We narrowed to 83,435 results for: myc
-
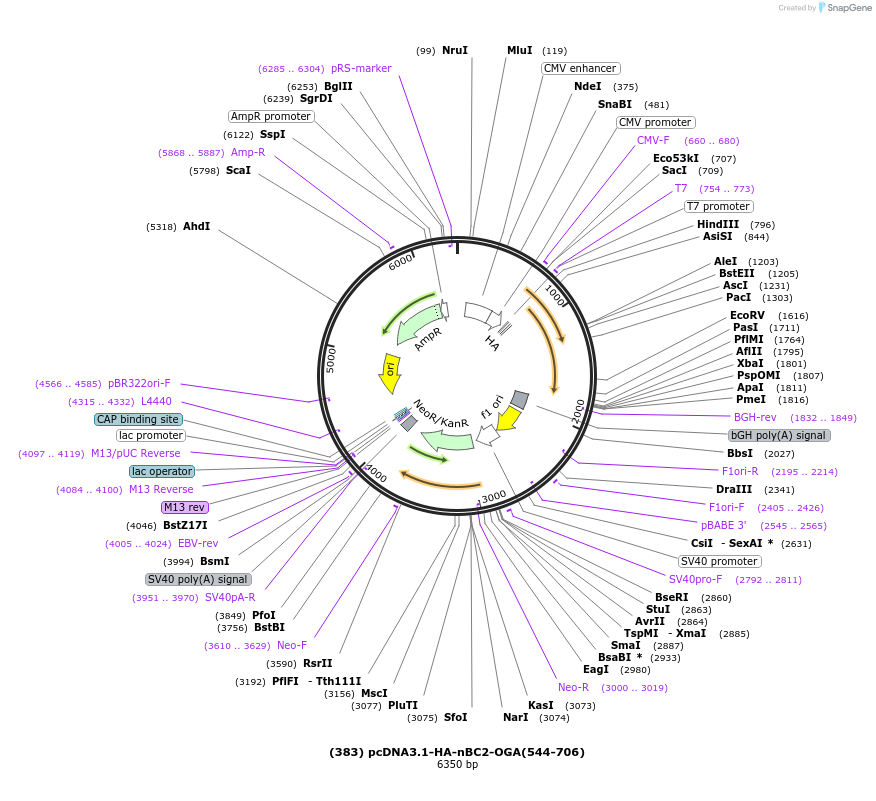
Plasmid#168100PurposeHA- tagged BC2 nanobody fused to human OGA stalk domain residues (544-706). Used in combination with myc-OGA(1-400) to create split-OGADepositorInsertHA-nBC2-OGA(544-706)
TagsHA-tagExpressionMammalianMutationOGA truncated to contain residues 544-706PromoterT7/CMVAvailable SinceMay 24, 2021AvailabilityAcademic Institutions and Nonprofits only -
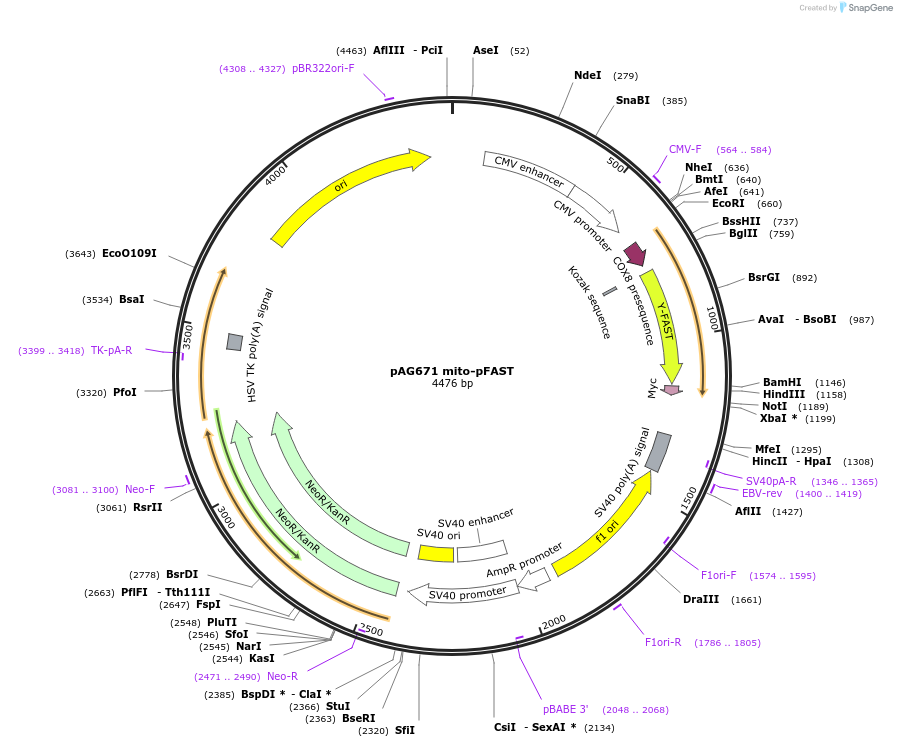
pAG671 mito-pFAST
Plasmid#172867PurposeExpresses mitochondrial targeting domain-pFAST in mammalian cellsDepositorInsertmito-pFAST
TagsMyc tag (C terminal on insert)ExpressionMammalianPromoterCMVAvailable SinceFeb. 3, 2022AvailabilityAcademic Institutions and Nonprofits only -
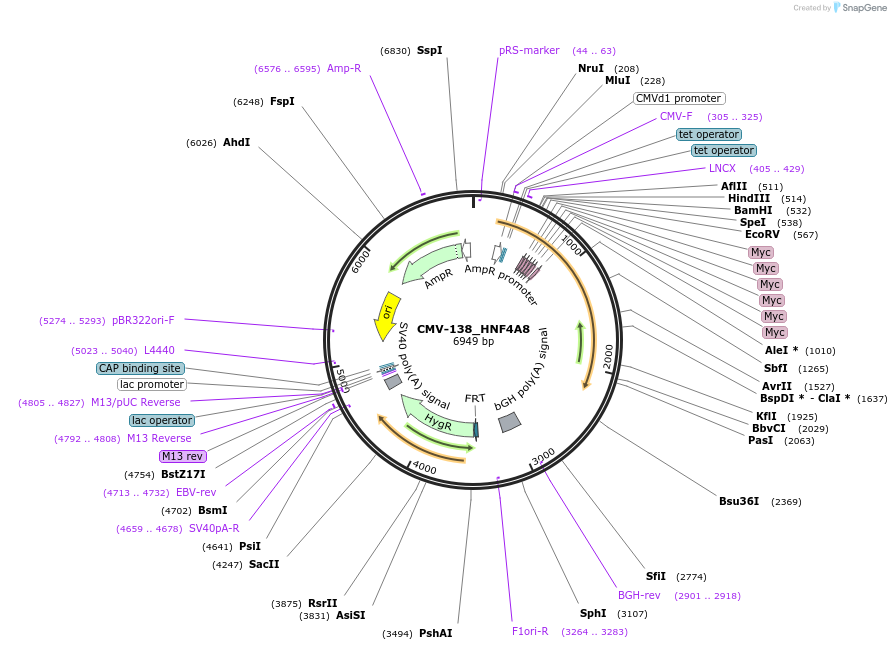
CMV-138_HNF4A8
Plasmid#31093DepositorInsertHNF4A (HNF4A Human)
UseFlp/frtTagsmycExpressionMammalianMutationsplice variant 8 of HNF4A, deletion of CMV promo…Available SinceOct. 14, 2011AvailabilityAcademic Institutions and Nonprofits only -
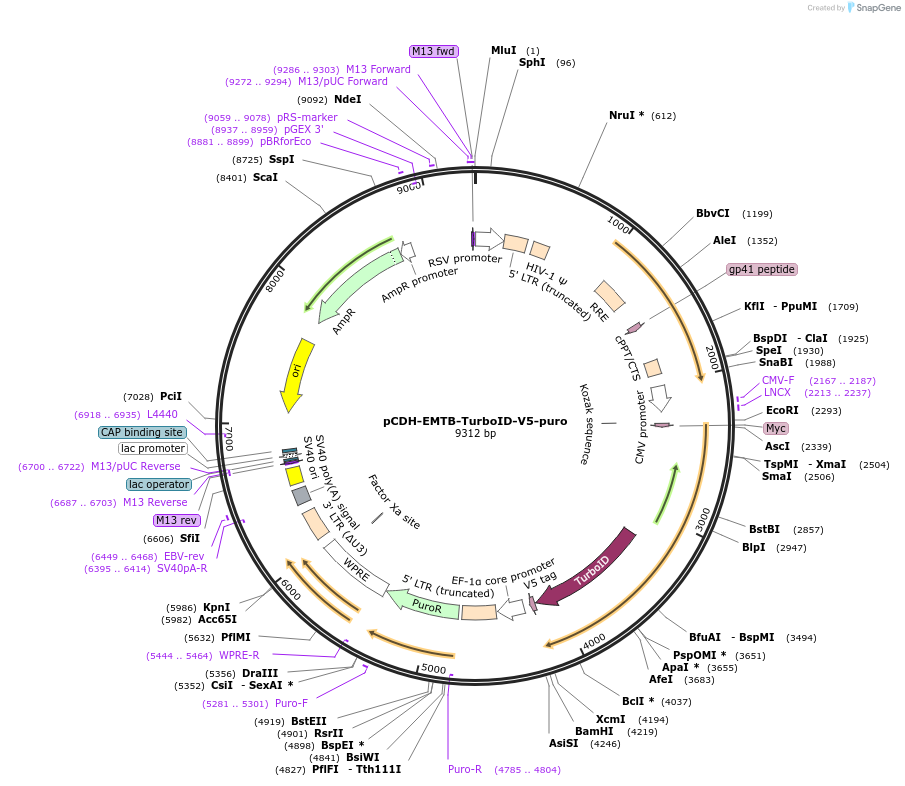
pCDH-EMTB-TurboID-V5-puro
Plasmid#190741PurposeA lentiviral plasmid encoding EMTB fusion to V5-tagged TurboID on microtubules (EMTB as the microtubule-targeting tag)DepositorInsertEMTB-TurboID-V5 (RPS14 EMTB is human ensconsin; TurboID is engineered BirA from E.coli, Human)
UseLentiviralTagsMycExpressionMammalianMutationMicrotubule binding-domain of ensconsin (amino ac…PromoterCMVAvailable SinceFeb. 2, 2023AvailabilityAcademic Institutions and Nonprofits only -
pcDNA3.1-HA-14-3-3 ϵ
Plasmid#48797Purposeexpresses human myc-14-3-3ϵ in mammalian cellsDepositorAvailable SinceJune 4, 2014AvailabilityAcademic Institutions and Nonprofits only -
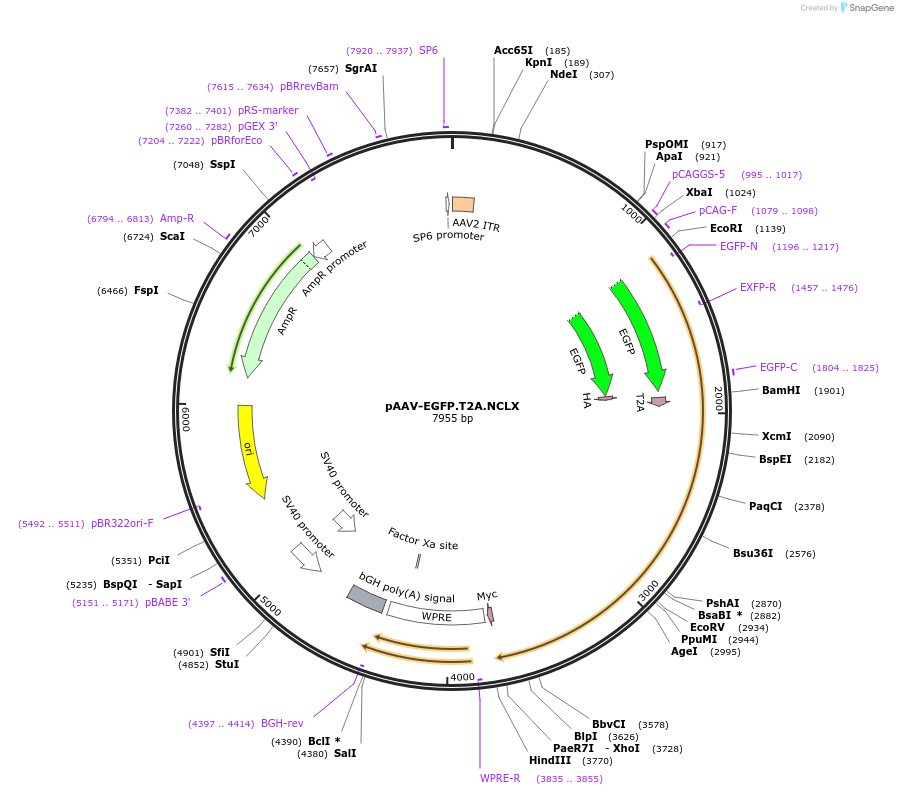
pAAV-EGFP.T2A.NCLX
Plasmid#181873PurposeExpresses EGFP and murine NCLX (separated by a T2A cleavage site) under control of the synthetic CAG promoterDepositorInsertsUseAAV and AdenoviralTagsHA and mycExpressionMammalianPromotersynthetic hybrid CAG promoter and synthetic hybri…Available SinceMarch 4, 2022AvailabilityAcademic Institutions and Nonprofits only -
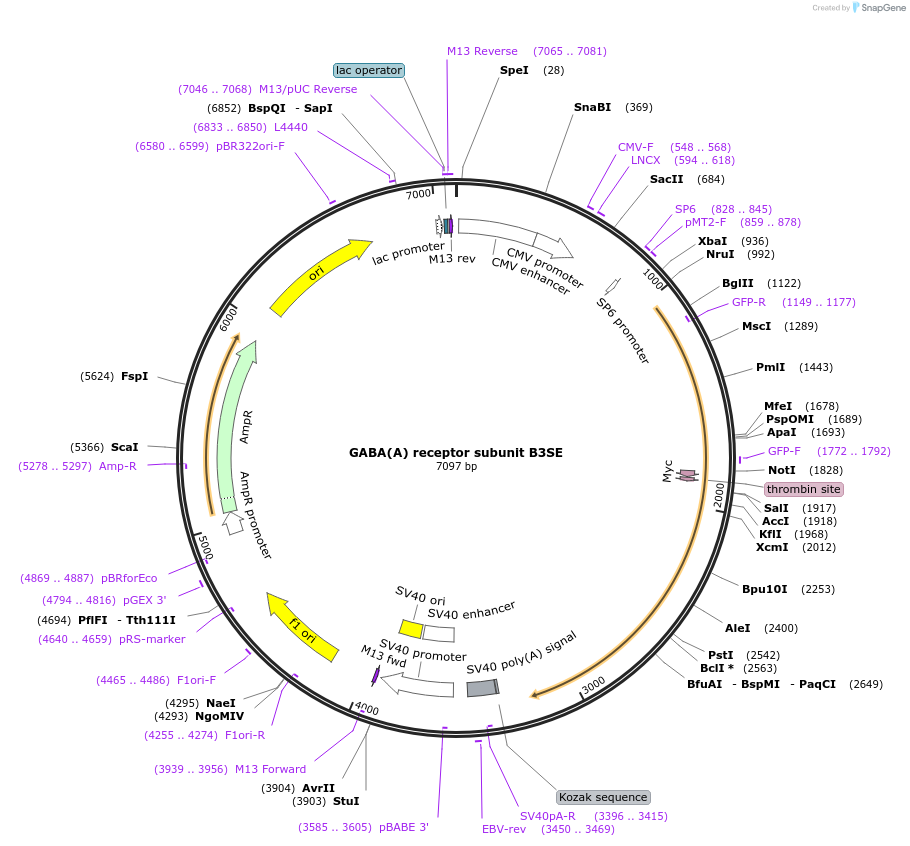
GABA(A) receptor subunit B3SE
Plasmid#49171PurposepHluorin-tagged GABA A receptor subunit (beta 3) produces minimal fluorescence during trafficking and a robust fluorescent signal at the cell surfaceDepositorInsertGABA(A) receptor subunit beta 3 (Gabrb3 Mouse)
Tagsmyc, pHGFP (pHluorin), and thrombin cleavage site…ExpressionMammalianMutationGTG (V) to TGC (C)PromoterCMVAvailable SinceDec. 10, 2013AvailabilityAcademic Institutions and Nonprofits only -
pESC-LEU-HsAIDSc
Plasmid#60810Purposeto express human AID recoded for yeast codon usageDepositorAvailable SinceDec. 19, 2014AvailabilityAcademic Institutions and Nonprofits only -
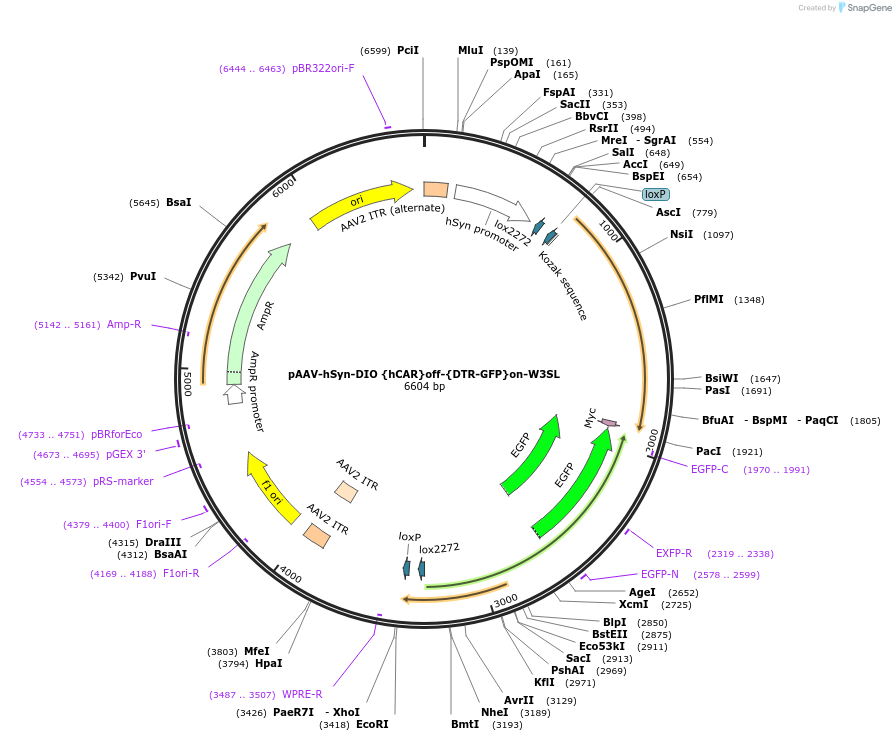
pAAV-hSyn-DIO {hCAR}off-{DTR-GFP}on-W3SL
Plasmid#111396PurposeAAV vector with hSynapsin promoter, Cre-OFF hCAR (for efficient CAV-2 infection), Cre-ON DTR-EGFP (diphtheria toxin receptor for cell ablation), and W3SL cassette (for maximize cloning capacity)DepositorInsertsUseAAVTagsEGFP and MycExpressionMammalianPromoterhSynapsinAvailable SinceJune 7, 2018AvailabilityAcademic Institutions and Nonprofits only -
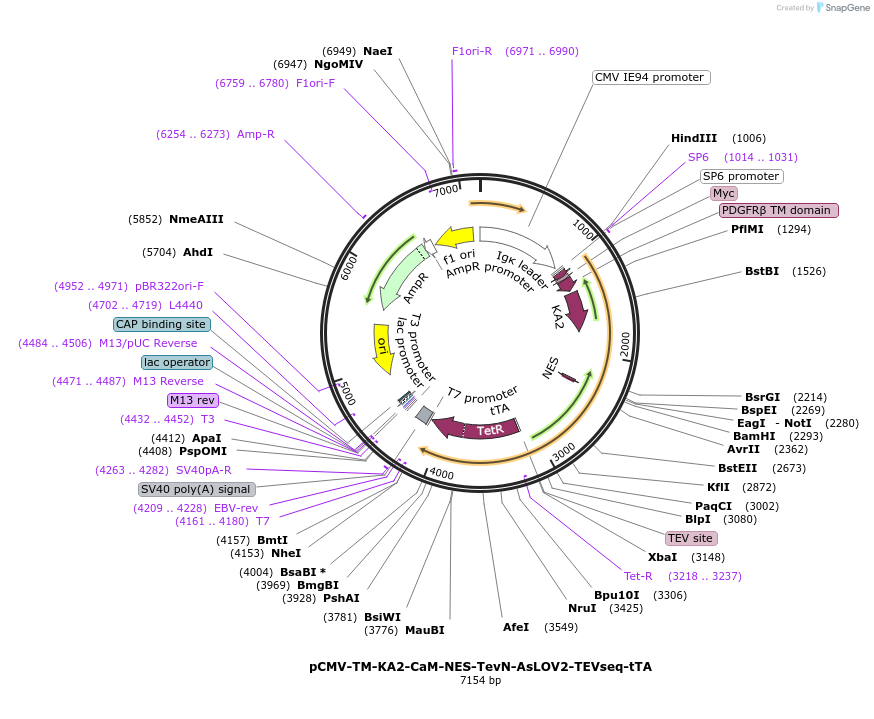
pCMV-TM-KA2-CaM-NES-TevN-AsLOV2-TEVseq-tTA
Plasmid#171615PurposeSoma-targeted Cal-Light with KA2DepositorInsertTM-KA2-CaM-NES-TevN-AsLOV2-TEVseq-tTA
UseSynthetic BiologyTagsMycExpressionMammalianPromoterCMVAvailable SinceOct. 15, 2021AvailabilityAcademic Institutions and Nonprofits only -
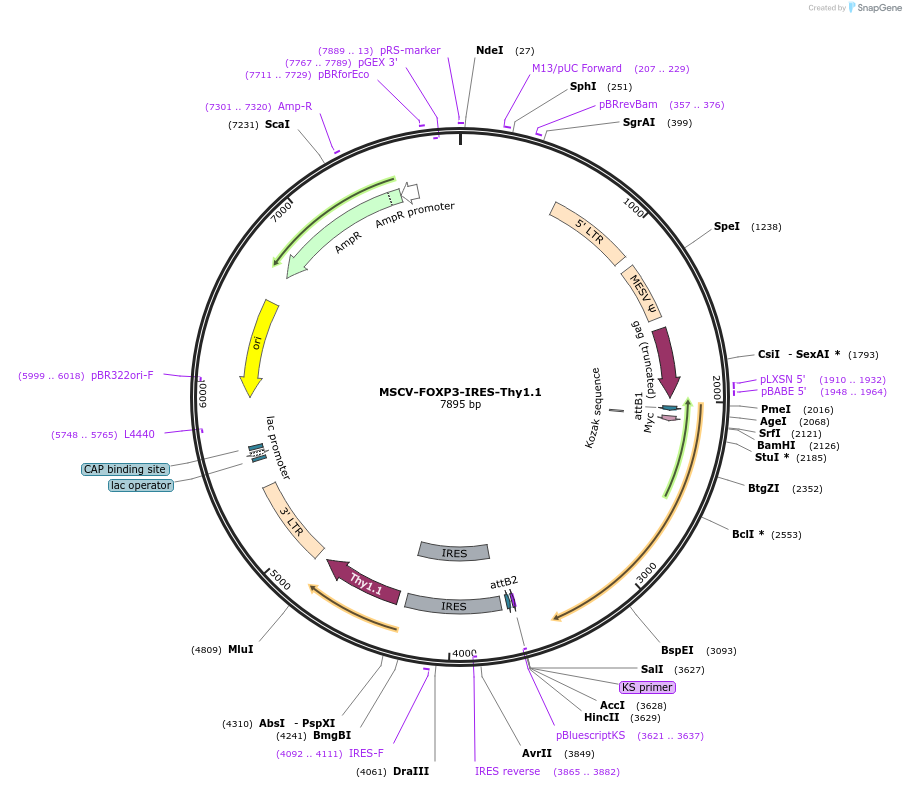
MSCV-FOXP3-IRES-Thy1.1
Plasmid#17443DepositorAvailable SinceMarch 12, 2008AvailabilityAcademic Institutions and Nonprofits only -
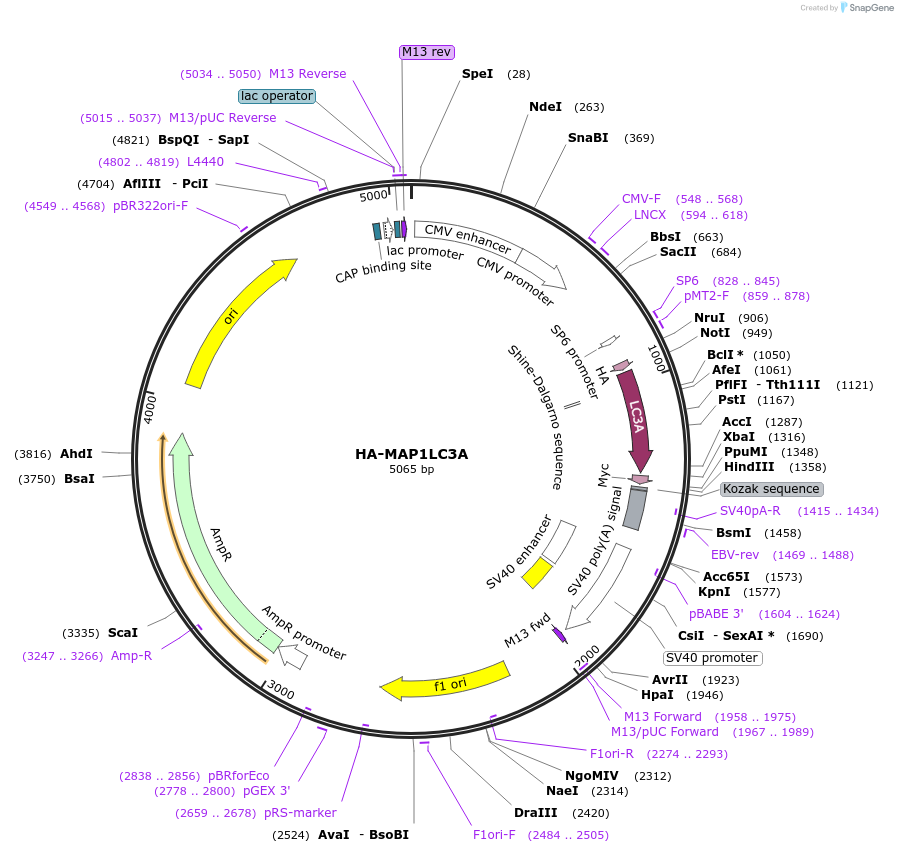
HA-MAP1LC3A
Plasmid#137756PurposeExpression of LC3ADepositorAvailable SinceFeb. 26, 2020AvailabilityAcademic Institutions and Nonprofits only -
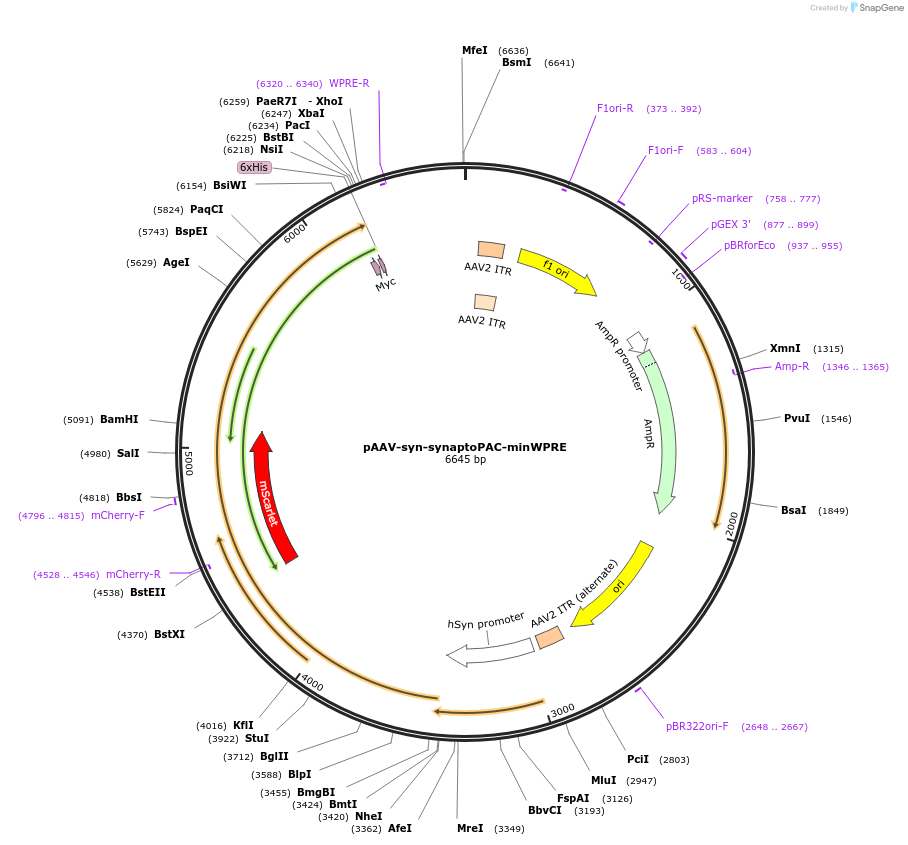
pAAV-syn-synaptoPAC-minWPRE
Plasmid#153085PurposeExpresses synaptoPAC, a fusion of synaptophysin, mScarlet, and bPAC in neuronsDepositorInsertsUseAAVTagsHis tag, mScarlet, and myc tagExpressionMammalianAvailable SinceSept. 8, 2020AvailabilityAcademic Institutions and Nonprofits only -
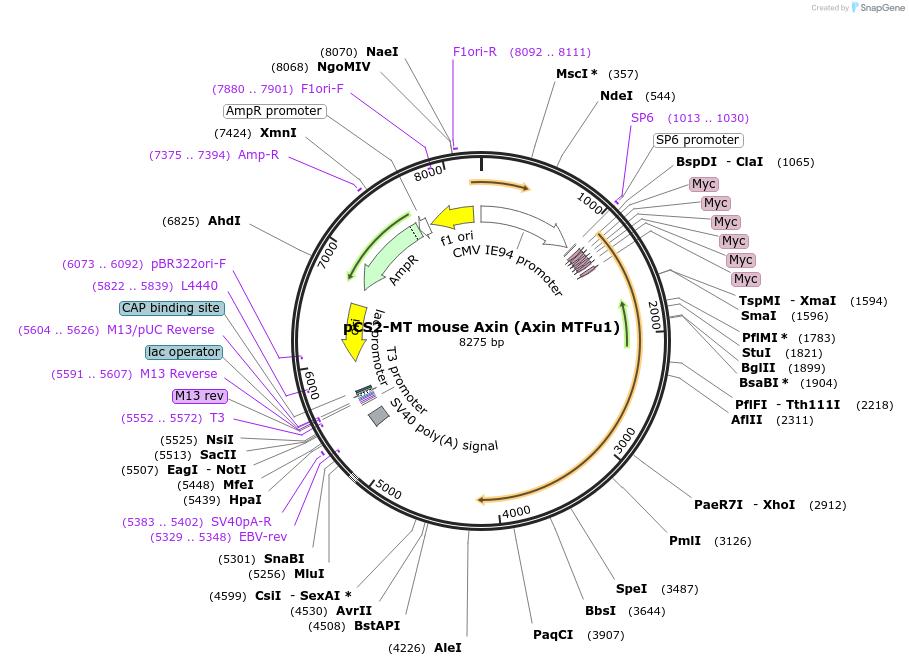
pCS2-MT mouse Axin (Axin MTFu1)
Plasmid#21287DepositorAvailable SinceDec. 15, 2009AvailabilityAcademic Institutions and Nonprofits only -
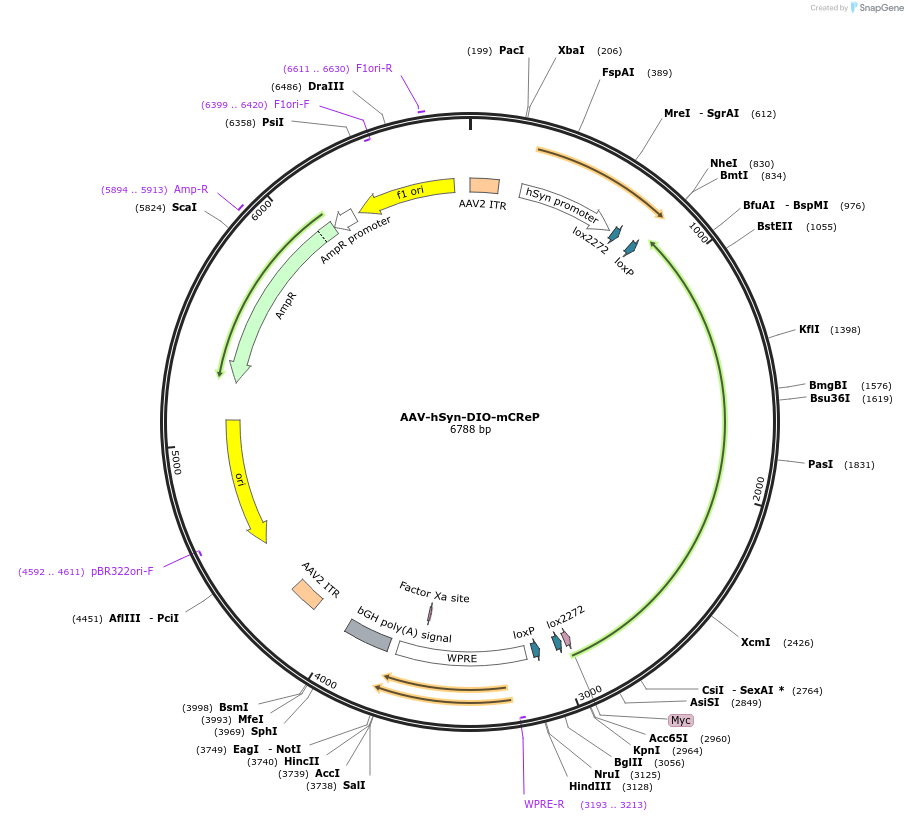
AAV-hSyn-DIO-mCReP
Plasmid#167503PurposeConditional expression of the eIF2alpha phophatase, CReP (Ppp1r15b)DepositorAvailable SinceJune 7, 2021AvailabilityAcademic Institutions and Nonprofits only -
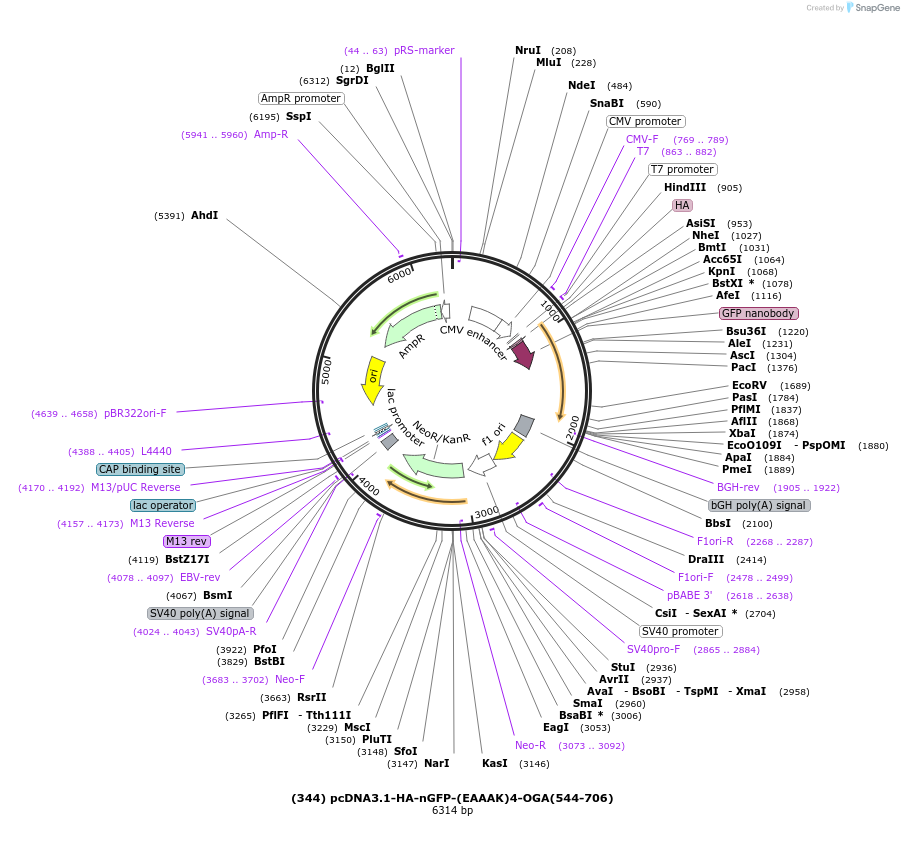
(344) pcDNA3.1-HA-nGFP-(EAAAK)4-OGA(544-706)
Plasmid#168097PurposeHA- tagged GFP nanobody fused to human OGA stalk domain residues (544-706). Used in combination with myc-OGA(1-400) to create split-OGA.DepositorInsertHA-nGFP-(EAAAK)4-OGA(544-706)
TagsHA-TagExpressionMammalianPromoterT7/CMVAvailable SinceMay 18, 2021AvailabilityAcademic Institutions and Nonprofits only -
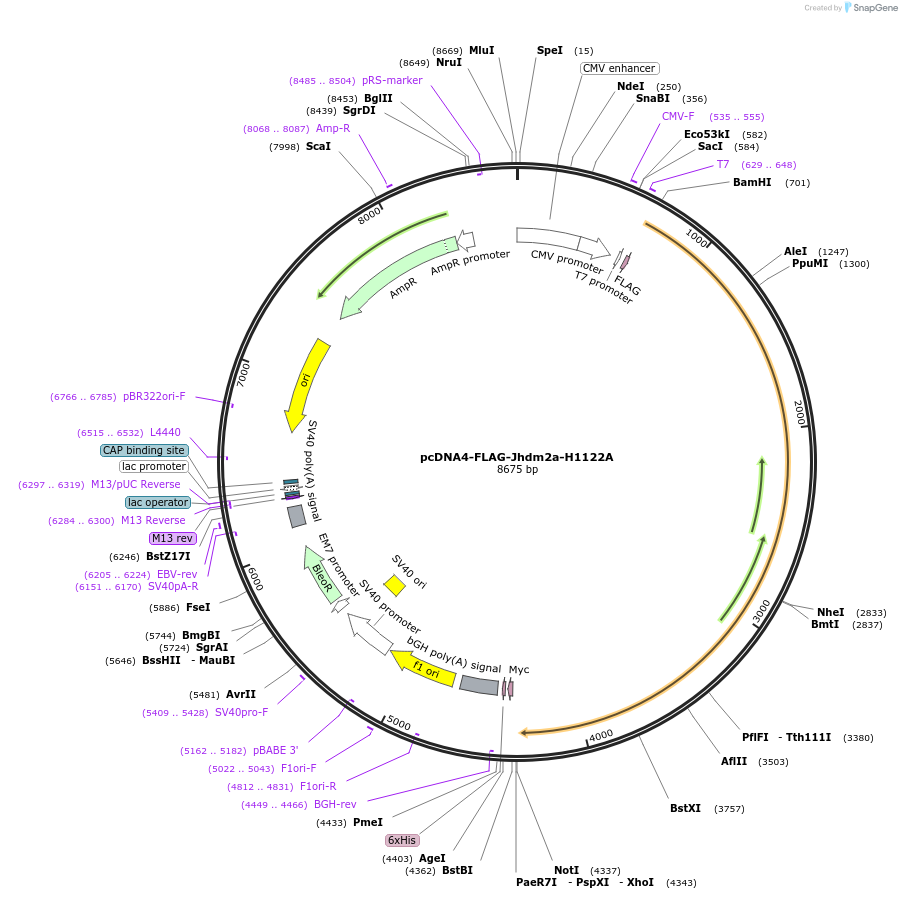
pcDNA4-FLAG-Jhdm2a-H1122A
Plasmid#38140DepositorInsertJhdm2a-H1122A (Kdm3a Mouse)
TagsFLAGExpressionMammalianMutationprotein starts at aa115; H1122APromoterCMVAvailable SinceSept. 28, 2012AvailabilityAcademic Institutions and Nonprofits only -
pAAV-hSyn-DIO {hCAR}off-{GCaMP6f}on-W3SL
Plasmid#111394PurposeAAV vector with hSynapsin promoter, Cre-OFF hCAR (for efficient CAV-2 infection), Cre-ON GCaMP6f (ultrasensitive calcium sensor), and W3SL regulatory cassette (for maximize cloning capacity)DepositorInsertsUseAAVTags6xHis, Myc, T7, and XpressExpressionMammalianPromoterhSynapsinAvailable SinceJune 8, 2018AvailabilityAcademic Institutions and Nonprofits only -
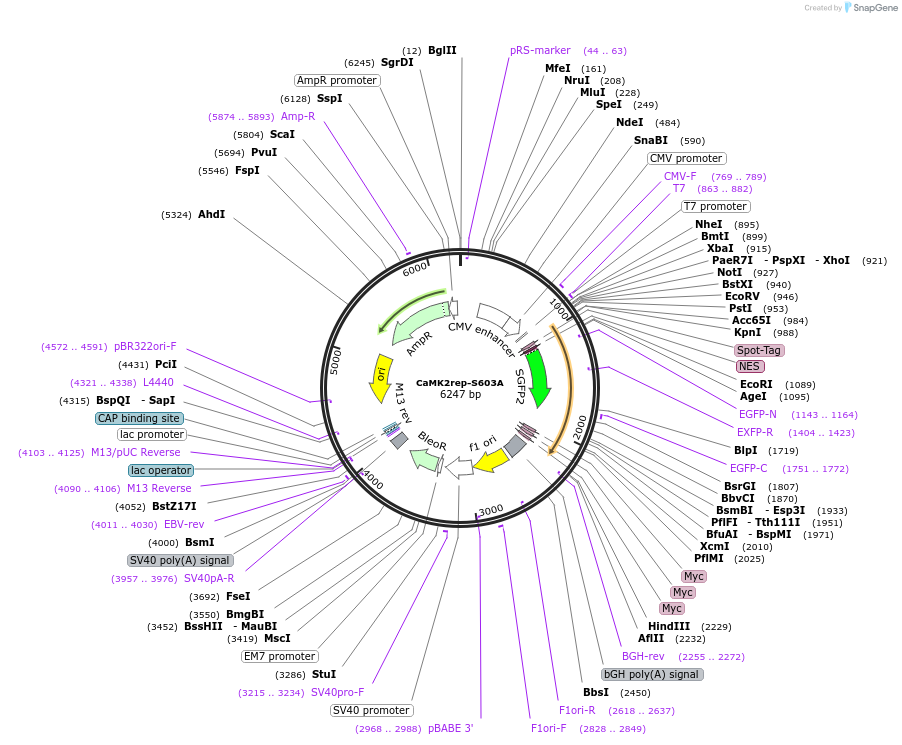
CaMK2rep-S603A
Plasmid#239623PurposeInactive CaMKII activity reporter. The plasmid contains sGFP2 fluorescent protein, 3xmyc tags and double mutated phospho-site 3 from rat Synapsin-1a.DepositorInsertSpotNES-sGFP2-SynP3-3xmyc (Syn1 Synthetic, Rat)
TagsSpot, myc, and sGFP2ExpressionMammalianMutationchanged Serine 603 to AlaninePromoterCMVAvailable SinceJuly 1, 2025AvailabilityAcademic Institutions and Nonprofits only -
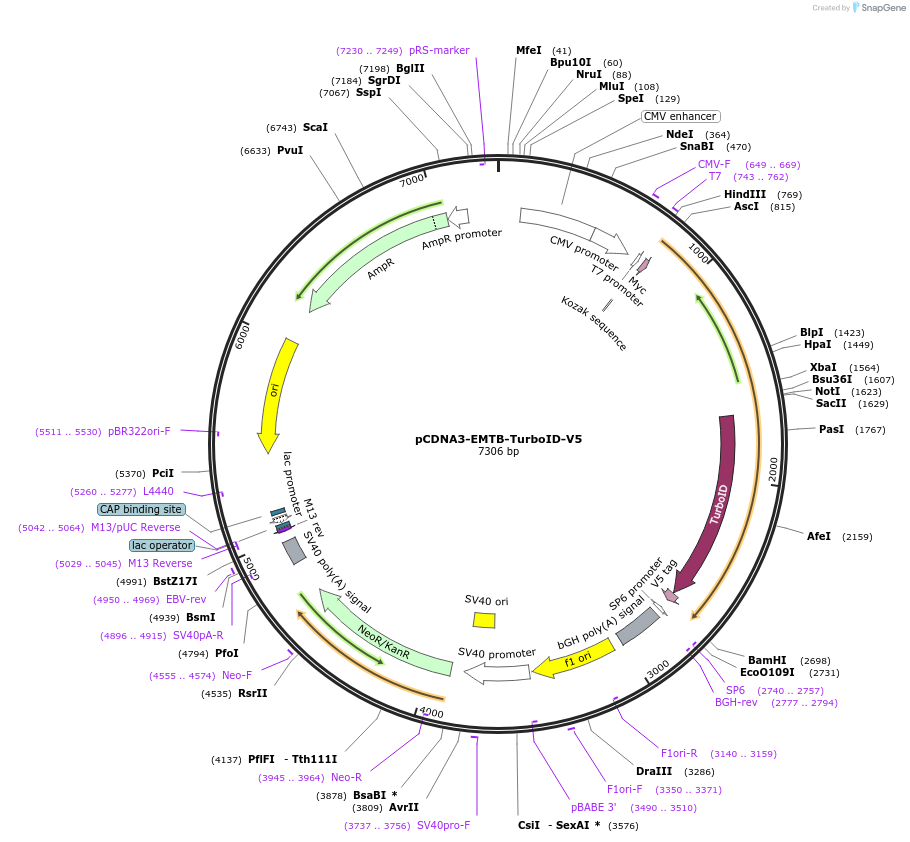
pCDNA3-EMTB-TurboID-V5
Plasmid#190736PurposeMammalian expression of EMTB fusion to V5-tagged TurboID on microtubules (EMTB as the microtubule-targeting tag)DepositorInsertensconsin (MAP7 Human)
TagsMyc, TurboID, and V5ExpressionMammalianMutationMicrotubule binding-domain of ensconsin (amino ac…PromoterCMVAvailable SinceDec. 8, 2022AvailabilityAcademic Institutions and Nonprofits only