We narrowed to 13,339 results for: ache
-
-
-
-
-
-
-
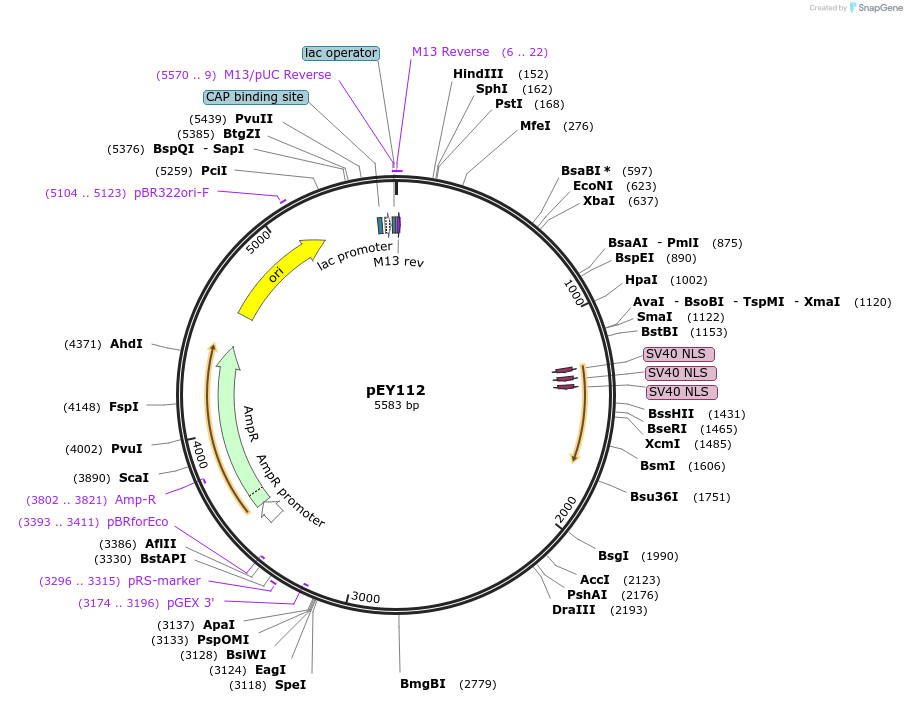
pEY102
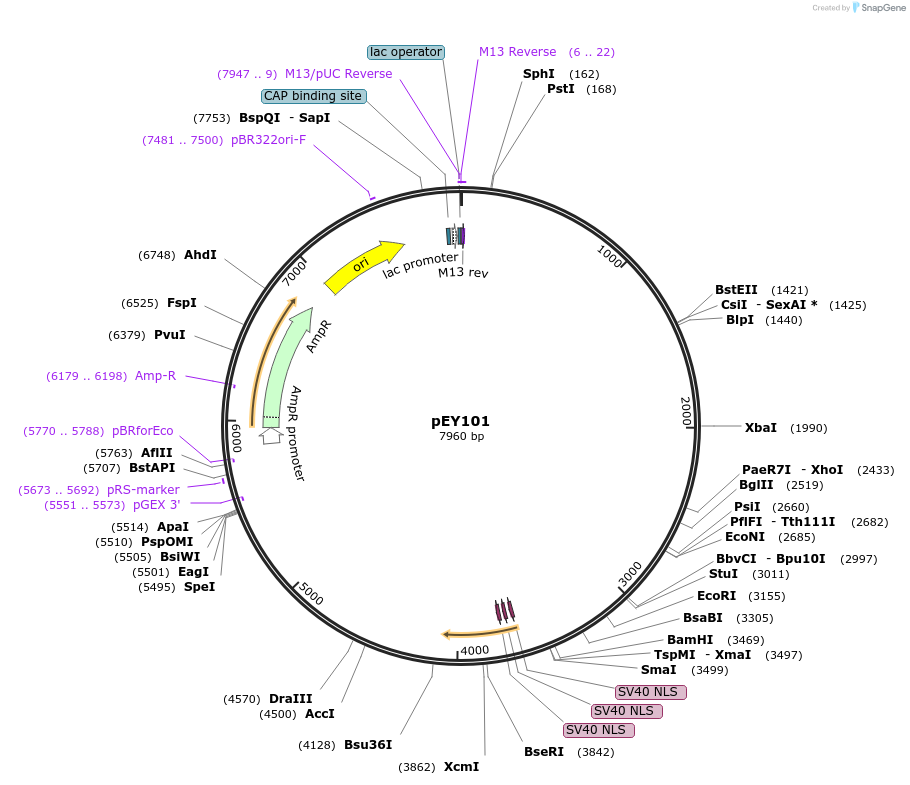
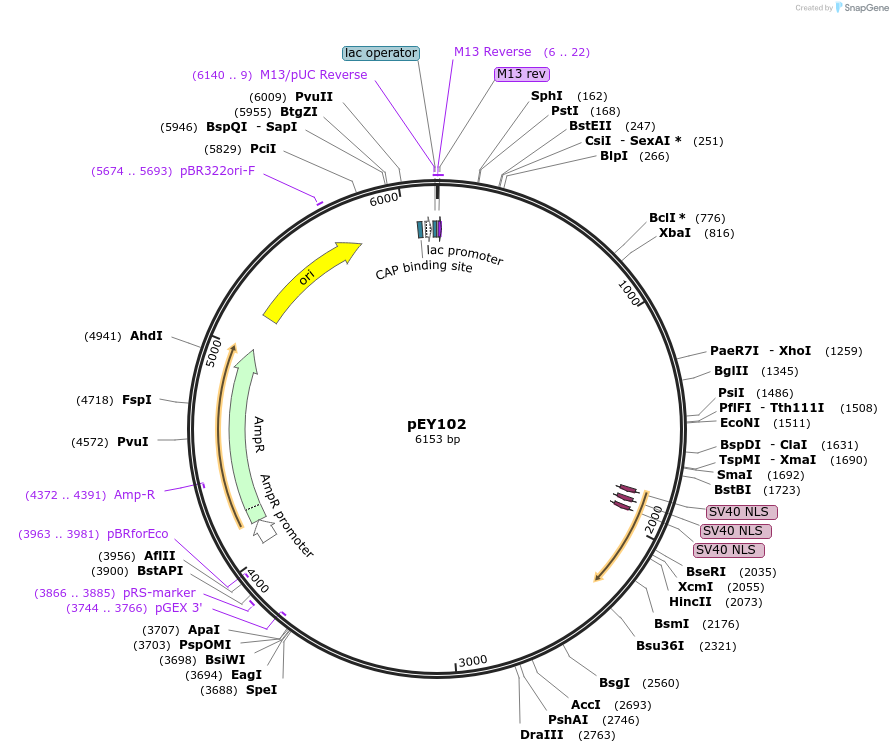
Plasmid#223449Purposelin-11(intron 3) fluorescent neural reporter driving nuclear mNeptune2.5 expression (refer to NeuroPAL paper for expression)DepositorInsertlin-11(intron 3) (lin-11 Nematode)
ExpressionWormAvailable SinceAug. 7, 2024AvailabilityAcademic Institutions and Nonprofits only -
-
-
-
-
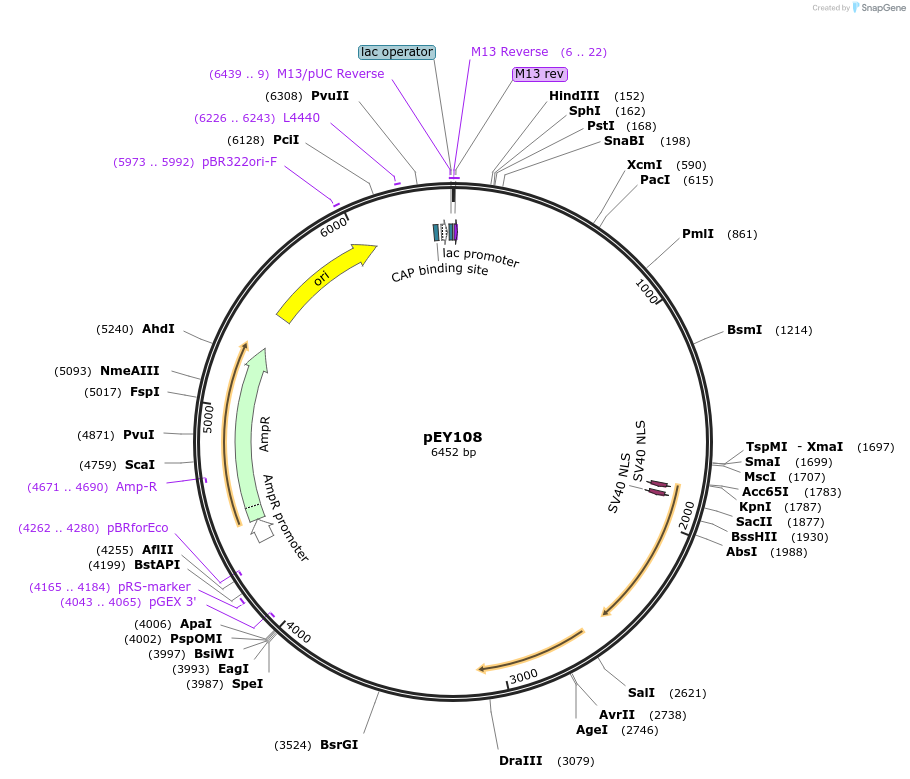
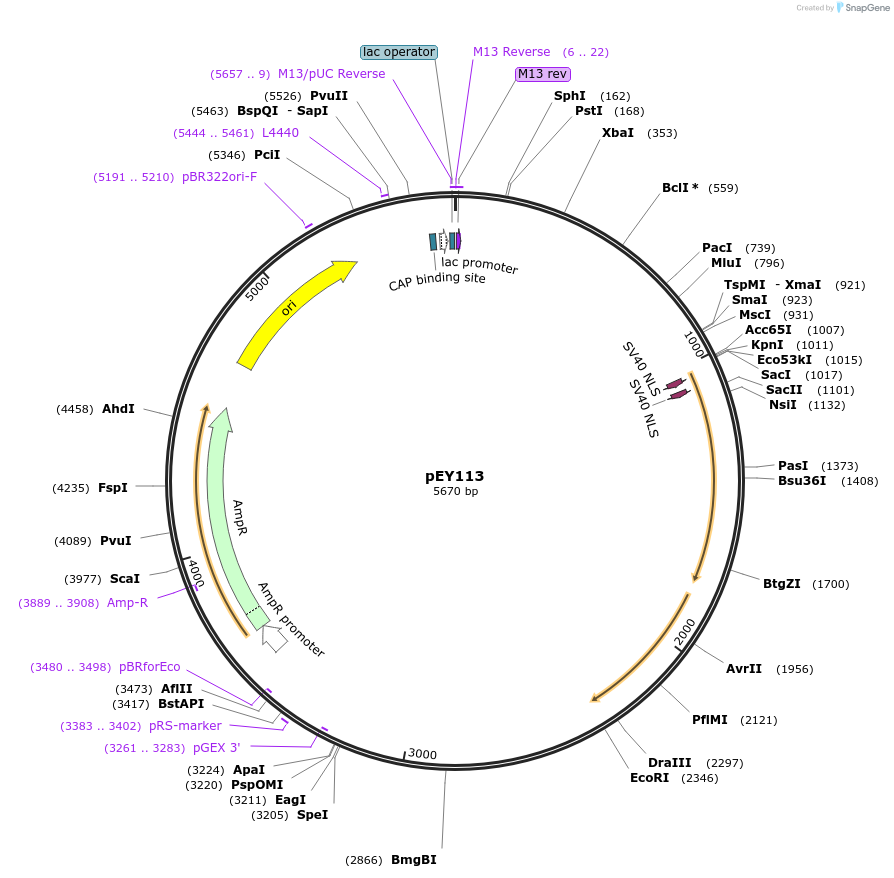
pEY108
Plasmid#223455Purposensy-5 (prom4) fluorescent neural reporter driving nuclear CyOFP1 expression (refer to NeuroPAL paper for expression)DepositorInsertnsy-5(prom4)
ExpressionWormAvailable SinceAug. 7, 2024AvailabilityAcademic Institutions and Nonprofits only -
-
-
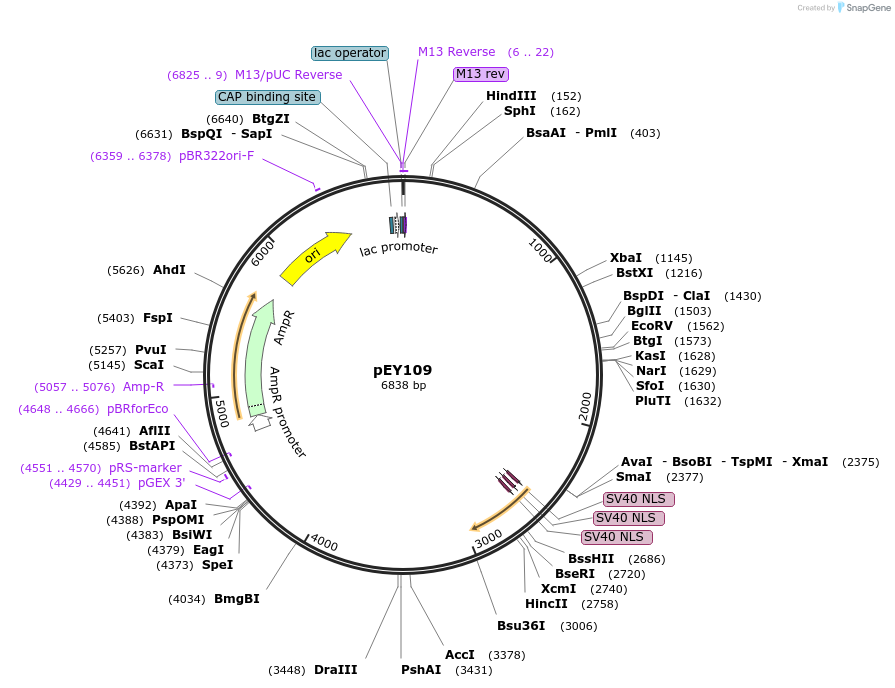
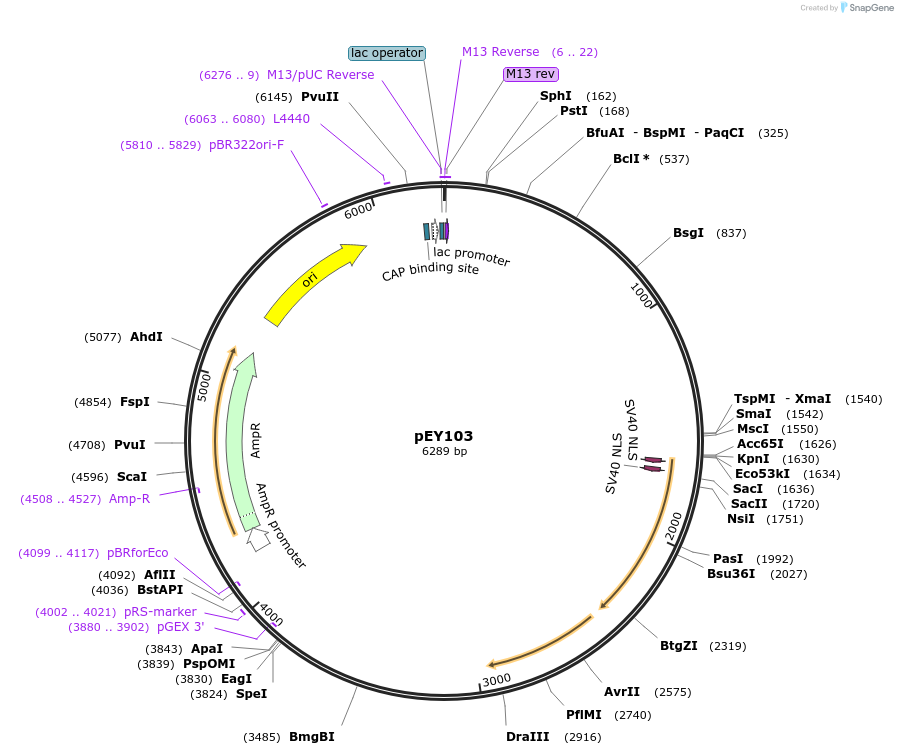
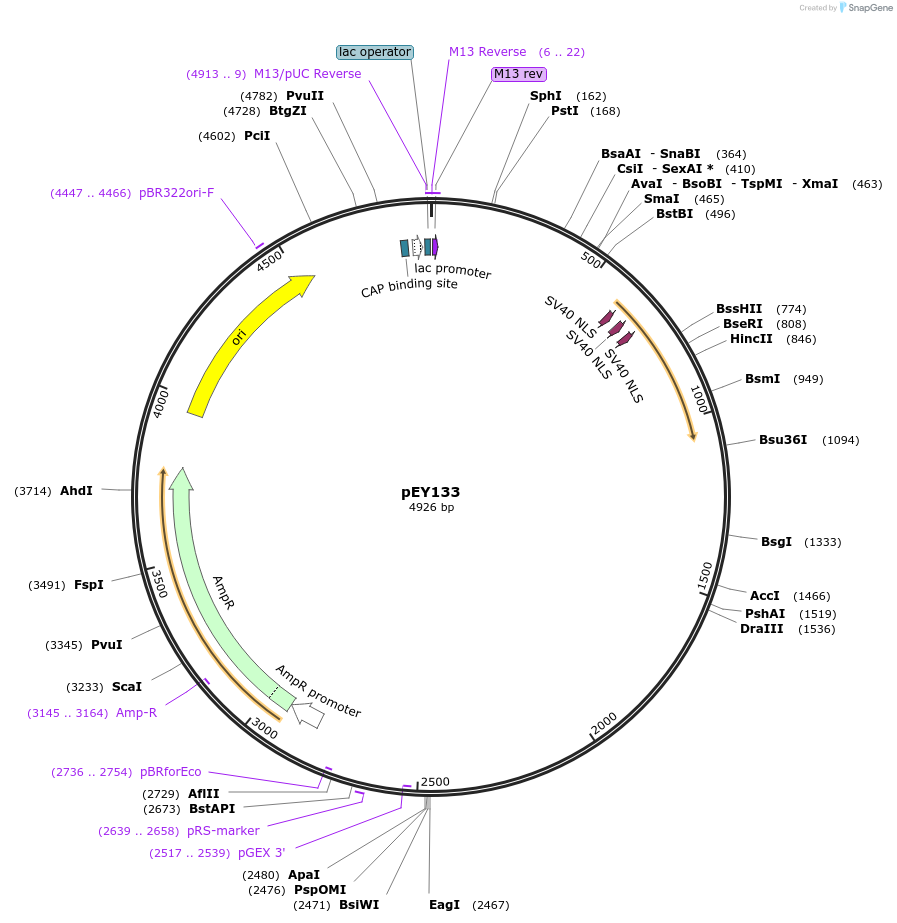
pEY103
Plasmid#223450Purposelin11(intron 7) fluorescent neural reporter driving nuclear mTagBFP2 expression (refer to NeuroPAL paper for expression)DepositorInsertlin-11(intron 7) (lin-11 Nematode)
ExpressionWormAvailable SinceAug. 7, 2024AvailabilityAcademic Institutions and Nonprofits only -
-
-
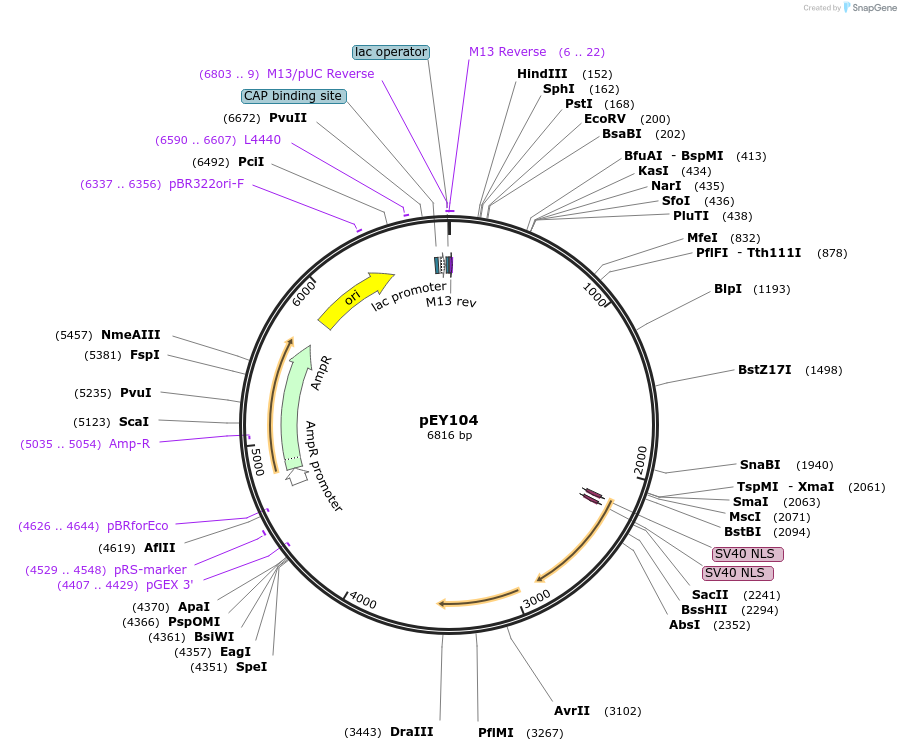
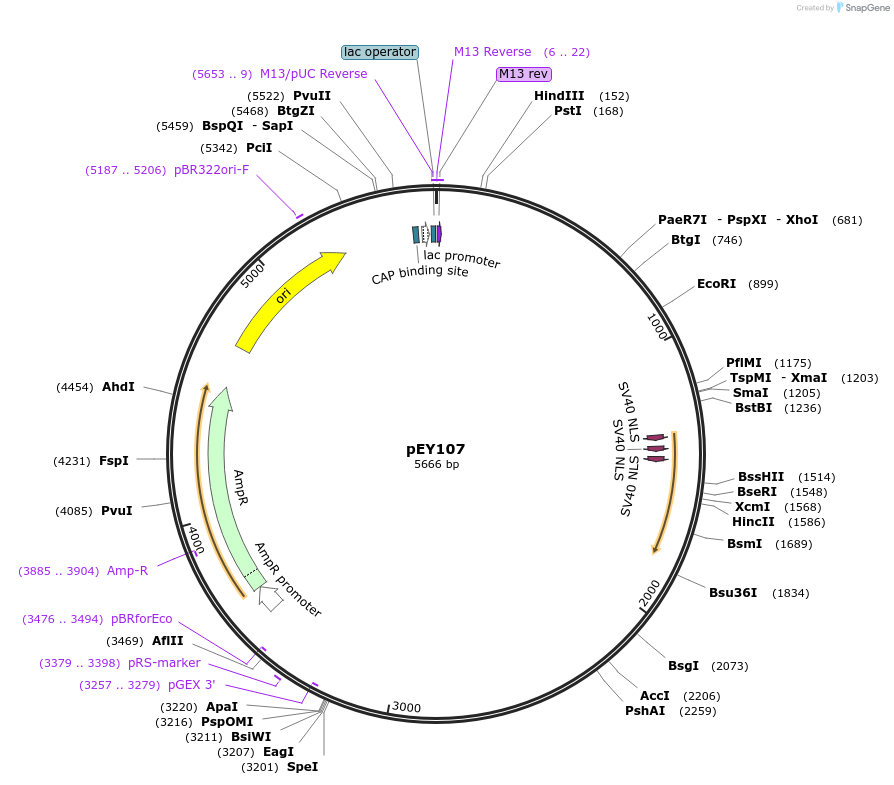
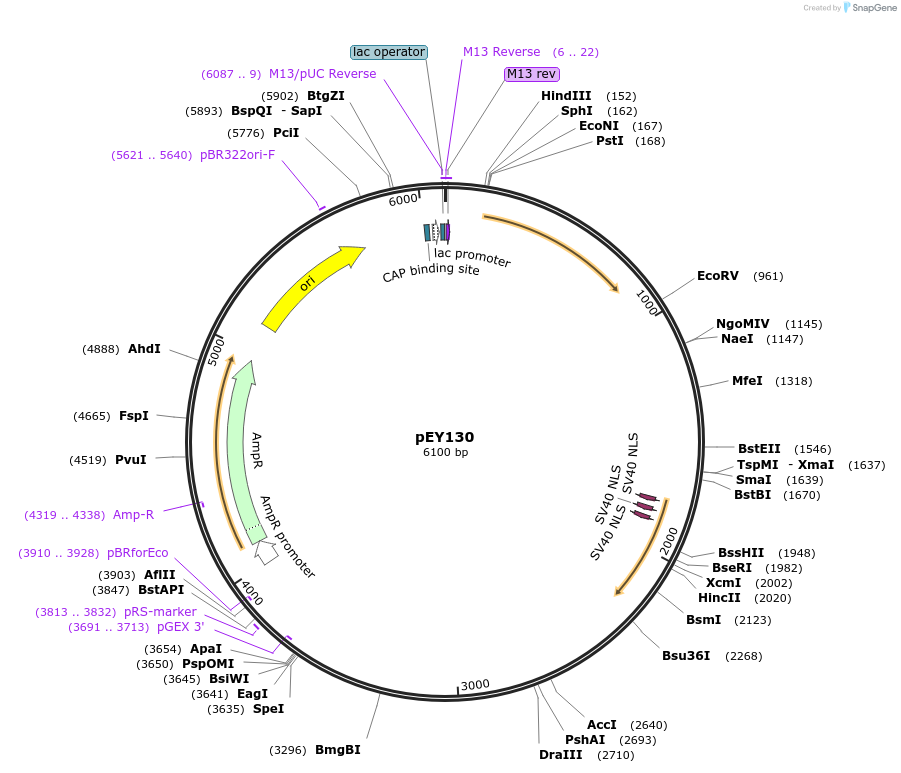
pEY107
Plasmid#223454Purposensy-5 (prom3) fluorescent neural reporter driving nuclear mNeptune2.5 expression (refer to NeuroPAL paper for expression)DepositorInsertnsy-5(prom3)
ExpressionWormAvailable SinceAug. 6, 2024AvailabilityAcademic Institutions and Nonprofits only -
-
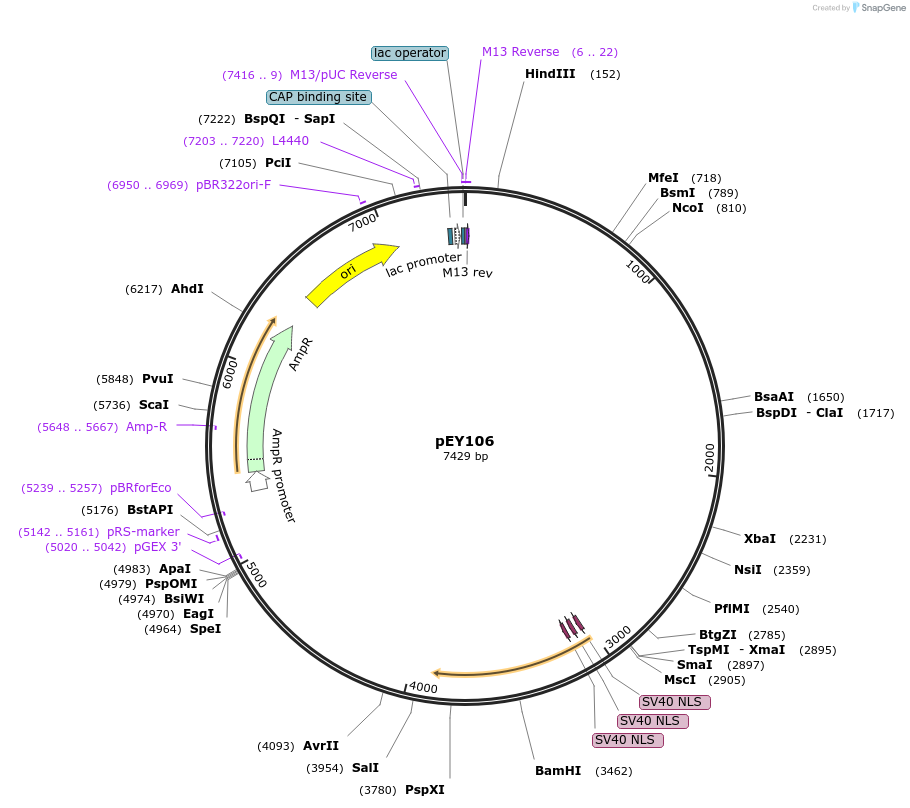
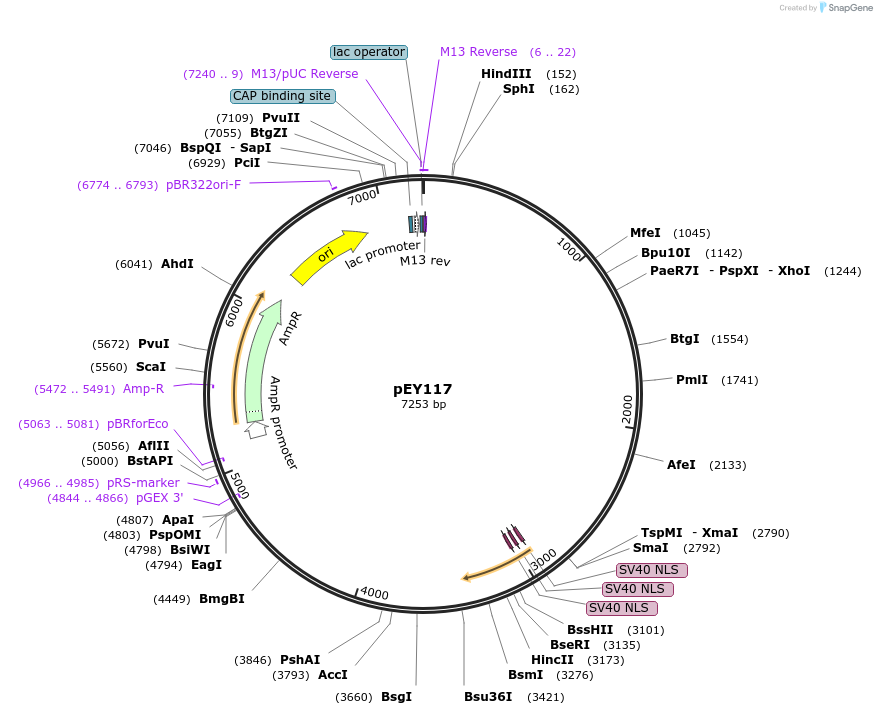
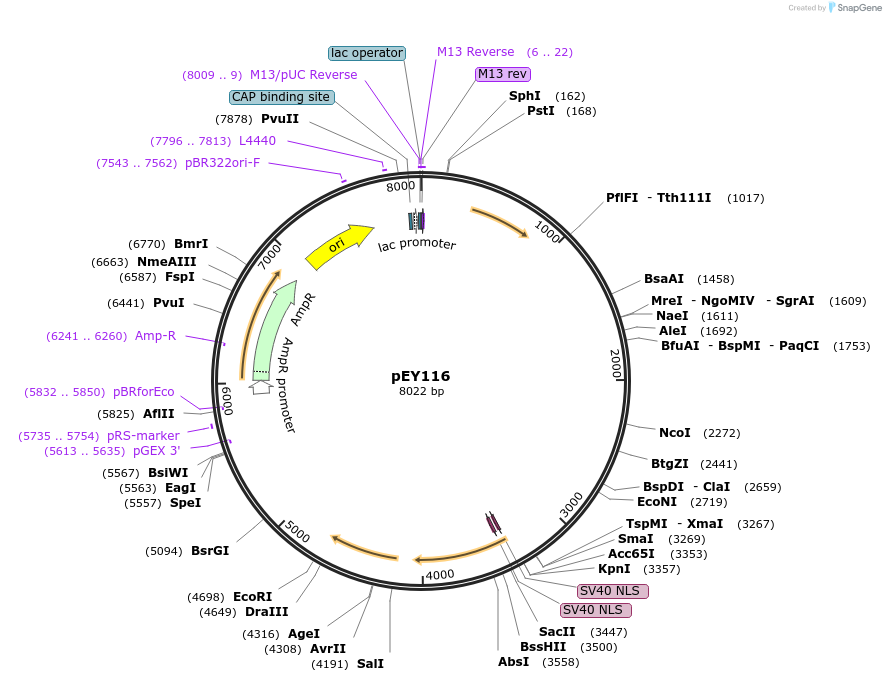
pEY117
Plasmid#223464Purposeser-2(prom1B) fluorescent neural reporter driving nuclear mNeptune2.5 expression (refer to NeuroPAL paper for expression)DepositorInsertser-2(prom1B) (ser-2 Nematode)
ExpressionWormAvailable SinceAug. 6, 2024AvailabilityAcademic Institutions and Nonprofits only -
-
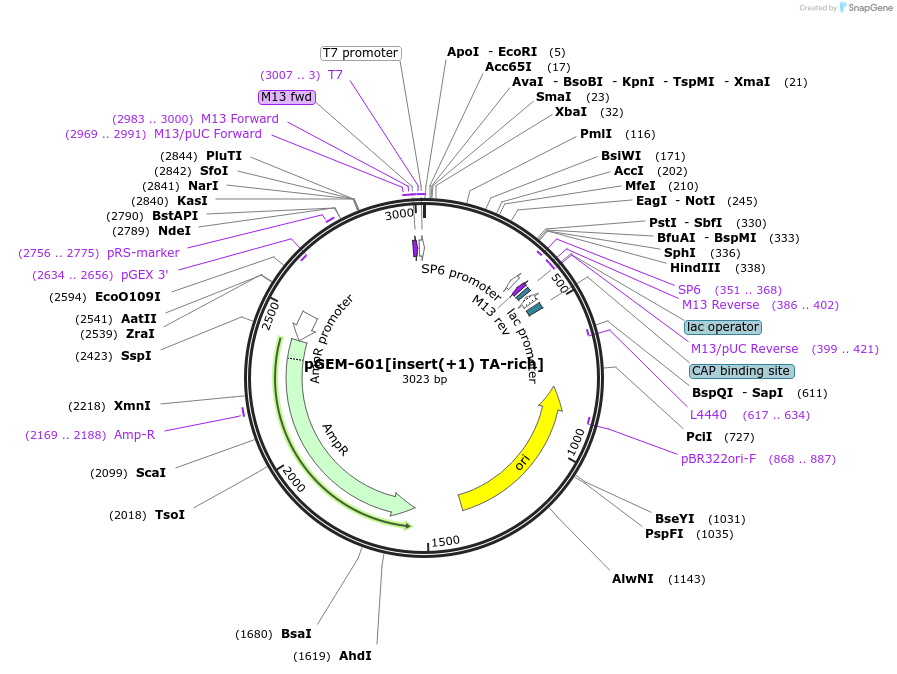
pGEM-601[insert(+1) TA-rich]
Plasmid#221196Purpose601 nucleosome positioning sequence with an additional base pair inserted 22 nt from the dyad on the TA-rich side of the 601DepositorInsert601[insert(+1)TA-rich]
UseUnspecifiedMutation601 has an additional mutation added 22 bp from d…Available SinceAug. 6, 2024AvailabilityAcademic Institutions and Nonprofits only -
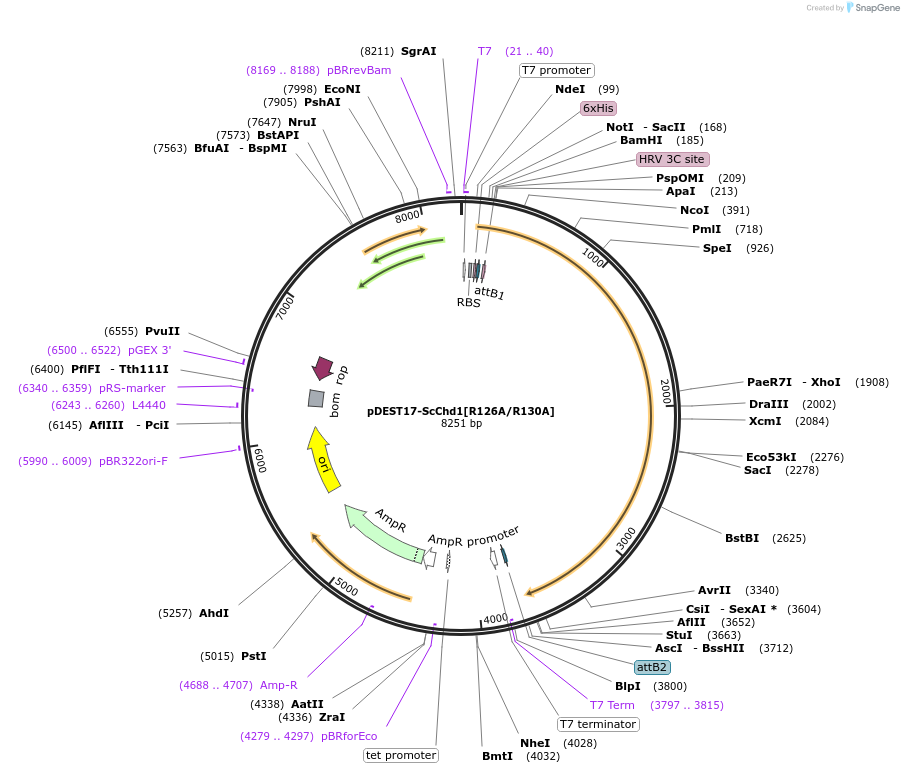
pDEST17-ScChd1[R126A/R130A]
Plasmid#221197PurposeScChd1-aa118-1274 chromatin remodeler with R126A and R130A mutationsDepositorInsertScChd1 (aa118-1274)
Tags6xHis-tag (Precission protease cleavable)ExpressionBacterialMutationR126A, R130APromoterT7Available SinceAug. 6, 2024AvailabilityAcademic Institutions and Nonprofits only -
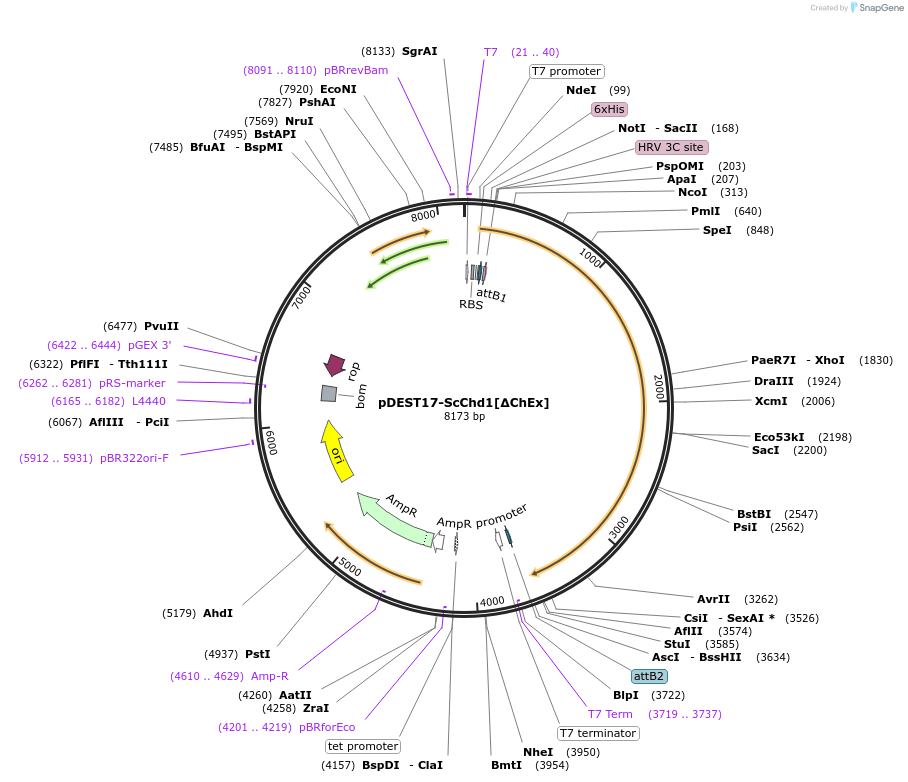
pDEST17-ScChd1[∆ChEx]
Plasmid#221199PurposeScChd1-aa142-1274 chromatin remodeler with ChEx domain deletedDepositorInsertScChd1 (aa142-1274)
Tags6xHis-tag (Precission protease cleavable)ExpressionBacterialMutationdeleted amino acids 1-117 and 1275-1468Available SinceAug. 6, 2024AvailabilityAcademic Institutions and Nonprofits only -
-
-
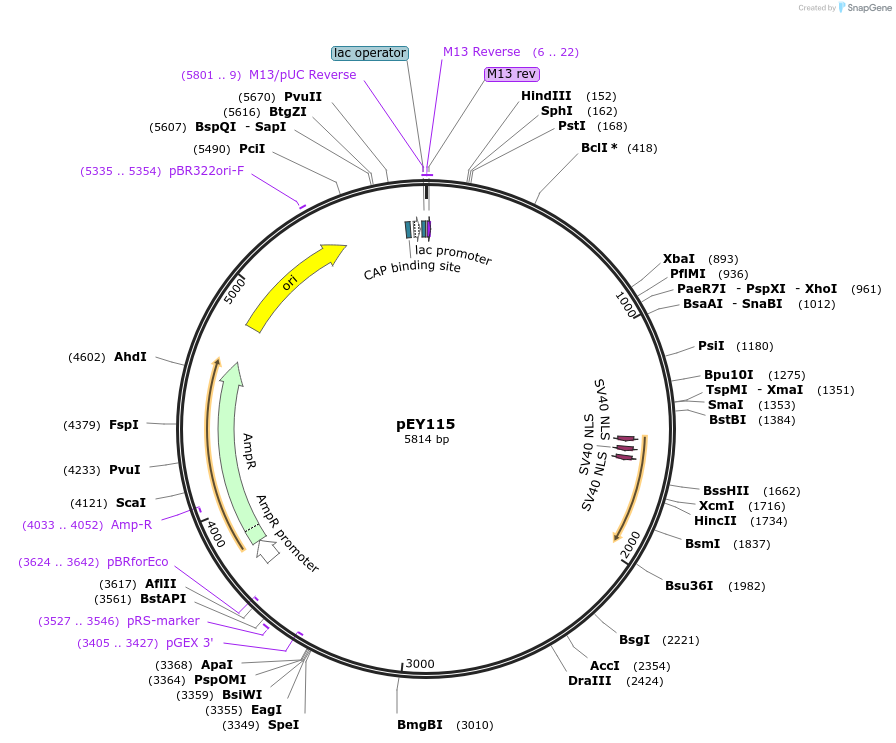
pEY115
Plasmid#223462Purposepdfr-1 (3-2K) fluorescent neural reporter driving nuclear mNeptune2.5 expression (refer to NeuroPAL paper for expression)DepositorInsertpdfr-1(3-2K) (pdfr-1 Nematode)
ExpressionWormAvailable SinceAug. 6, 2024AvailabilityAcademic Institutions and Nonprofits only -
-
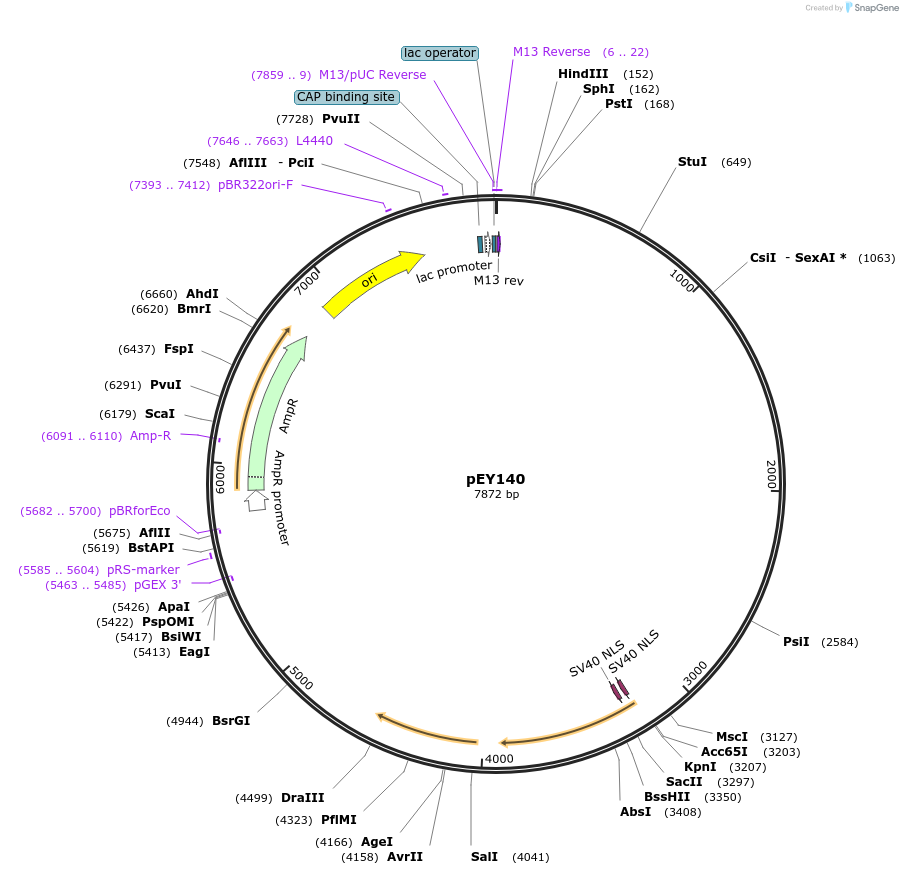
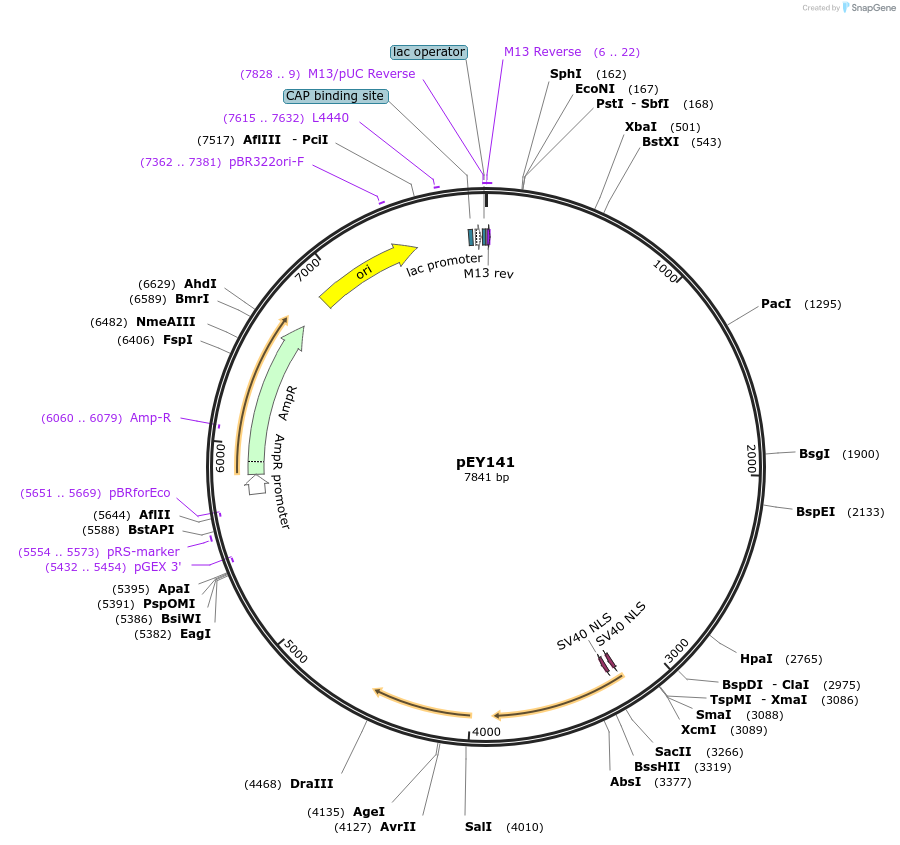
pEY141
Plasmid#223488Purposetrp-4(intron 1) fluorescent neural reporter driving nuclear mCyOFP1 expression (refer to NeuroPAL paper for expression)DepositorInserttrp-4(intron 1) (trp-4 Nematode)
ExpressionWormAvailable SinceAug. 6, 2024AvailabilityAcademic Institutions and Nonprofits only -
-
-
-
-
-
-
-
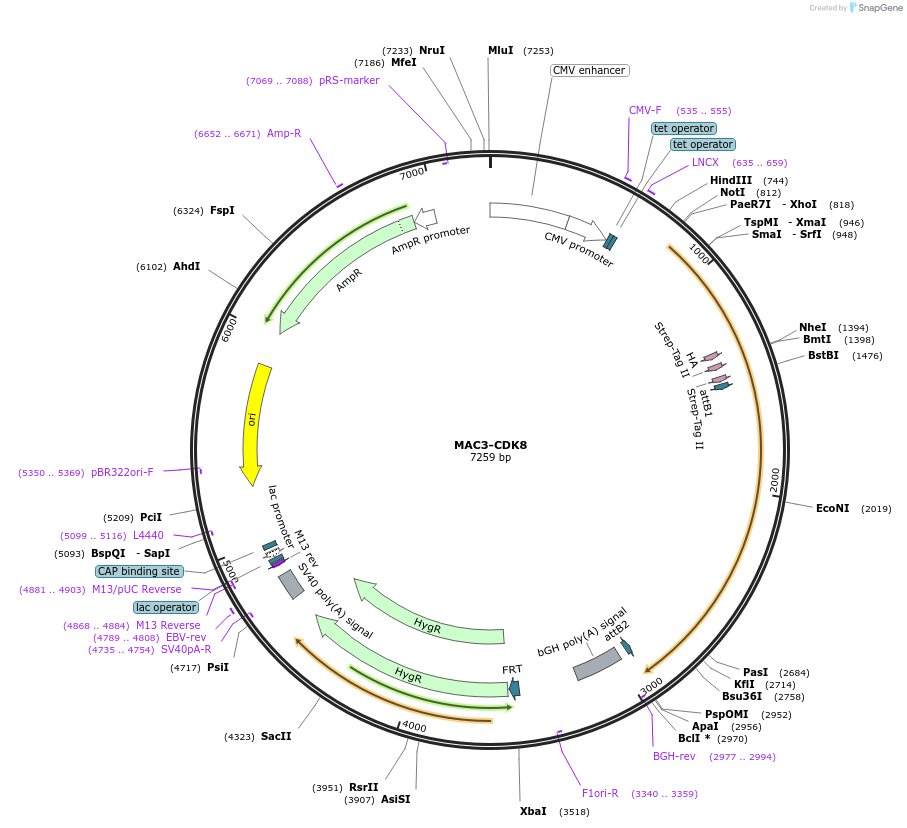
MAC3-CDK8
Plasmid#222609PurposeExpression of MAC3-tagged CDK8DepositorAvailable SinceAug. 6, 2024AvailabilityIndustry, Academic Institutions, and Nonprofits -
-
-
-
-
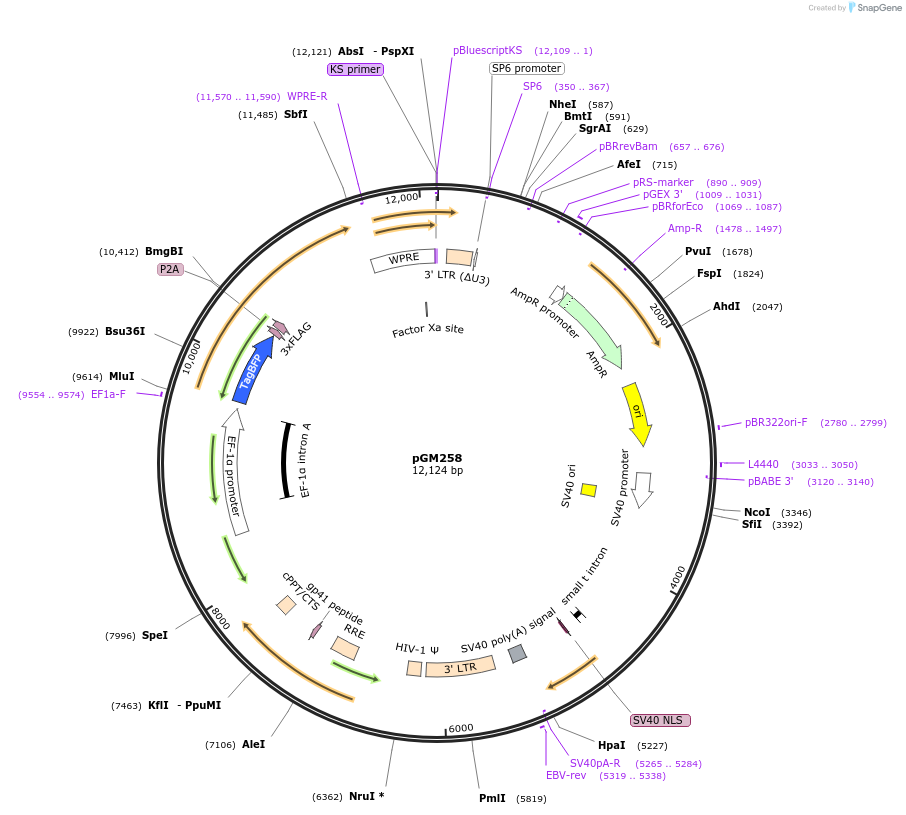
pGM258
Plasmid#220176PurposeExpresses exogenous C-terminal mutant TTC1 (I271D)DepositorAvailable SinceJune 7, 2024AvailabilityAcademic Institutions and Nonprofits only -
-
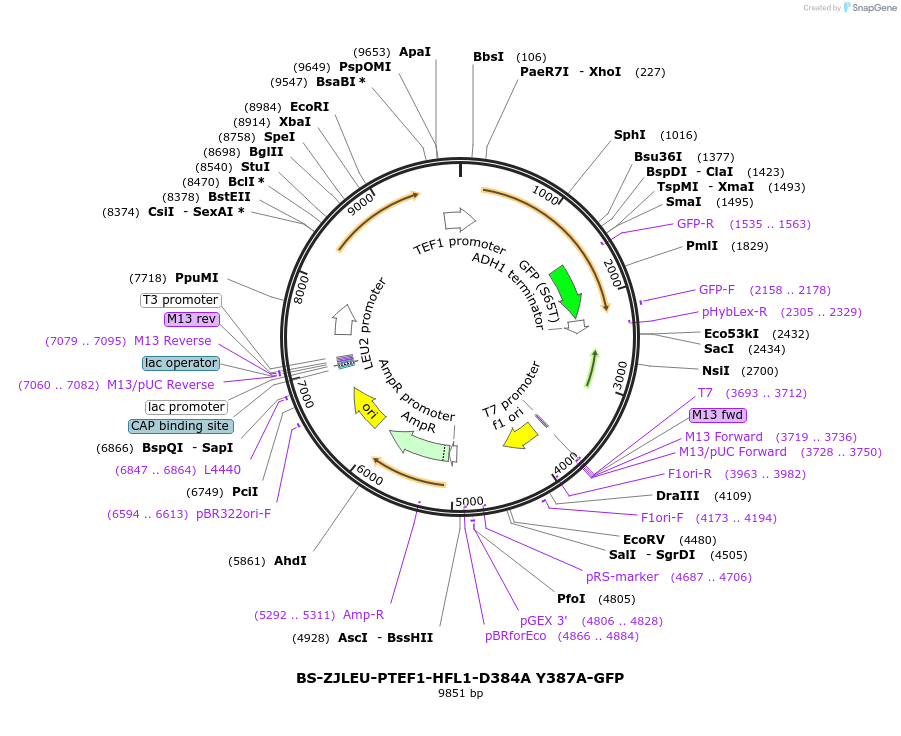
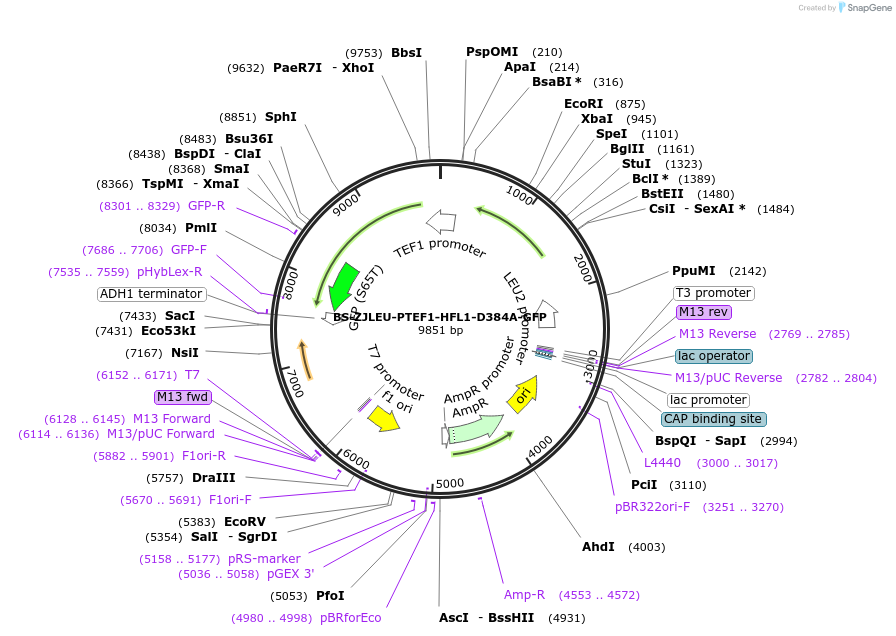
BS-ZJLEU-PTEF1-HFL1-D384A Y387A-GFP
Plasmid#207024PurposeExpression of Hfl1 point mutant. Uses auxotrophic marker LEU2 (Saccharomyces cerevisiae).DepositorInsertHFL1
TagsGFPExpressionYeastMutationD384A Y387APromoterpTEF1Available SinceJune 4, 2024AvailabilityAcademic Institutions and Nonprofits only -
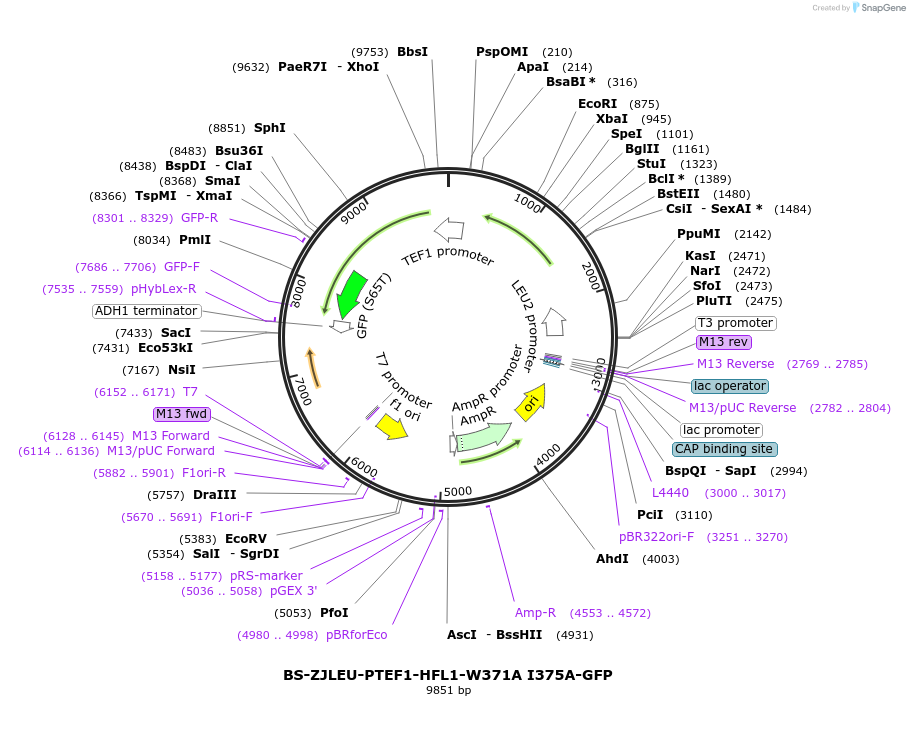
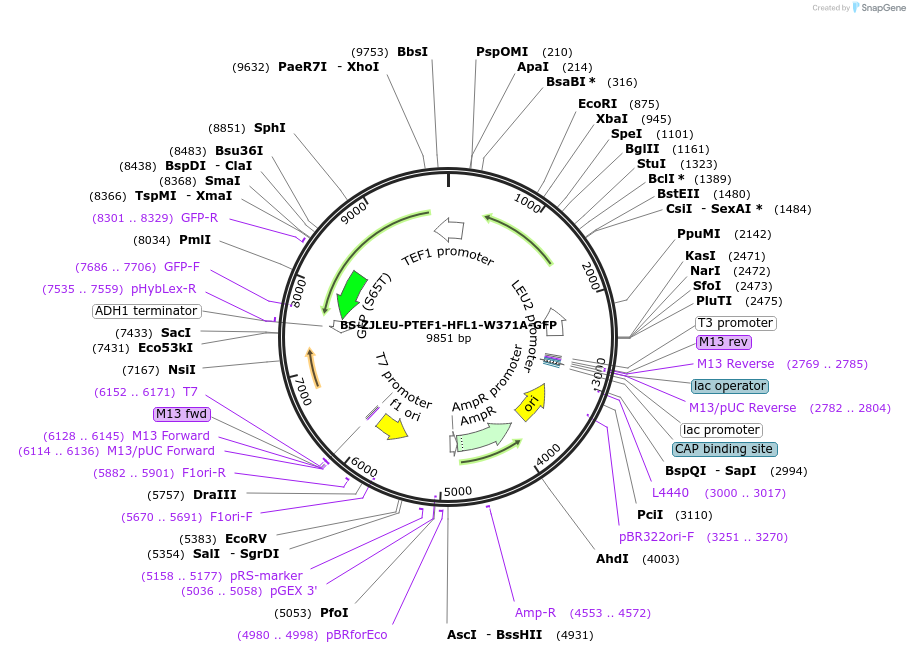
BS-ZJLEU-PTEF1-HFL1-W371A I375A-GFP
Plasmid#207023PurposeExpression of Hfl1 point mutant. Uses auxotrophic marker LEU2 (Saccharomyces cerevisiae).DepositorInsertHFL1
TagsGFPExpressionYeastMutationW371A I375APromoterpTEF1Available SinceJune 4, 2024AvailabilityAcademic Institutions and Nonprofits only -
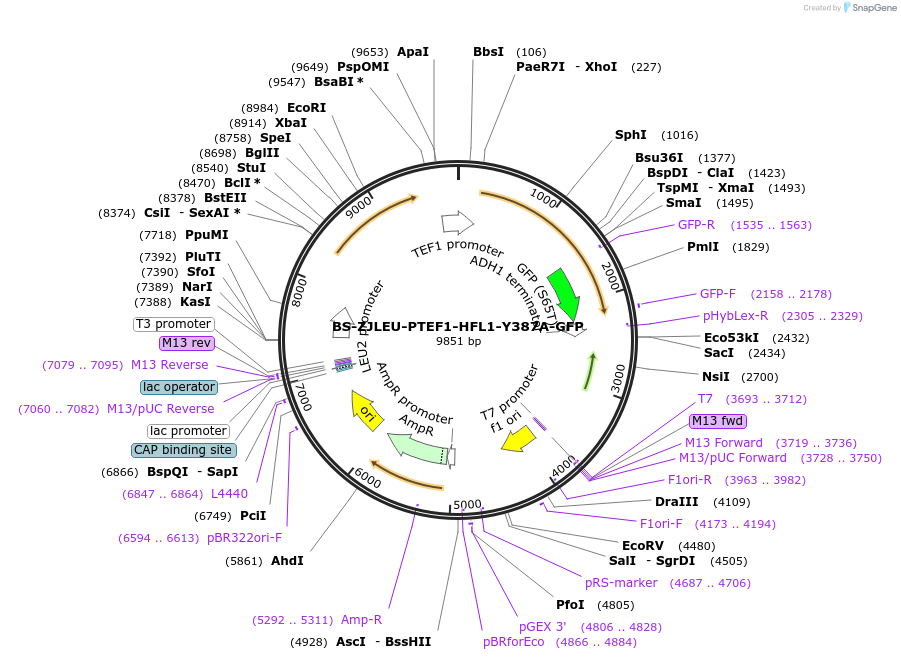
BS-ZJLEU-PTEF1-HFL1-Y387A-GFP
Plasmid#207022PurposeExpression of Hfl1 point mutant. Uses auxotrophic marker LEU2 (Saccharomyces cerevisiae).DepositorInsertHFL1
TagsGFPExpressionYeastMutationY387APromoterpTEF1Available SinceJune 4, 2024AvailabilityAcademic Institutions and Nonprofits only -
BS-ZJLEU-PTEF1-HFL1-D384A-GFP
Plasmid#207021PurposeExpression of Hfl1 point mutant. Uses auxotrophic marker LEU2 (Saccharomyces cerevisiae).DepositorInsertHFL1
TagsGFPExpressionYeastMutationD384APromoterpTEF1Available SinceJune 4, 2024AvailabilityAcademic Institutions and Nonprofits only -
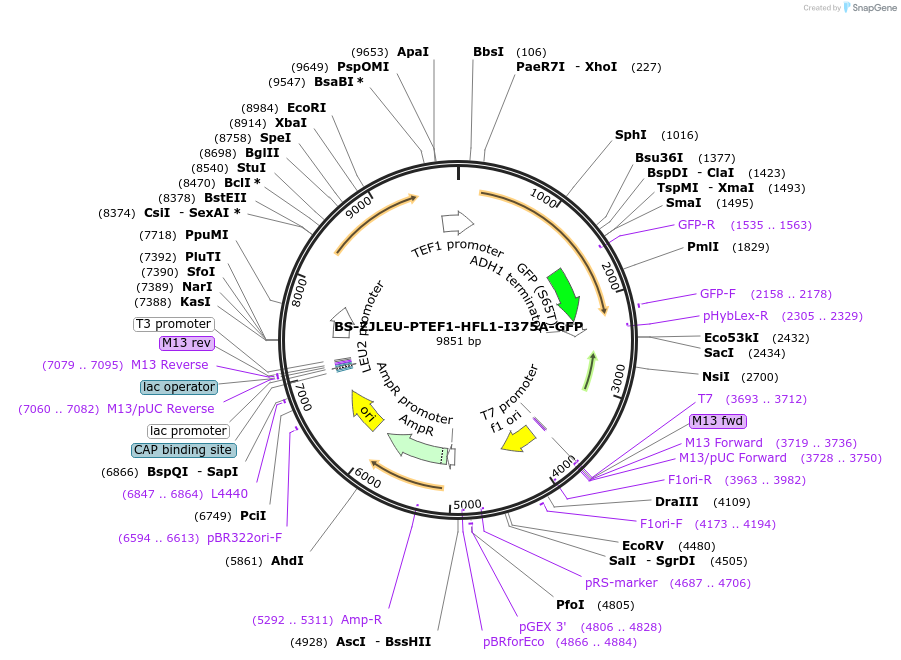
BS-ZJLEU-PTEF1-HFL1-I375A-GFP
Plasmid#207020PurposeExpression of Hfl1 point mutant. Uses auxotrophic marker LEU2 (Saccharomyces cerevisiae).DepositorInsertHFL1
TagsGFPExpressionYeastMutationI375APromoterpTEF1Available SinceJune 4, 2024AvailabilityAcademic Institutions and Nonprofits only -
BS-ZJLEU-PTEF1-HFL1-W371A-GFP
Plasmid#207019PurposeExpression of Hfl1 point mutant. Uses auxotrophic marker LEU2 (Saccharomyces cerevisiae).DepositorInsertHFL1
TagsGFPExpressionYeastMutationW371APromoterpTEF1Available SinceJune 4, 2024AvailabilityAcademic Institutions and Nonprofits only -
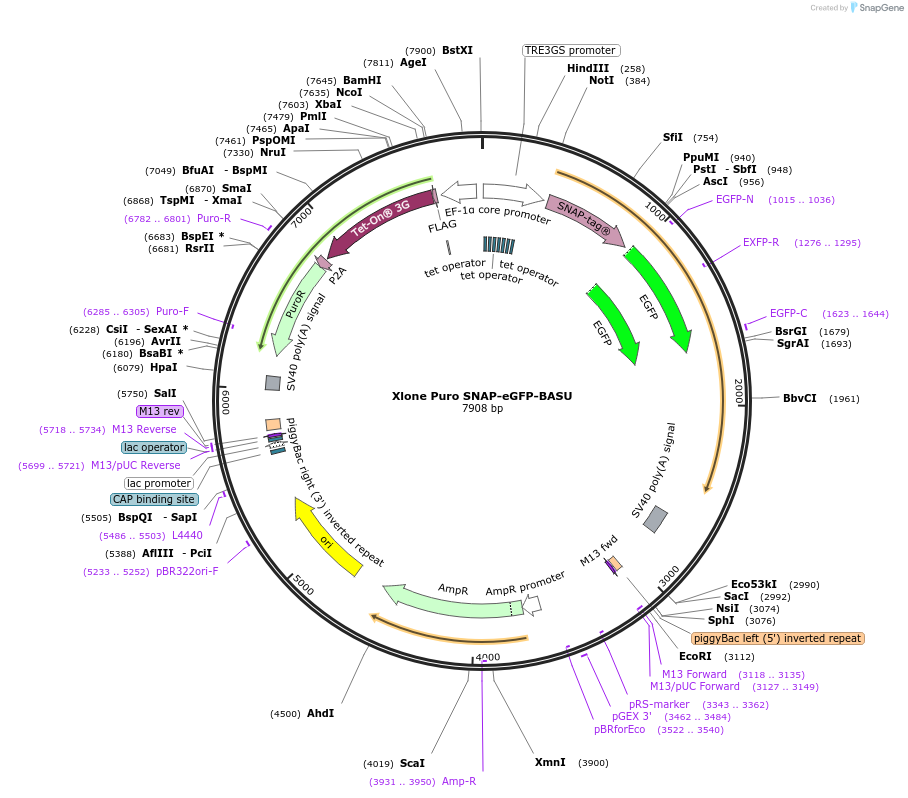
Xlone Puro SNAP-eGFP-BASU
Plasmid#218452Purposeexpresses the SNAP-eGFP-BASU fusion protein via stable PiggyBac integrationDepositorInsertSNAP-eGFP-BASU
ExpressionMammalianPromoterTRE3GSAvailable SinceJune 4, 2024AvailabilityAcademic Institutions and Nonprofits only