We narrowed to 83,312 results for: MYC
-
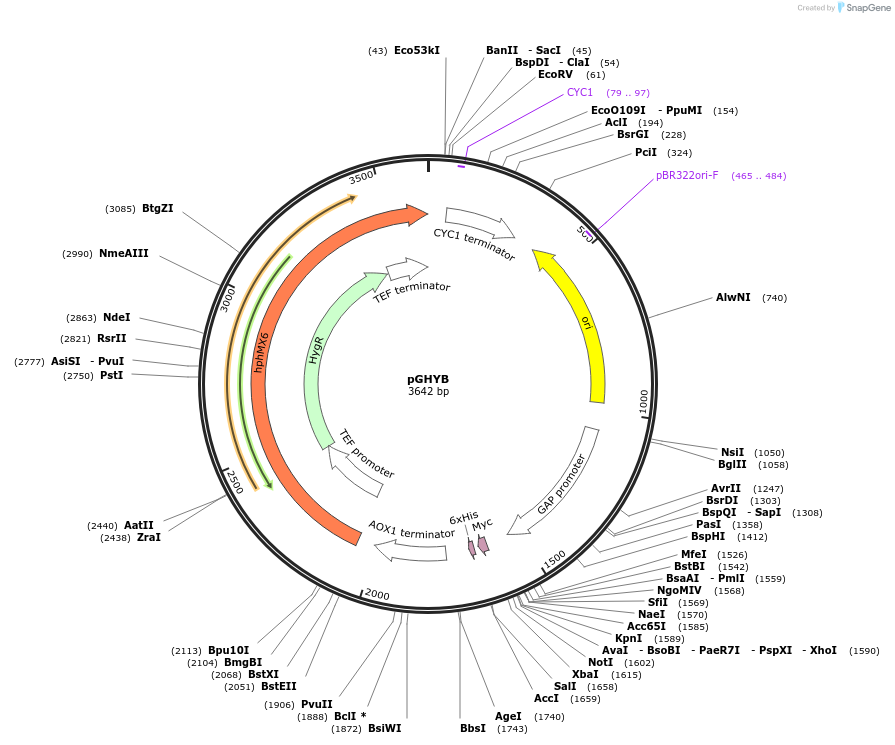
Plasmid#87447PurposePichia pastoris expression plasmidDepositorTypeEmpty backboneExpressionYeastPromoterGAPAvailable SinceMarch 17, 2017AvailabilityAcademic Institutions and Nonprofits only
-
pMN437
Plasmid#32362DepositorInsertgfpm2+
UseMycobacterial gfp expression vectorAvailable SinceNov. 18, 2011AvailabilityAcademic Institutions and Nonprofits only -
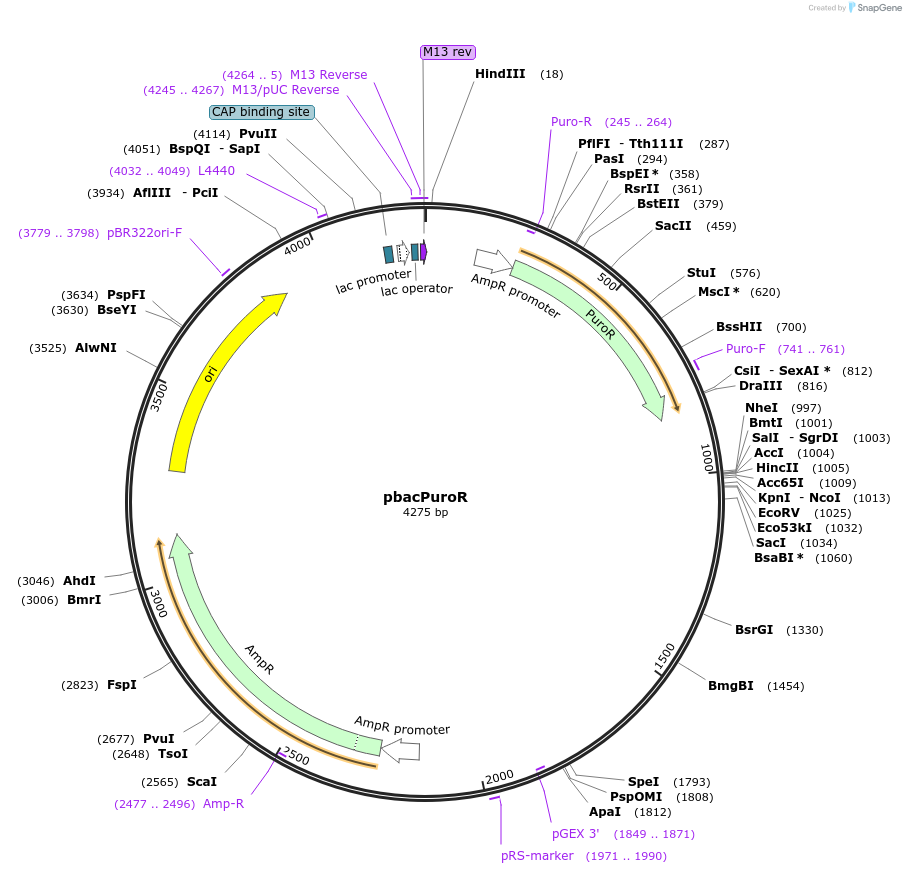
pbacPuroR
Plasmid#26348DepositorInsertBacterial promoter::PuroR::bacterial 3p flank
ExpressionBacterialAvailable SinceDec. 3, 2010AvailabilityAcademic Institutions and Nonprofits only -
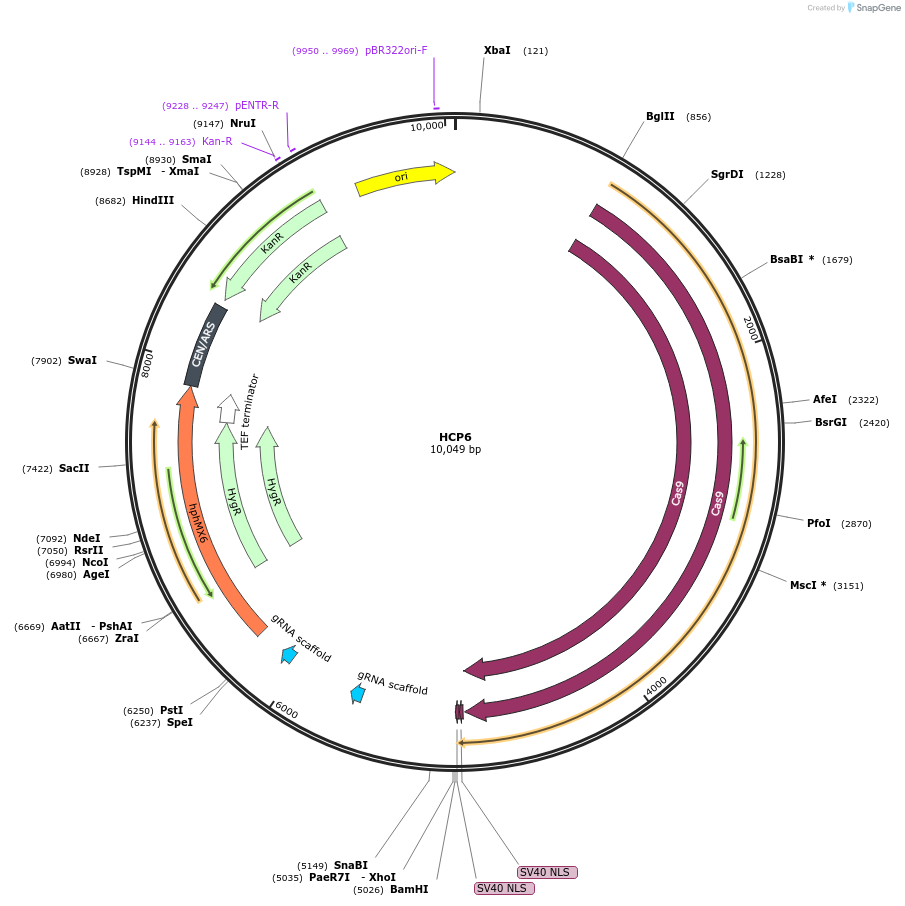
HCP6
Plasmid#166108PurposeThis plasmid encodes a Cas9 protein as well as two sgRNAs, one targets the C-terminus of Cyr1 and the other targets a non-coding region on chromosome X (location X1)DepositorAvailable SinceApril 13, 2021AvailabilityAcademic Institutions and Nonprofits only -
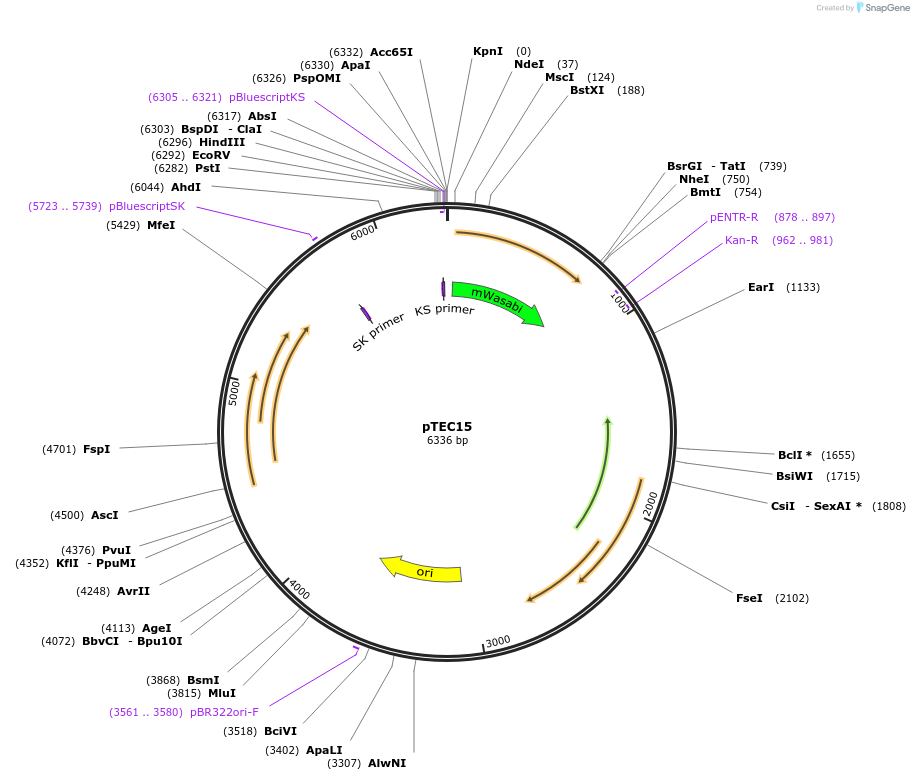
pTEC15
Plasmid#30174DepositorInsertMycobacterium Strong Promoter (MSP)
UseMycobacteria expressionTagsWasabiAvailable SinceJune 30, 2011AvailabilityAcademic Institutions and Nonprofits only -
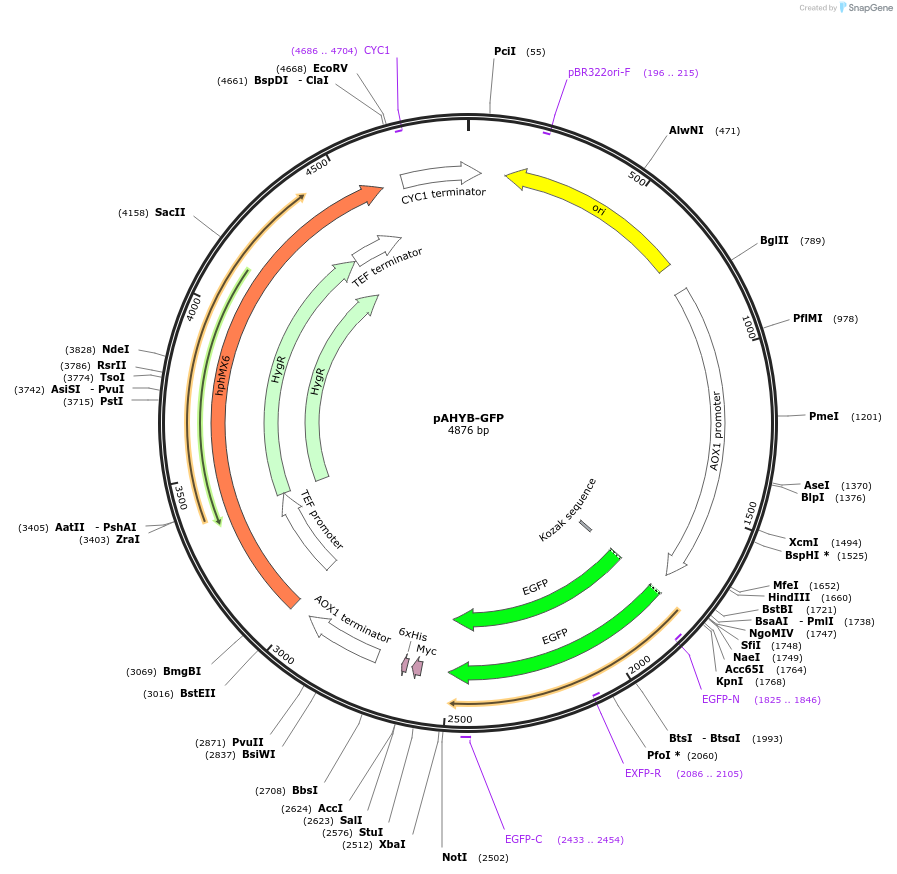
pAHYB-GFP
Plasmid#85074PurposeExpression of GFP in Pichia pastorisDepositorInsertGFP
ExpressionYeastPromoterAOX1Available SinceMarch 17, 2017AvailabilityAcademic Institutions and Nonprofits only -
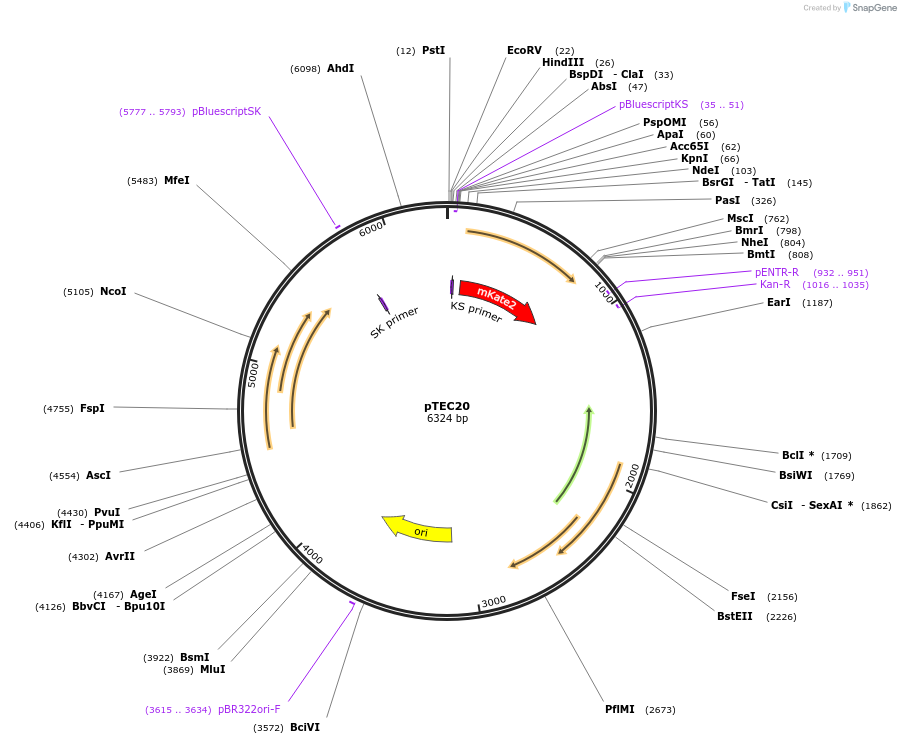
pTEC20
Plasmid#30179DepositorInsertMycobacterium Strong Promoter (MSP)
UseMycobacteria expressionTagsmKate2Available SinceJune 3, 2011AvailabilityAcademic Institutions and Nonprofits only -
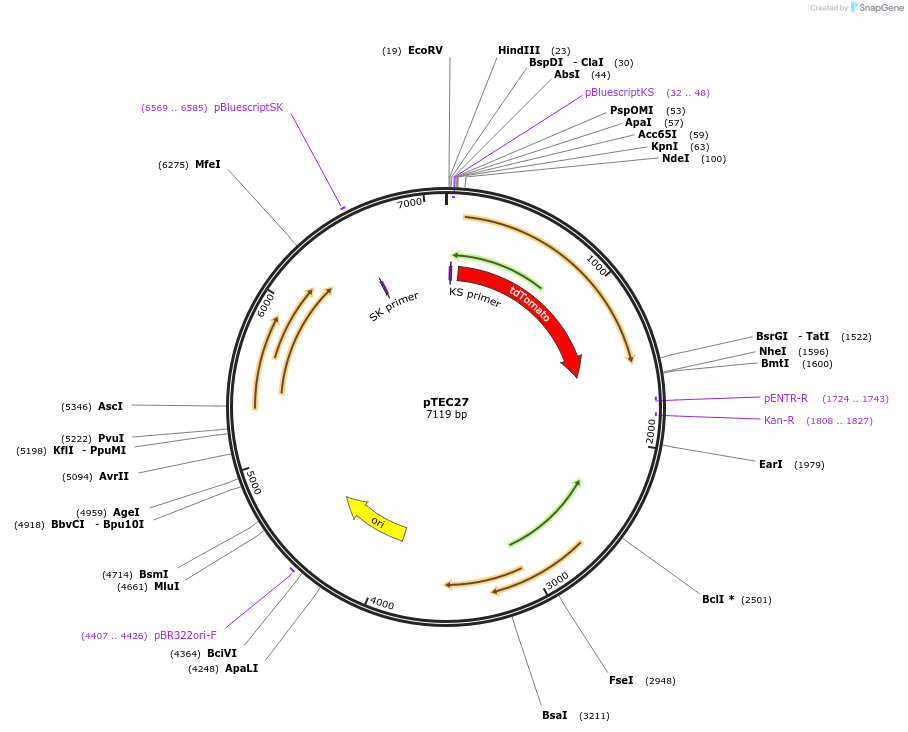
pTEC27
Plasmid#30182DepositorInsertMycobacterium Strong Promoter (MSP)
UseMycobacteria expressionTagstdTomatoAvailable SinceJune 3, 2011AvailabilityAcademic Institutions and Nonprofits only -
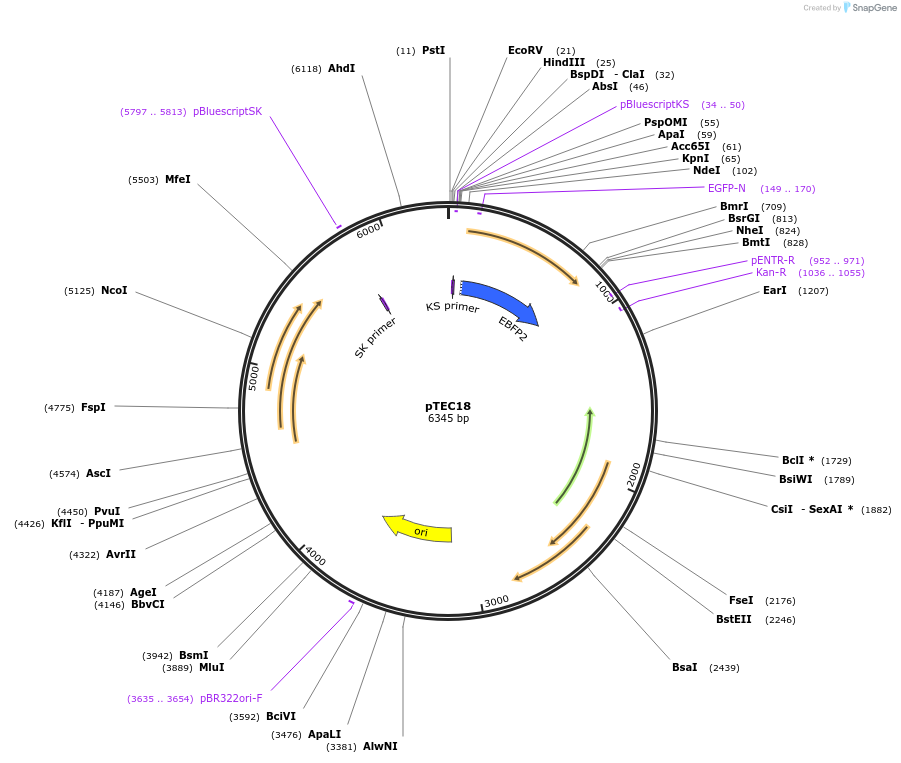
pTEC18
Plasmid#30177DepositorInsertMycobacterium Strong Promoter (MSP)
UseMycobacteria expressionTagsEBFP2Available SinceJune 3, 2011AvailabilityAcademic Institutions and Nonprofits only -
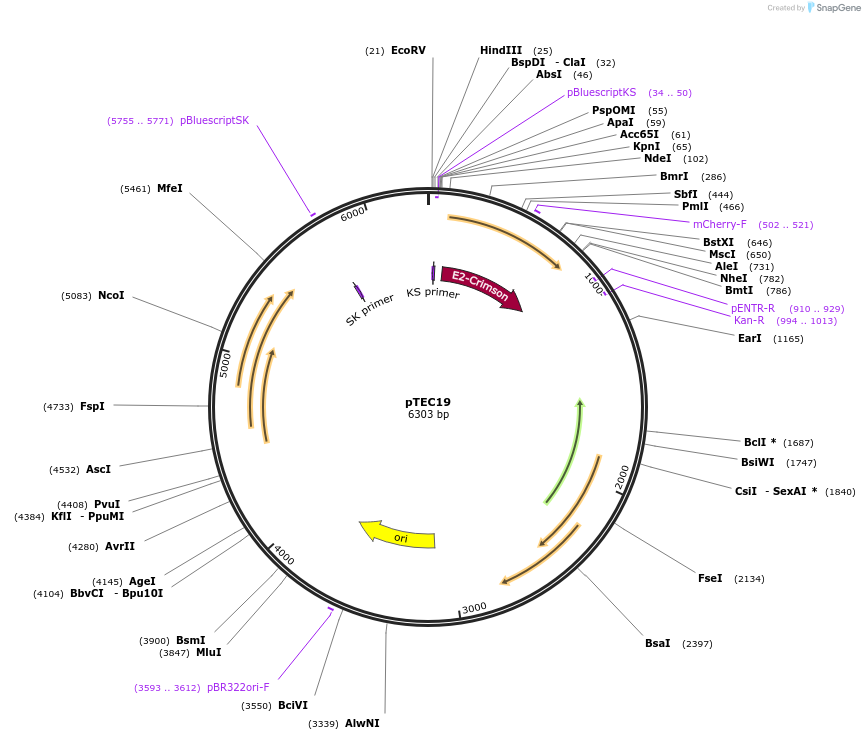
pTEC19
Plasmid#30178DepositorInsertMycobacterium Strong Promoter (MSP)
UseMycobacteria expressionTagsCrimsonAvailable SinceJune 30, 2011AvailabilityAcademic Institutions and Nonprofits only -
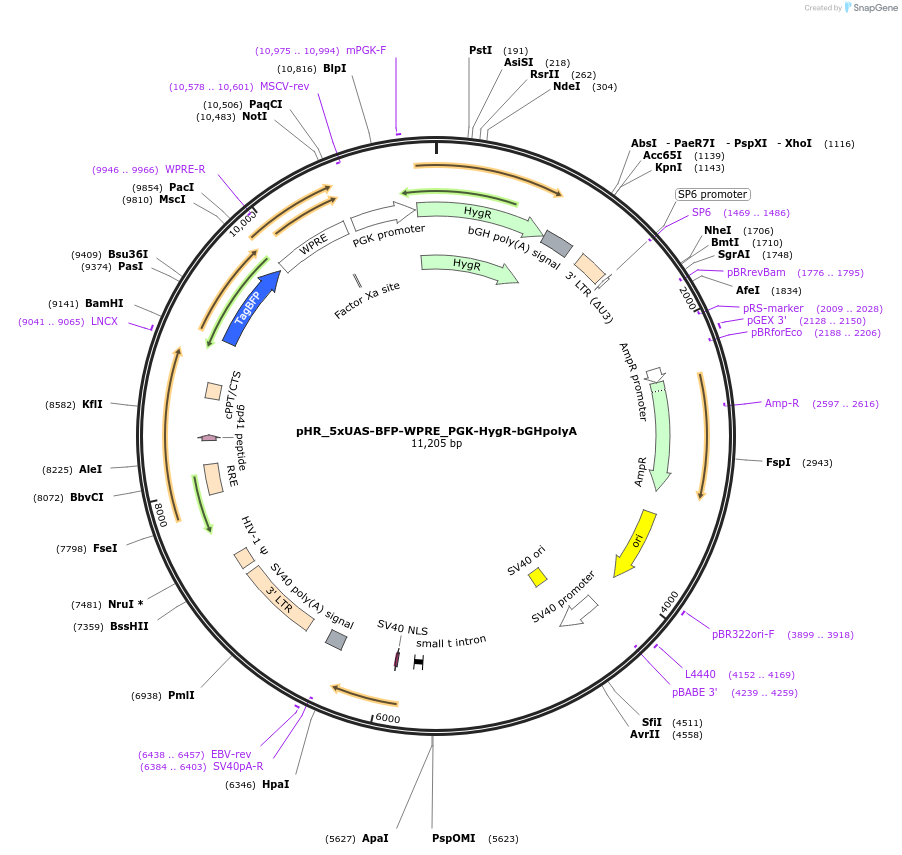
pHR_5xUAS-BFP-WPRE_PGK-HygR-bGHpolyA
Plasmid#216665PurposeExpresses Gal4 inducible BFP, with hygromycin resistanceDepositorInsertsBFP
HygromycinR
UseLentiviralAvailable SinceNov. 14, 2024AvailabilityAcademic Institutions and Nonprofits only -
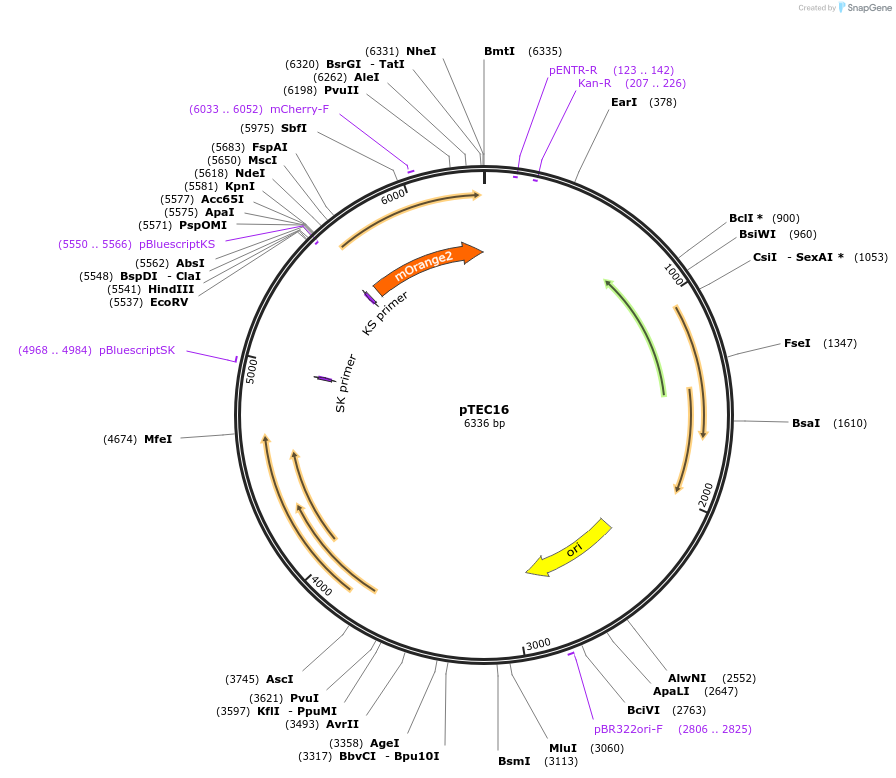
pTEC16
Plasmid#30175DepositorInsertMycobacterium Strong Promoter (MSP)
UseMycobacteria expressionTagsmOrange2Available SinceJune 3, 2011AvailabilityAcademic Institutions and Nonprofits only -
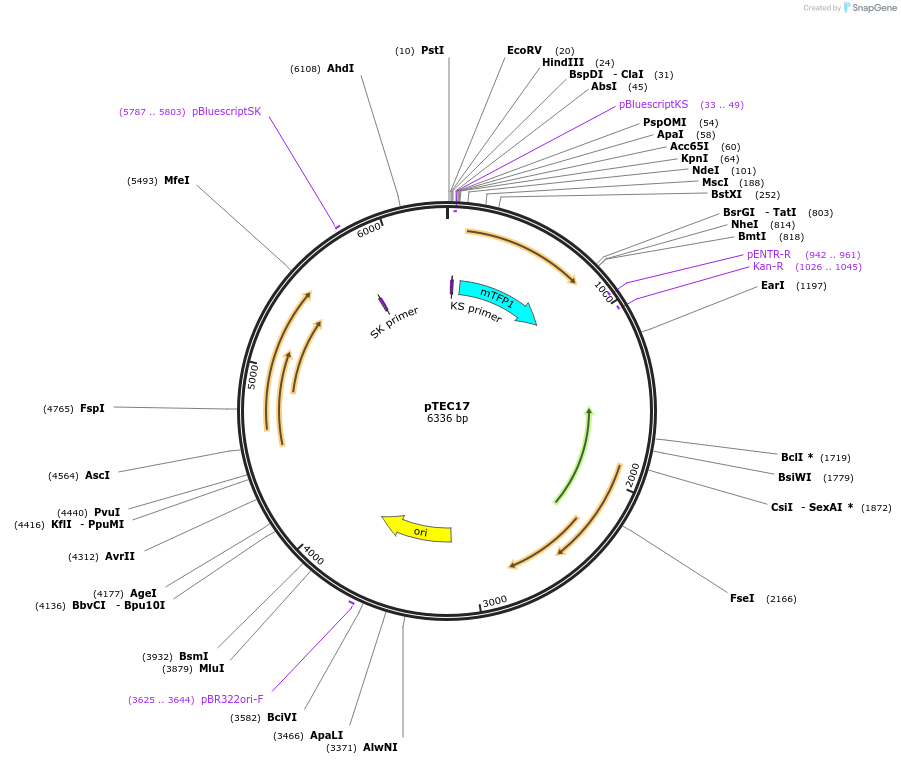
pTEC17
Plasmid#30176DepositorInsertMycobacterium Strong Promoter (MSP)
UseMycobacteria expressionTagsTealAvailable SinceJune 3, 2011AvailabilityAcademic Institutions and Nonprofits only -
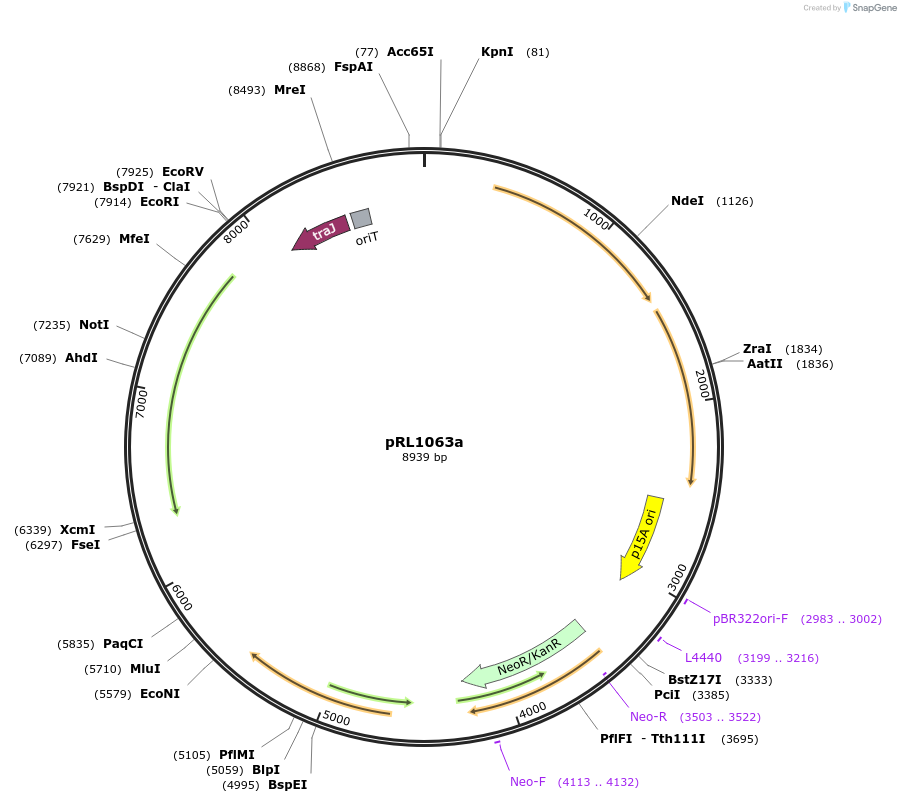
pRL1063a
Plasmid#70687PurposepRL1058 with Vibrio fischeri luxAB for sensingDepositorInsertTransposon Tn5-1063
Available SinceJune 8, 2016AvailabilityAcademic Institutions and Nonprofits only -
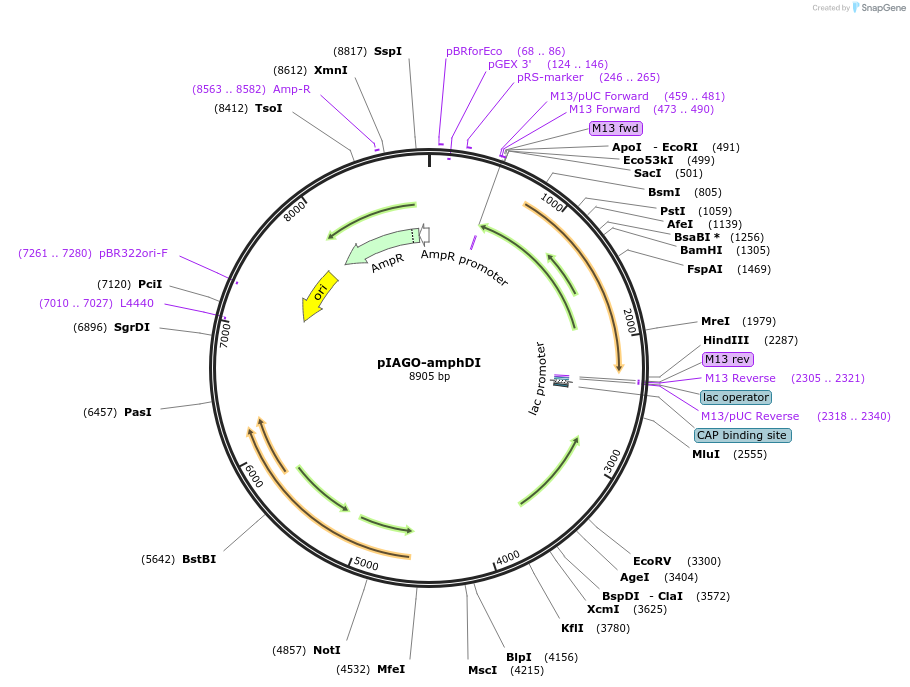
pIAGO-amphDI
Plasmid#114459PurposeExpression of AmphDI glycosyltansferase in streptomycetesDepositorInsertamphDI
ExpressionBacterialPromoterermEAvailable SinceFeb. 21, 2019AvailabilityAcademic Institutions and Nonprofits only -
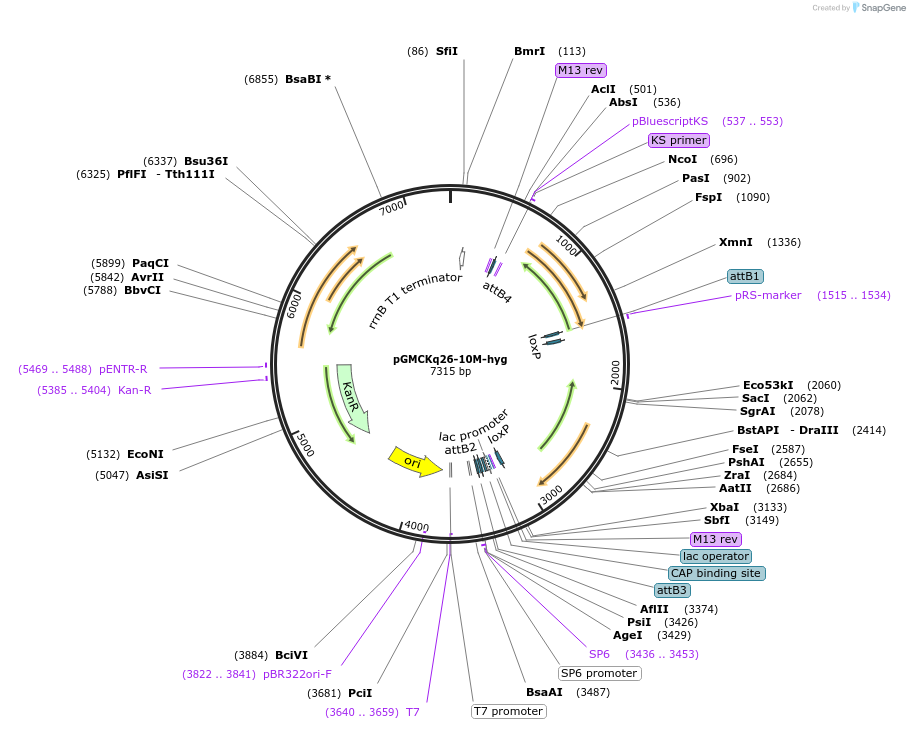
pGMCKq26-10M-hyg
Plasmid#27337DepositorInserttetR(sB), hygR
UseGateway CloningMutationN/AAvailable SinceAug. 24, 2011AvailabilityAcademic Institutions and Nonprofits only -
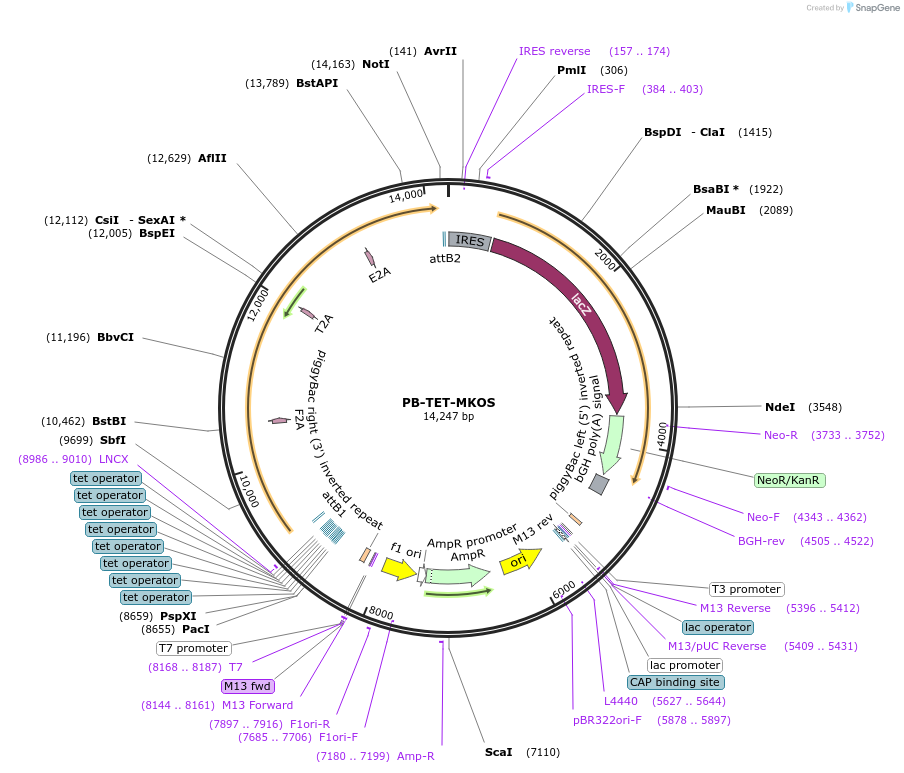
PB-TET-MKOS
Plasmid#20959PurposepiggyBac vector with tet inducible cMyc, KLF4, Oct4, Sox2 for creation of induced pluripotent stem cellsDepositorUsepiggybacTagsIRES-bGeoExpressionMammalianAvailable SinceApril 23, 2009AvailabilityAcademic Institutions and Nonprofits only -
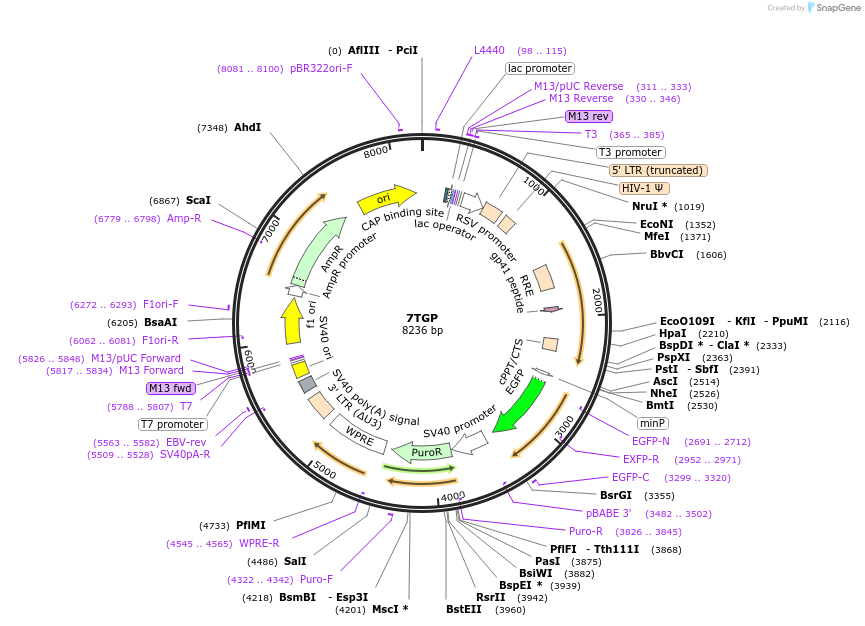
7TGP
Plasmid#24305DepositorInsert7xTcf-eGFP // SV40-PuroR
UseLentiviralMutationEGFP is GFP with S65T and F64L mutationsAvailable SinceApril 5, 2010AvailabilityAcademic Institutions and Nonprofits only -
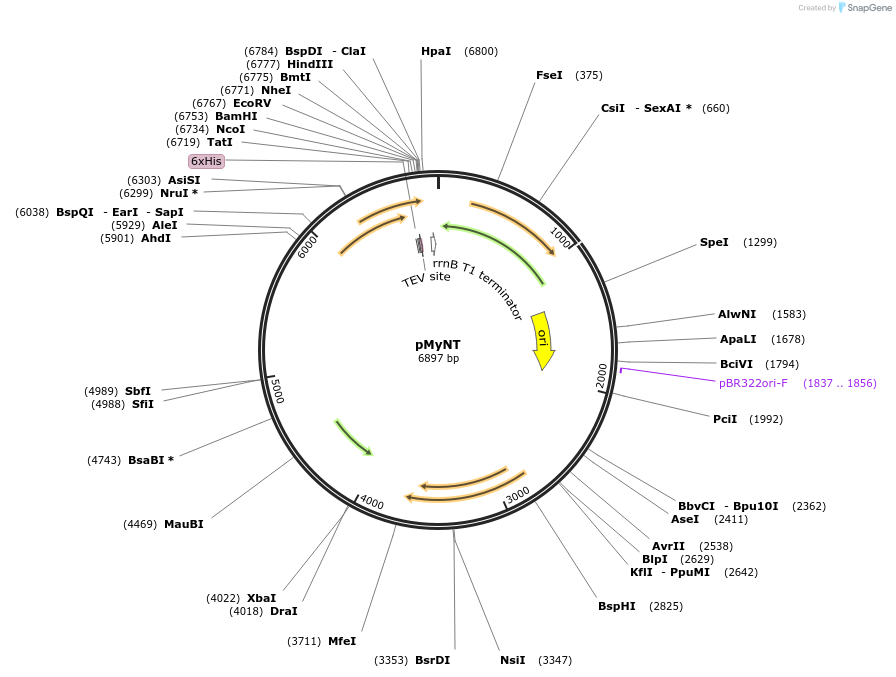
pMyNT
Plasmid#42191DepositorTypeEmpty backboneTagsHis6ExpressionBacterialPromoterColE1Available SinceAug. 5, 2013AvailabilityAcademic Institutions and Nonprofits only -
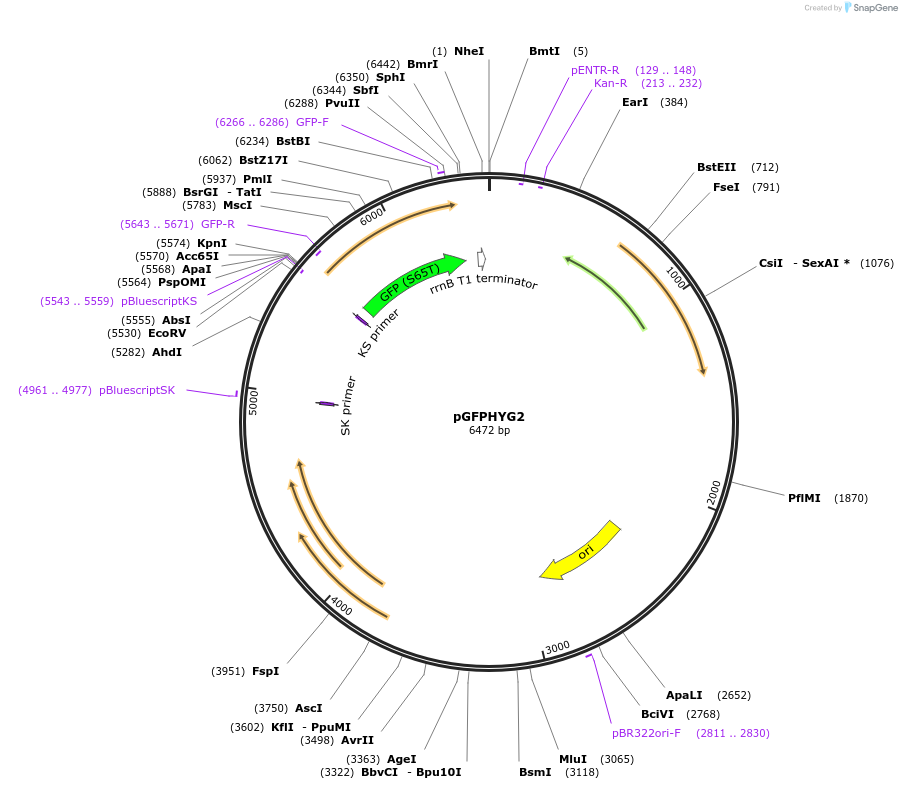
pGFPHYG2
Plasmid#30173DepositorInsertMycobacterium Strong Promoter (MSP)
UseMycobacteria expressionTagsGFPMutationGFPmut3 allele has improved brightness.Available SinceJune 6, 2011AvailabilityAcademic Institutions and Nonprofits only