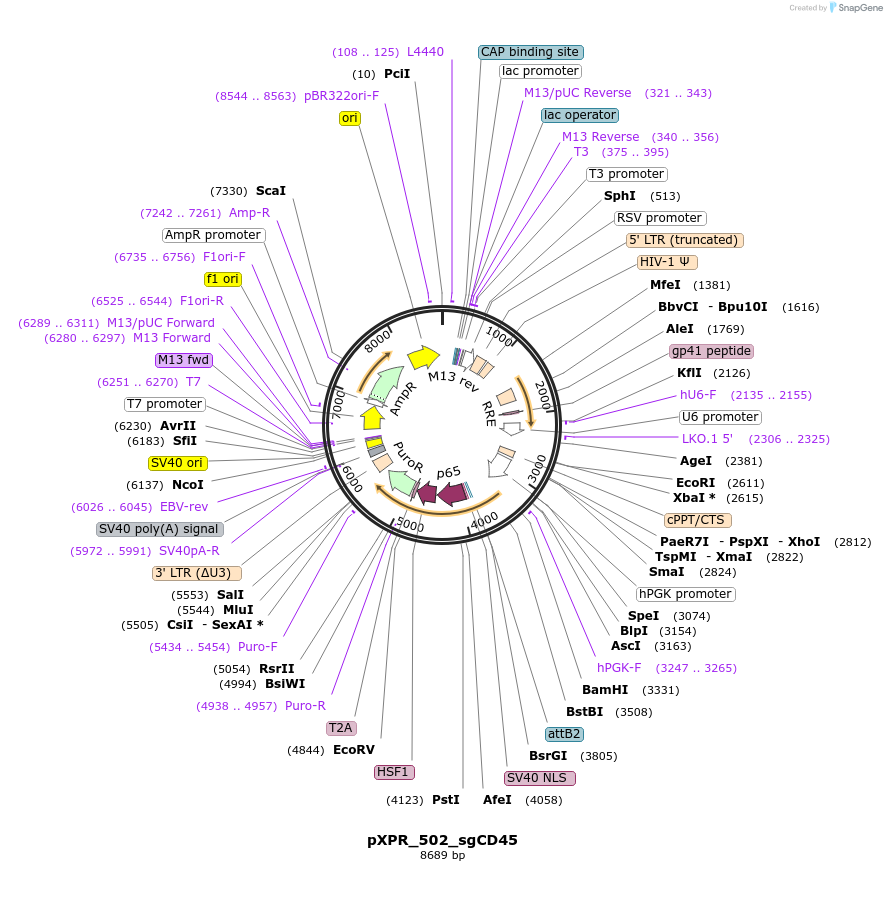
pXPR_502_sgCD45
Plasmid#158705PurposeCD45 CRISPRa controlDepositorInsertCD45 (PTPRC Human)
UseLentiviralAvailable SinceAug. 18, 2020AvailabilityAcademic Institutions and Nonprofits only -
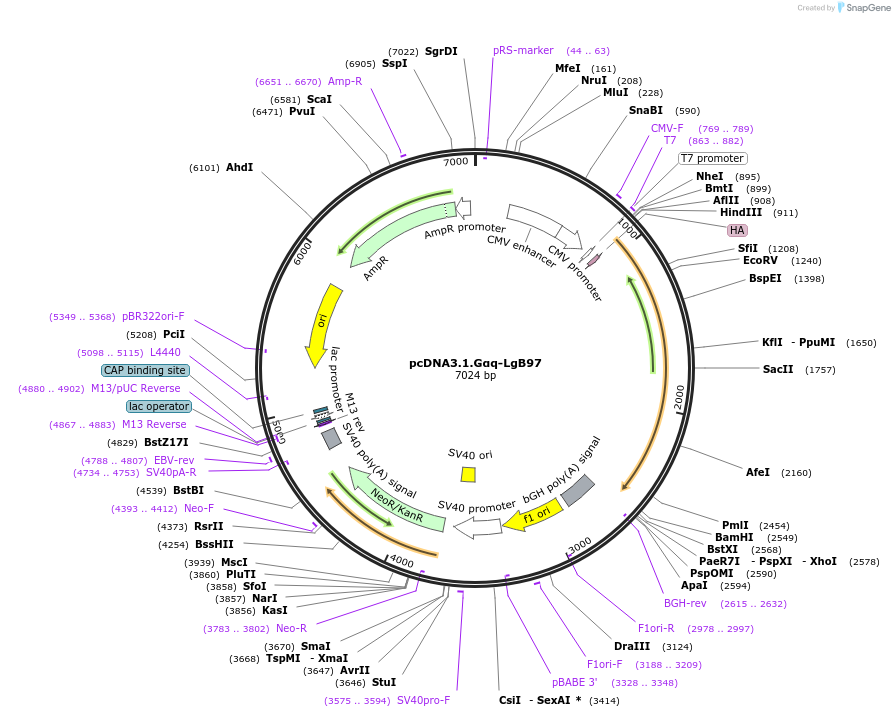
pcDNA3.1.Gαq-LgB97
Plasmid#134360PurposeNanoluc complementation assay. Expression of Gαq protein fused with the LgBiT(LgB)of the nanoluciferase inserted between residues 97 and 98 of Gαq. Addition of the HA epitope at N terminus of Gαq.DepositorAvailable SinceJuly 22, 2020AvailabilityAcademic Institutions and Nonprofits only -
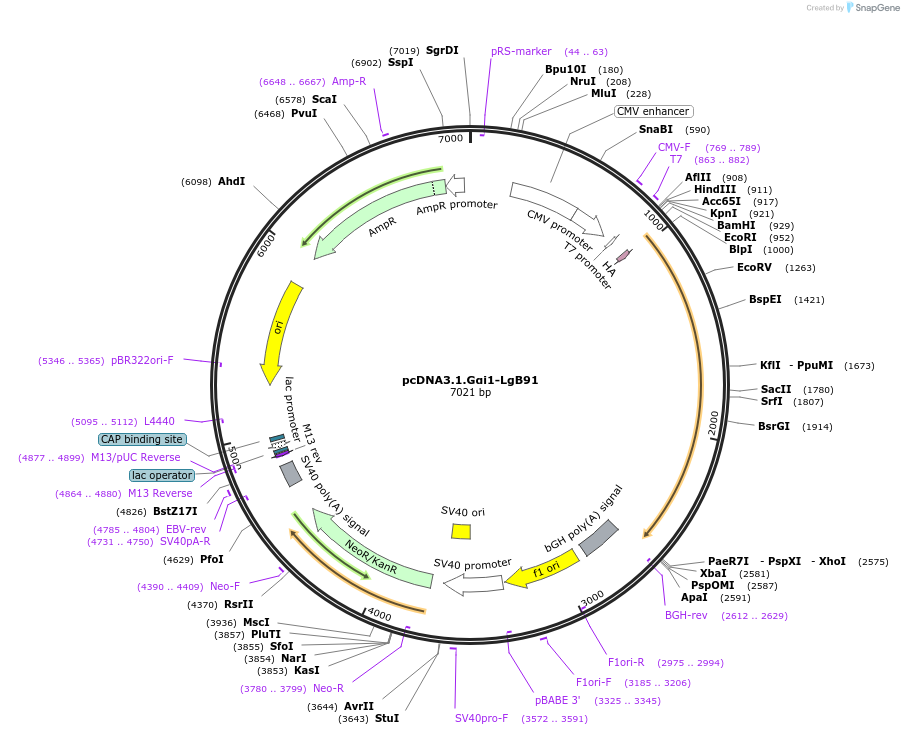
pcDNA3.1.Gαi1-LgB91
Plasmid#134342PurposeNanoluc complementation assay. Expression of Gαi1 protein fused with the LgBiT(LgB)of the nanoluciferase inserted between residues 91 and 92 of Gαi1. Addition of the HA epitope at N terminus of Gαi1.DepositorAvailable SinceJuly 22, 2020AvailabilityAcademic Institutions and Nonprofits only -
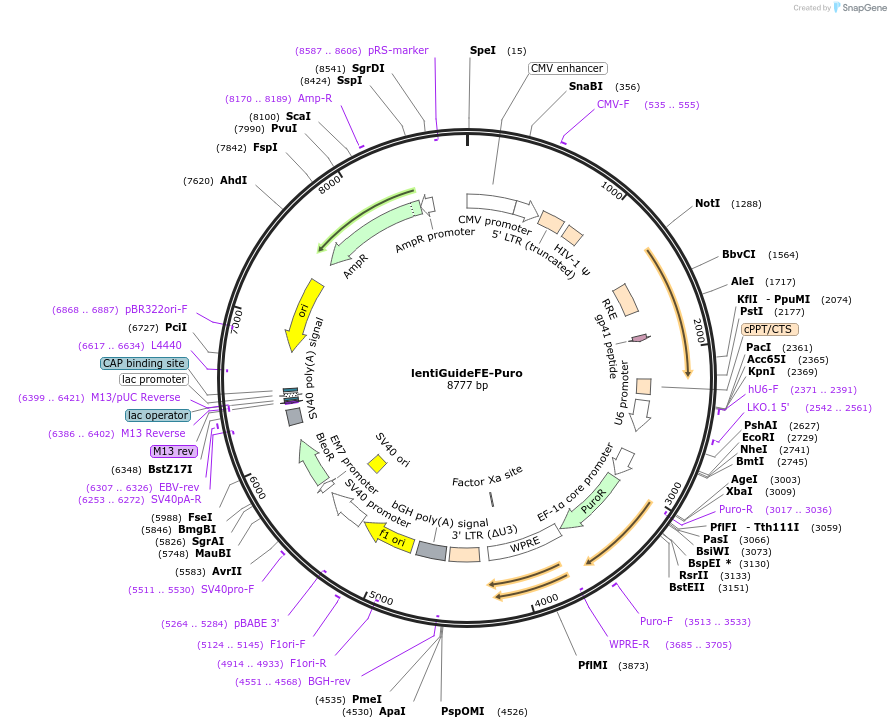
lentiGuideFE-Puro
Plasmid#170069PurposeExpresses S. pyogenes CRISPR chimeric RNA element (with F+E modifications) with customizable gRNA from U6 promoter and puromycin resistance from EFS-NS promoterDepositorInsertCas9 guide RNA scaffold with the F+E scaffold modification
UseCRISPR and LentiviralExpressionMammalianPromoterU6Available SinceOct. 12, 2022AvailabilityAcademic Institutions and Nonprofits only -
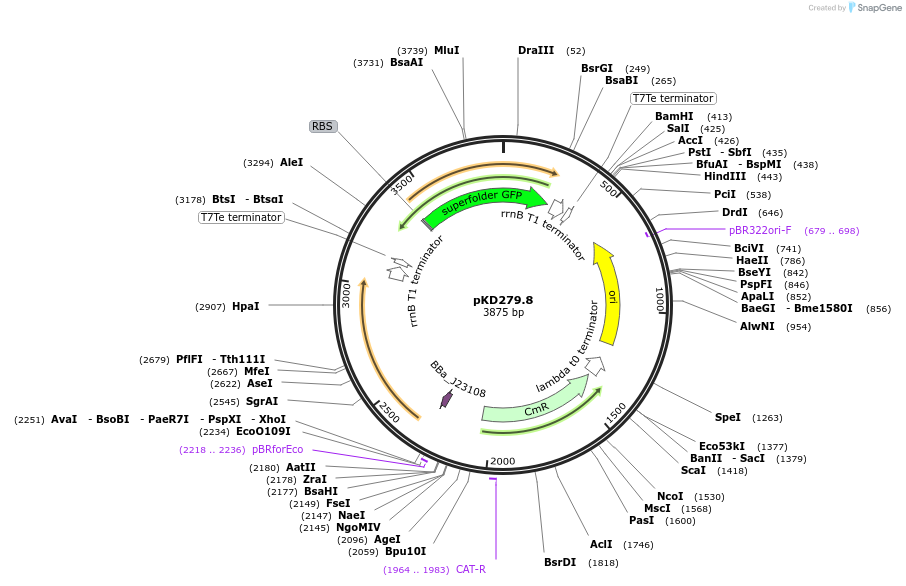
pKD279.8
Plasmid#173171PurposeEncodes expression of SO_4388(REC)-PsdR(DBD) 137 and an sfGFP reporterDepositorInsertsSO_4388(REC)-PsdR(DBD) 137
sfGFP
ExpressionBacterialPromoterJ23108 and PpsdA110Available SinceSept. 10, 2021AvailabilityAcademic Institutions and Nonprofits only -
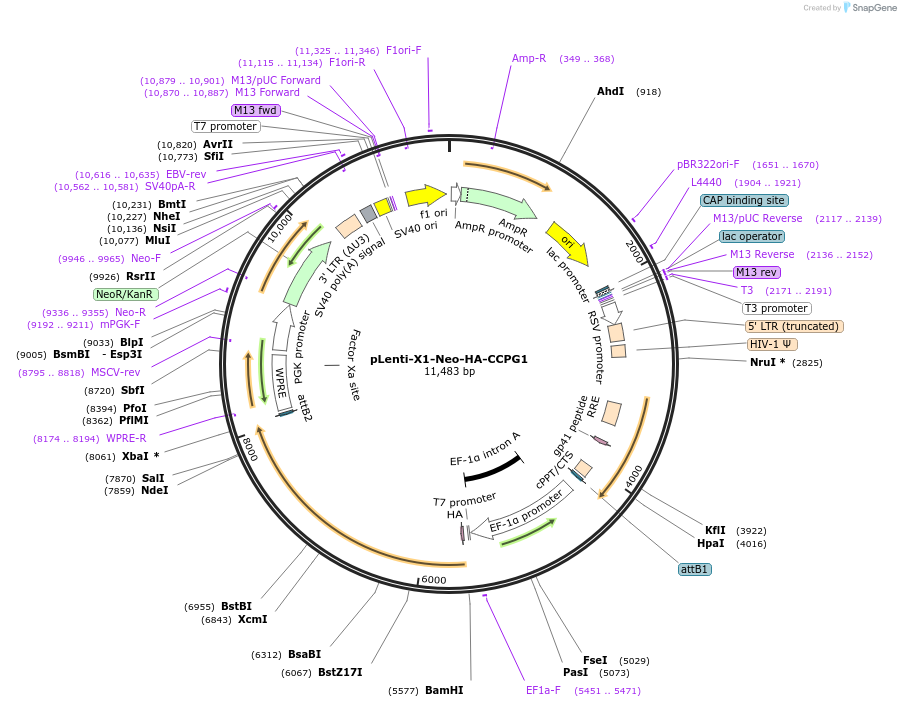
pLenti-X1-Neo-HA-CCPG1
Plasmid#139867PurposeLentiviral expression of HA-tagged human CCPG1DepositorAvailable SinceMarch 24, 2020AvailabilityAcademic Institutions and Nonprofits only -
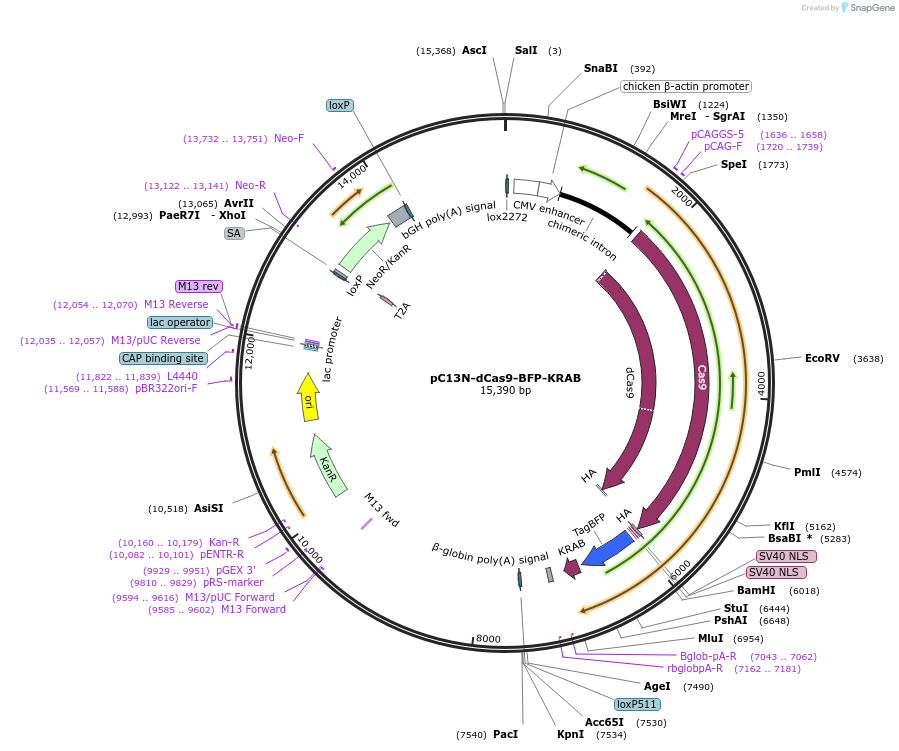
pC13N-dCas9-BFP-KRAB
Plasmid#127968Purposeconstitutive expression of dCas9-BFP-KRAB from the CLYBL locus (Ward lab)DepositorInsertCLYBL-CAG-dCas9-NLS-BFP-KRAB
UseTALENExpressionMammalianPromoterCAGAvailable SinceAug. 28, 2019AvailabilityAcademic Institutions and Nonprofits only