We narrowed to 11,549 results for: phen
-
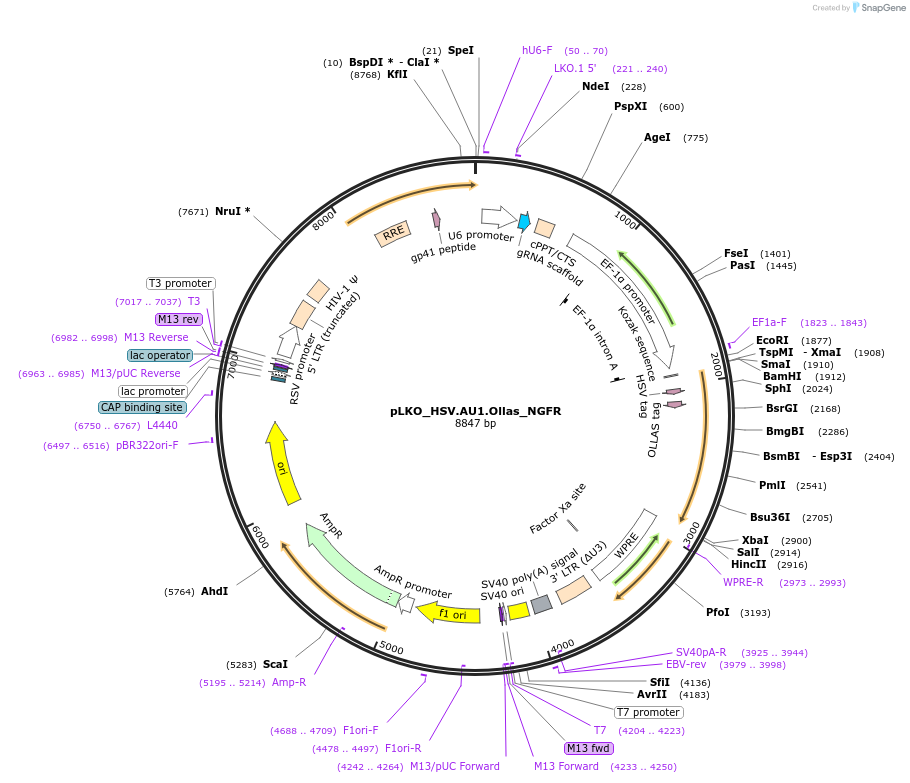
Plasmid#158273PurposeProtein Barcode (Pro-Code) for vector/cell tracking. Contains U6 trRNA cassette for sgRNA cloning. Enables phenotypic CRISPR screens at a single cell resolution (using cytometry).DepositorInsertPro-Code Tagged human dNGFR (NGFR Human)
UseCRISPR and LentiviralTagsHSV.AU1.OllasMutationTruncation of the signaling domainPromoterEF1aAvailable SinceSept. 3, 2021AvailabilityAcademic Institutions and Nonprofits only -
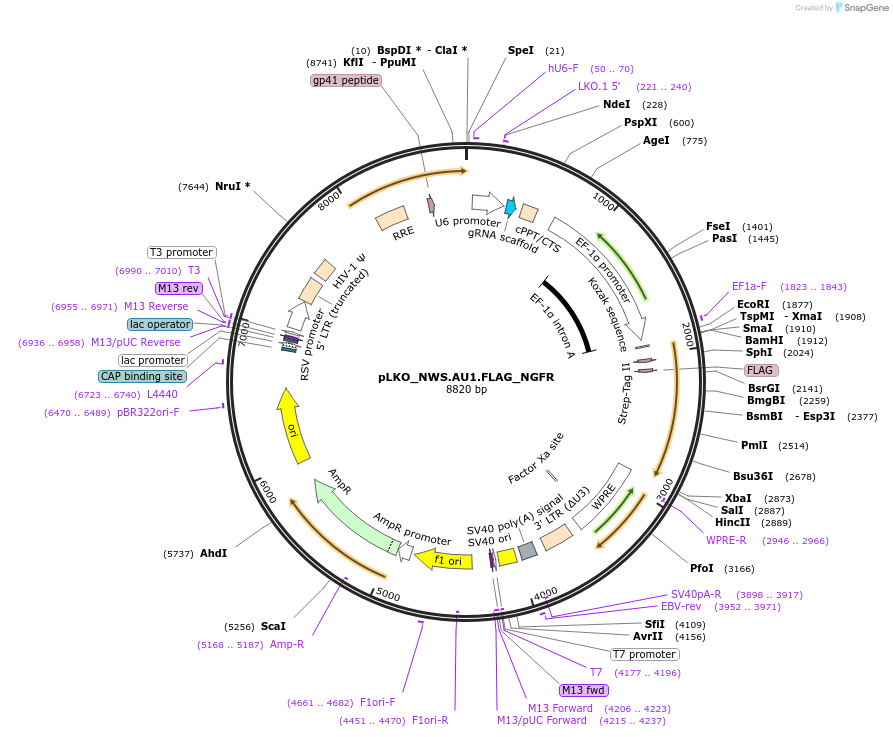
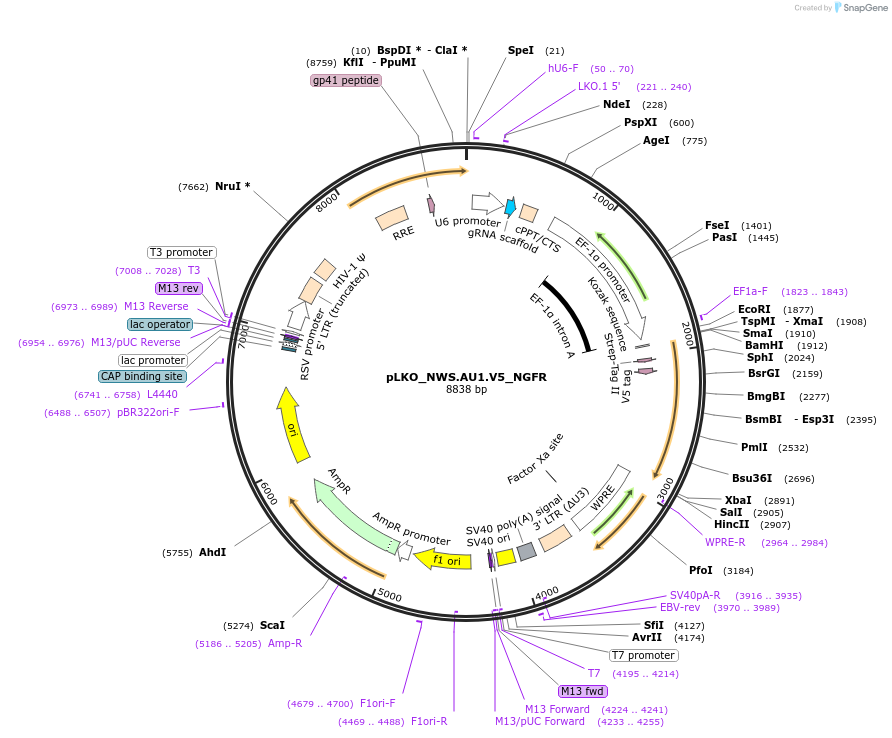
pLKO_NWS.AU1.FLAG_NGFR
Plasmid#158238PurposeProtein Barcode (Pro-Code) for vector/cell tracking. Contains U6 trRNA cassette for sgRNA cloning. Enables phenotypic CRISPR screens at a single cell resolution (using cytometry).DepositorInsertPro-Code Tagged human dNGFR (NGFR Human)
UseCRISPR and LentiviralTagsNWS.AU1.FLAGMutationTruncation of the signaling domainPromoterEF1aAvailable SinceSept. 3, 2021AvailabilityAcademic Institutions and Nonprofits only -
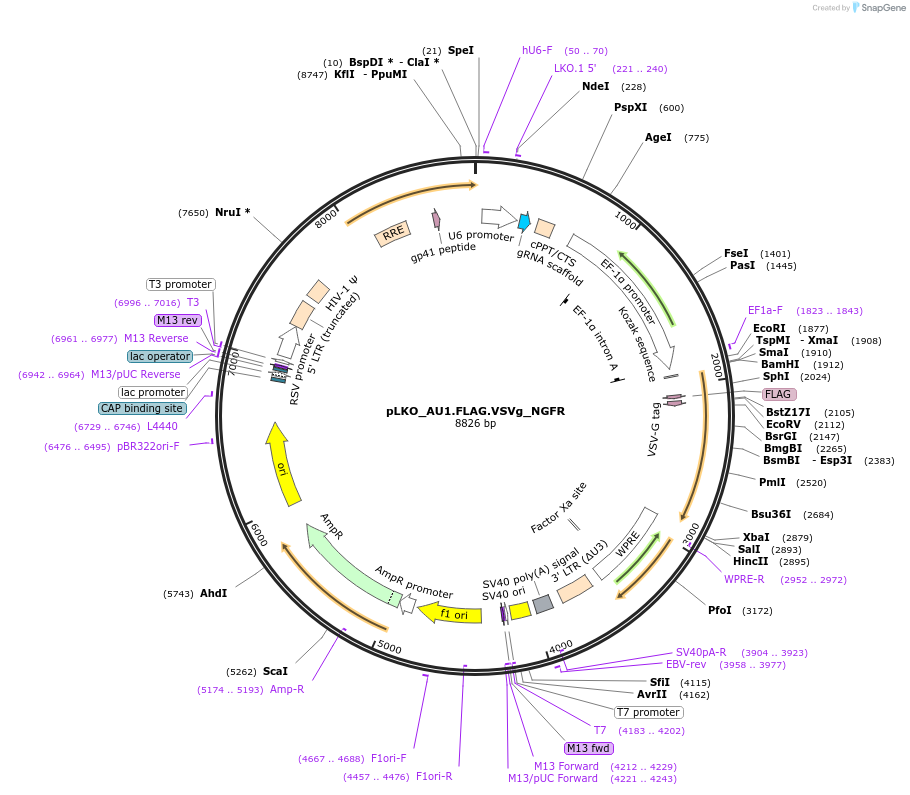
pLKO_AU1.FLAG.VSVg_NGFR
Plasmid#158242PurposeProtein Barcode (Pro-Code) for vector/cell tracking. Contains U6 trRNA cassette for sgRNA cloning. Enables phenotypic CRISPR screens at a single cell resolution (using cytometry).DepositorInsertPro-Code Tagged human dNGFR (NGFR Human)
UseCRISPR and LentiviralTagsAU1.FLAG.VSVgMutationTruncation of the signaling domainPromoterEF1aAvailable SinceSept. 3, 2021AvailabilityAcademic Institutions and Nonprofits only -
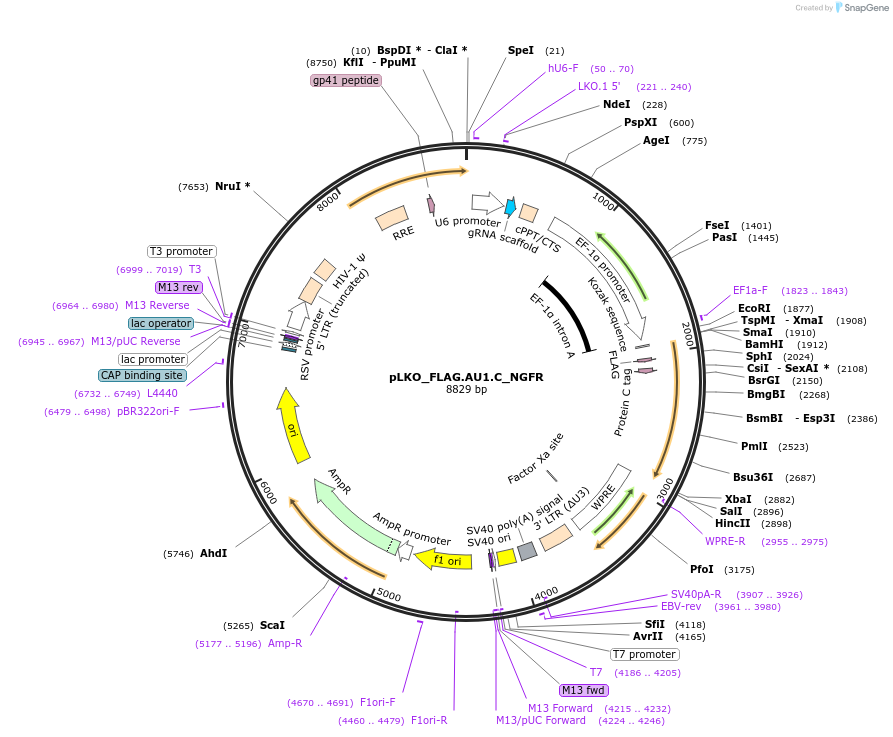
pLKO_FLAG.AU1.C_NGFR
Plasmid#158252PurposeProtein Barcode (Pro-Code) for vector/cell tracking. Contains U6 trRNA cassette for sgRNA cloning. Enables phenotypic CRISPR screens at a single cell resolution (using cytometry).DepositorInsertPro-Code Tagged human dNGFR (NGFR Human)
UseCRISPR and LentiviralTagsFLAG.AU1.CMutationTruncation of the signaling domainPromoterEF1aAvailable SinceSept. 3, 2021AvailabilityAcademic Institutions and Nonprofits only -
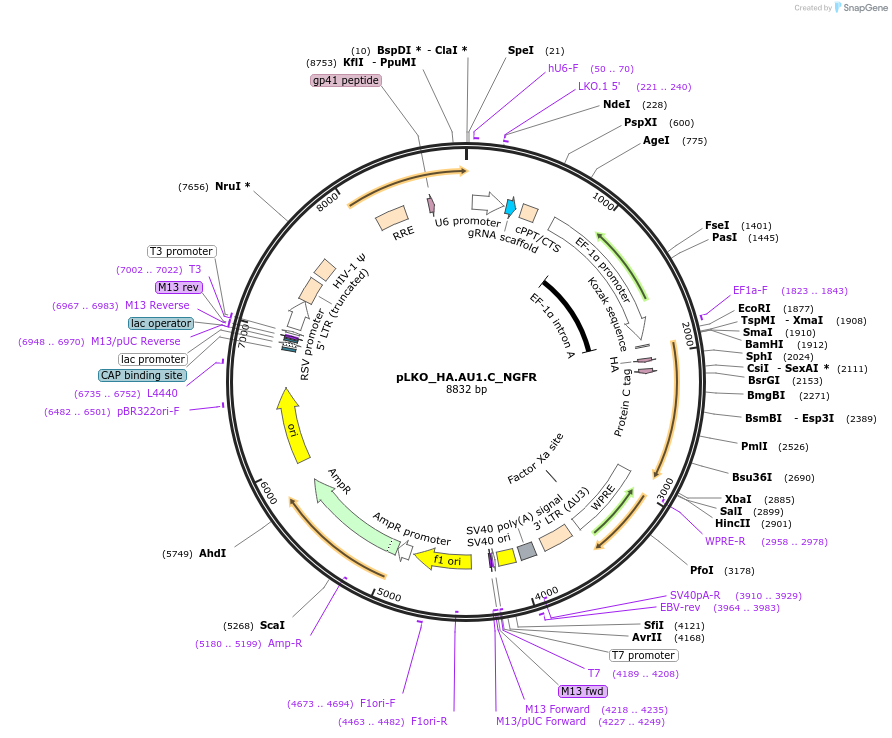
pLKO_HA.AU1.C_NGFR
Plasmid#158253PurposeProtein Barcode (Pro-Code) for vector/cell tracking. Contains U6 trRNA cassette for sgRNA cloning. Enables phenotypic CRISPR screens at a single cell resolution (using cytometry).DepositorInsertPro-Code Tagged human dNGFR (NGFR Human)
UseCRISPR and LentiviralTagsHA.AU1.CMutationTruncation of the signaling domainPromoterEF1aAvailable SinceSept. 3, 2021AvailabilityAcademic Institutions and Nonprofits only -
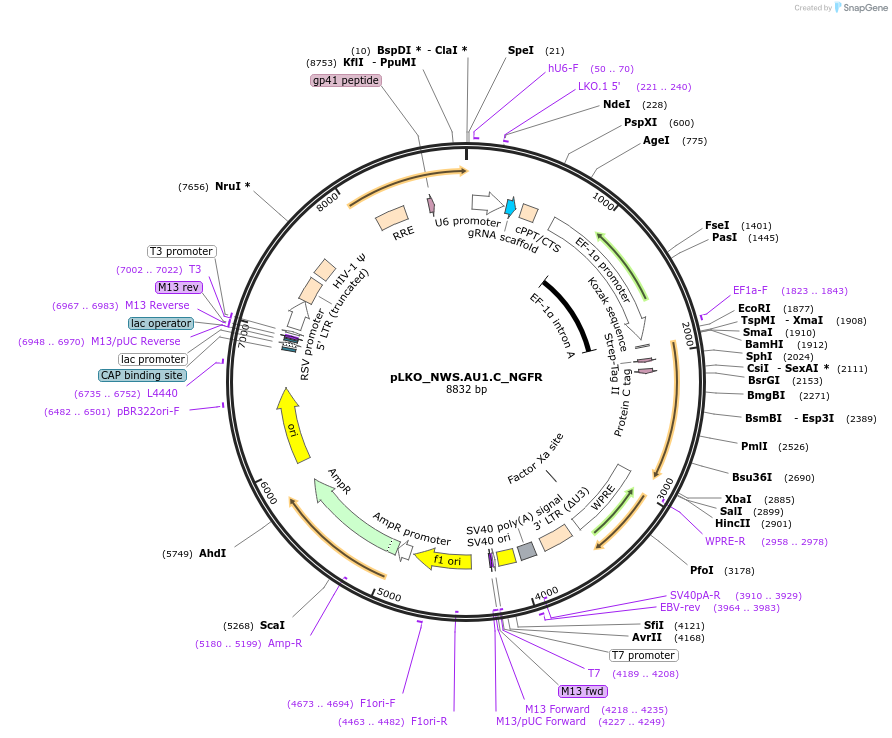
pLKO_NWS.AU1.C_NGFR
Plasmid#158254PurposeProtein Barcode (Pro-Code) for vector/cell tracking. Contains U6 trRNA cassette for sgRNA cloning. Enables phenotypic CRISPR screens at a single cell resolution (using cytometry).DepositorInsertPro-Code Tagged human dNGFR (NGFR Human)
UseCRISPR and LentiviralTagsNWS.AU1.CMutationTruncation of the signaling domainPromoterEF1aAvailable SinceSept. 3, 2021AvailabilityAcademic Institutions and Nonprofits only -
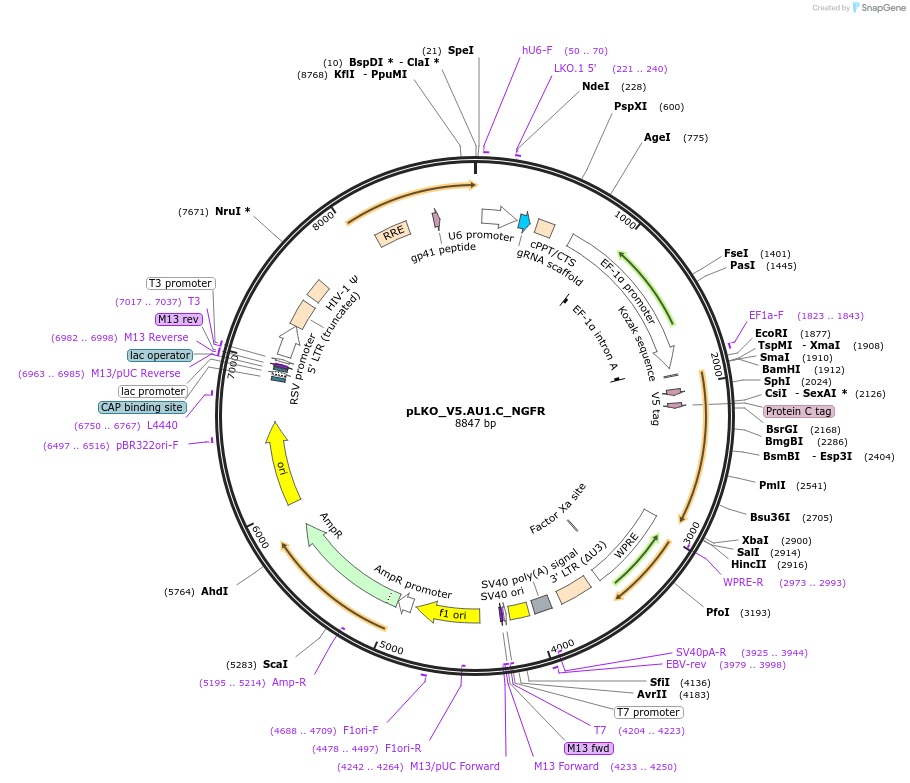
pLKO_V5.AU1.C_NGFR
Plasmid#158255PurposeProtein Barcode (Pro-Code) for vector/cell tracking. Contains U6 trRNA cassette for sgRNA cloning. Enables phenotypic CRISPR screens at a single cell resolution (using cytometry).DepositorInsertPro-Code Tagged human dNGFR (NGFR Human)
UseCRISPR and LentiviralTagsV5.AU1.CMutationTruncation of the signaling domainPromoterEF1aAvailable SinceSept. 3, 2021AvailabilityAcademic Institutions and Nonprofits only -
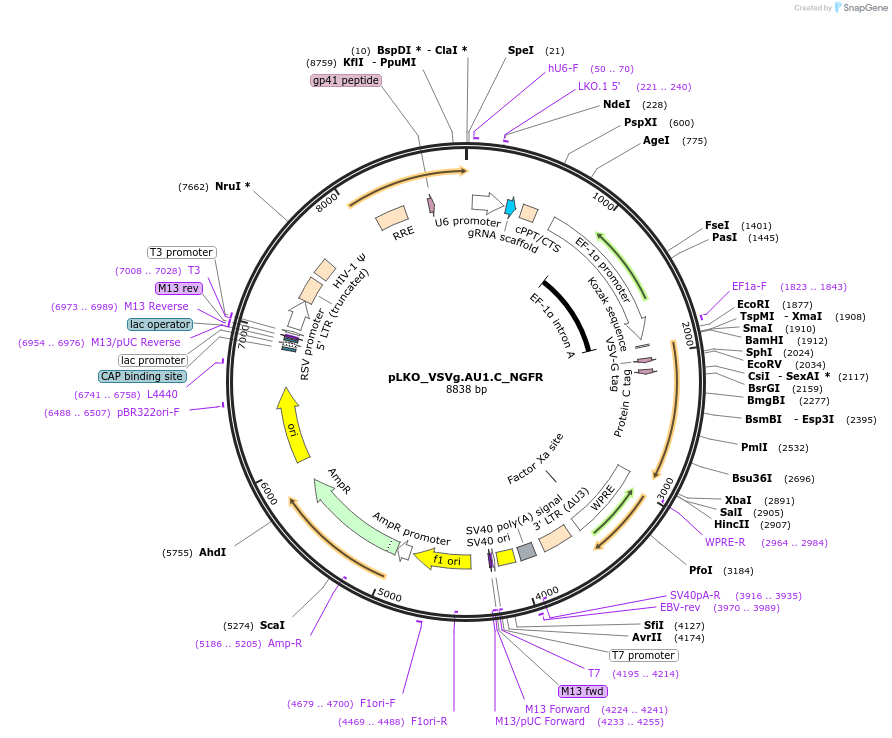
pLKO_VSVg.AU1.C_NGFR
Plasmid#158256PurposeProtein Barcode (Pro-Code) for vector/cell tracking. Contains U6 trRNA cassette for sgRNA cloning. Enables phenotypic CRISPR screens at a single cell resolution (using cytometry).DepositorInsertPro-Code Tagged human dNGFR (NGFR Human)
UseCRISPR and LentiviralTagsVSVg.AU1.CMutationTruncation of the signaling domainPromoterEF1aAvailable SinceSept. 3, 2021AvailabilityAcademic Institutions and Nonprofits only -
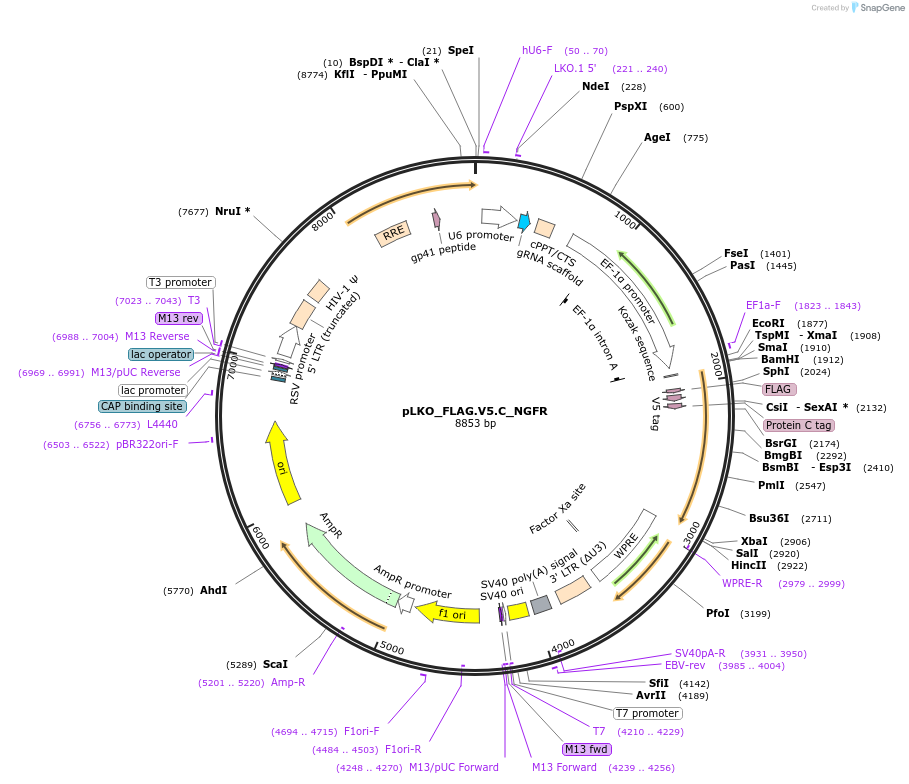
pLKO_FLAG.V5.C_NGFR
Plasmid#158259PurposeProtein Barcode (Pro-Code) for vector/cell tracking. Contains U6 trRNA cassette for sgRNA cloning. Enables phenotypic CRISPR screens at a single cell resolution (using cytometry).DepositorInsertPro-Code Tagged human dNGFR (NGFR Human)
UseCRISPR and LentiviralTagsFLAG.V5.CMutationTruncation of the signaling domainPromoterEF1aAvailable SinceSept. 3, 2021AvailabilityAcademic Institutions and Nonprofits only -
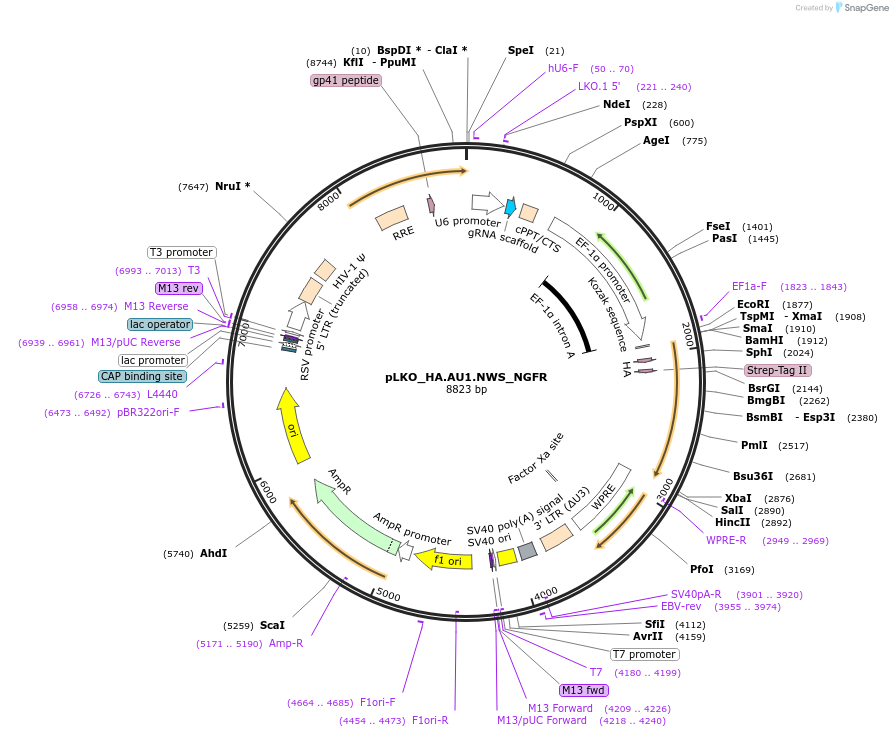
pLKO_HA.AU1.NWS_NGFR
Plasmid#158234PurposeProtein Barcode (Pro-Code) for vector/cell tracking. Contains U6 trRNA cassette for sgRNA cloning. Enables phenotypic CRISPR screens at a single cell resolution (using cytometry).DepositorInsertPro-Code Tagged human dNGFR (NGFR Human)
UseCRISPR and LentiviralTagsHA.AU1.NWSMutationTruncation of the signaling domainPromoterEF1aAvailable SinceSept. 3, 2021AvailabilityAcademic Institutions and Nonprofits only -
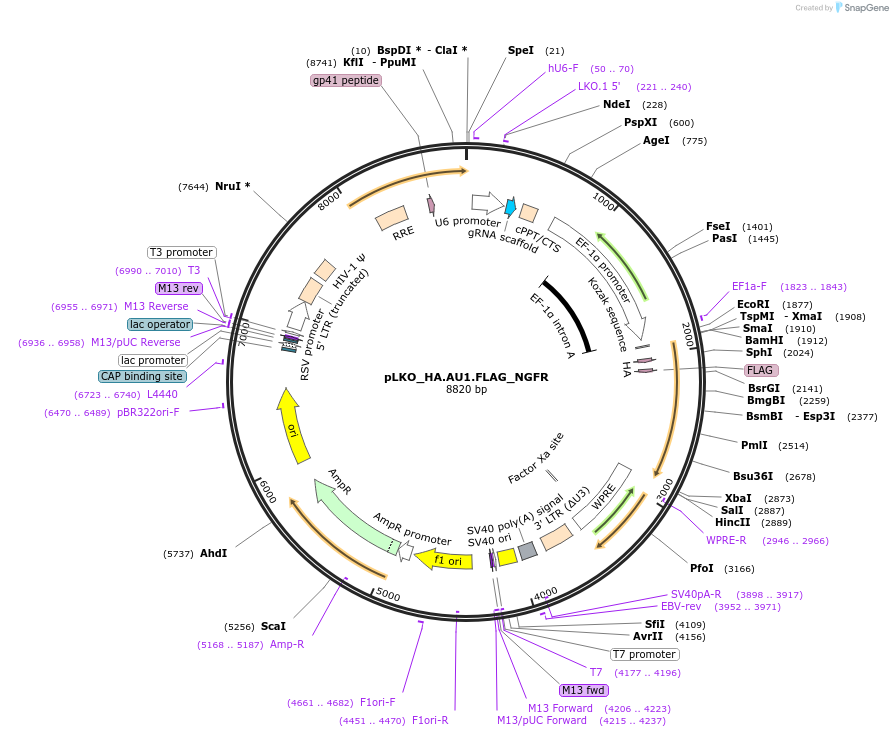
pLKO_HA.AU1.FLAG_NGFR
Plasmid#158236PurposeProtein Barcode (Pro-Code) for vector/cell tracking. Contains U6 trRNA cassette for sgRNA cloning. Enables phenotypic CRISPR screens at a single cell resolution (using cytometry).DepositorInsertPro-Code Tagged human dNGFR (NGFR Human)
UseCRISPR and LentiviralTagsHA.AU1.FLAGMutationTruncation of the signaling domainPromoterEF1aAvailable SinceSept. 3, 2021AvailabilityAcademic Institutions and Nonprofits only -
pLKO_NWS.AU1.V5_NGFR
Plasmid#158237PurposeProtein Barcode (Pro-Code) for vector/cell tracking. Contains U6 trRNA cassette for sgRNA cloning. Enables phenotypic CRISPR screens at a single cell resolution (using cytometry).DepositorInsertPro-Code Tagged human dNGFR (NGFR Human)
UseCRISPR and LentiviralTagsNWS.AU1.V5MutationTruncation of the signaling domainPromoterEF1aAvailable SinceSept. 3, 2021AvailabilityAcademic Institutions and Nonprofits only -
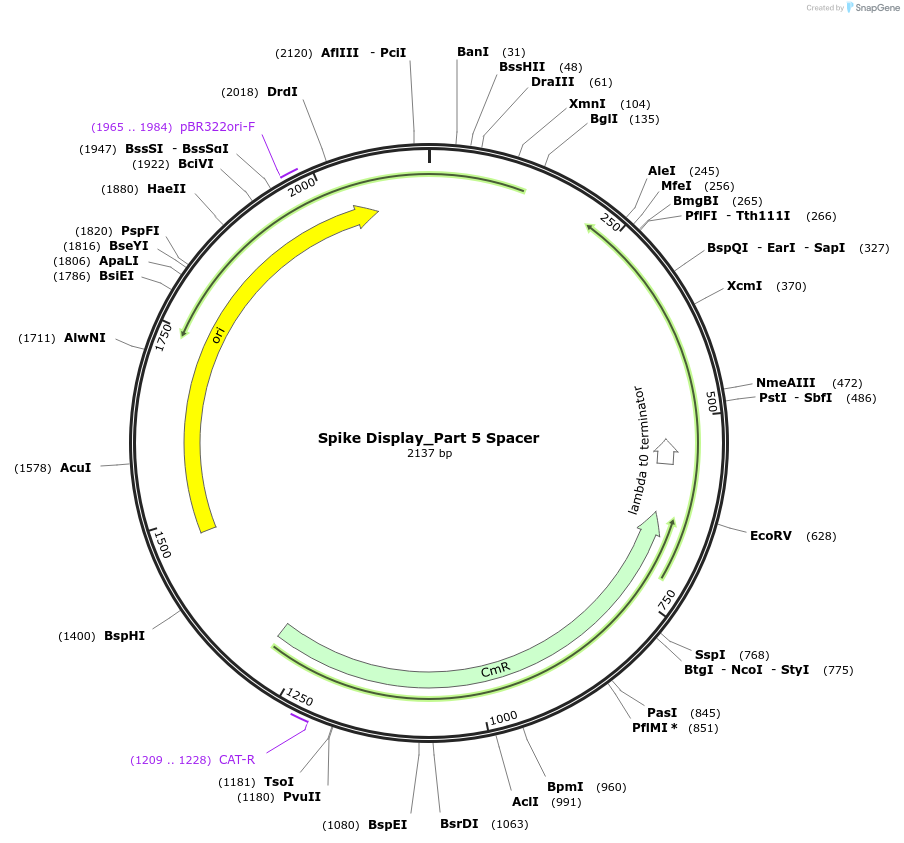
Spike Display_Part 5 Spacer
Plasmid#172733PurposeEncodes Part 5 of HexaPro-D614G spike to be used in the Spike Display systemDepositorInsertSARS-CoV-2 S HexaPro-D614G (S Severe acute respiratory syndrome coronavirus 2)
UseSynthetic BiologyMutationEctodomain only (AAs 1-1208); 682-685 (furin site…Available SinceAug. 17, 2021AvailabilityIndustry, Academic Institutions, and Nonprofits -
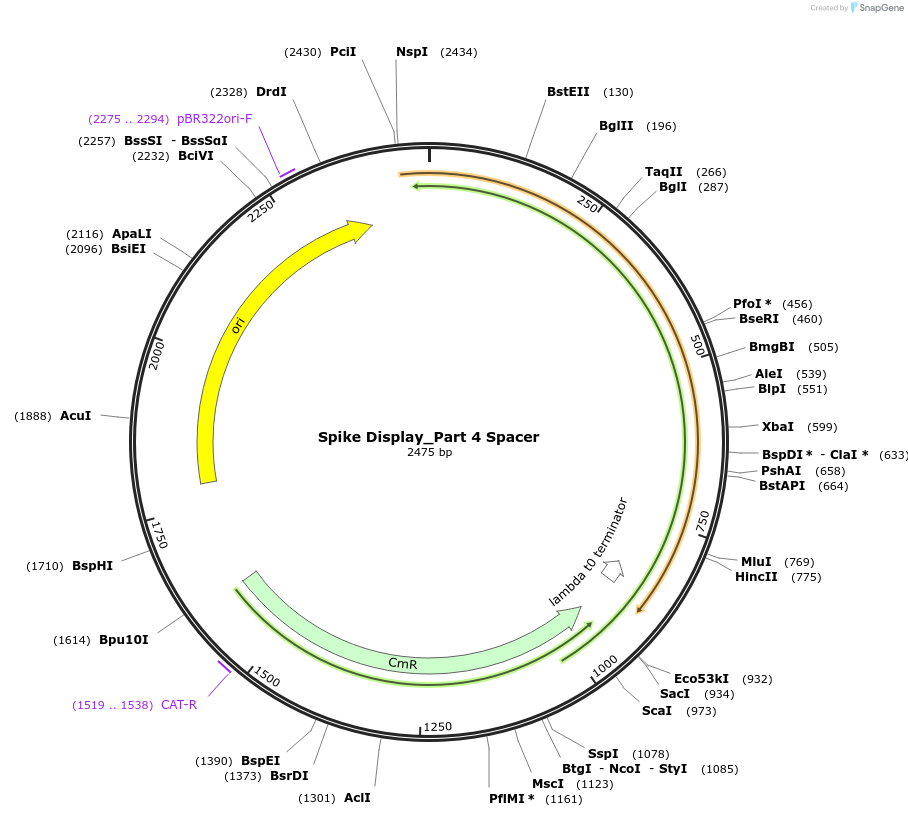
Spike Display_Part 4 Spacer
Plasmid#172732PurposeEncodes Part 4 of HexaPro-D614G spike to be used in the Spike Display systemDepositorInsertSARS-CoV-2 S HexaPro-D614G (S Severe acute respiratory syndrome coronavirus 2)
UseSynthetic BiologyMutationEctodomain only (AAs 1-1208); 682-685 (furin site…Available SinceAug. 17, 2021AvailabilityIndustry, Academic Institutions, and Nonprofits -
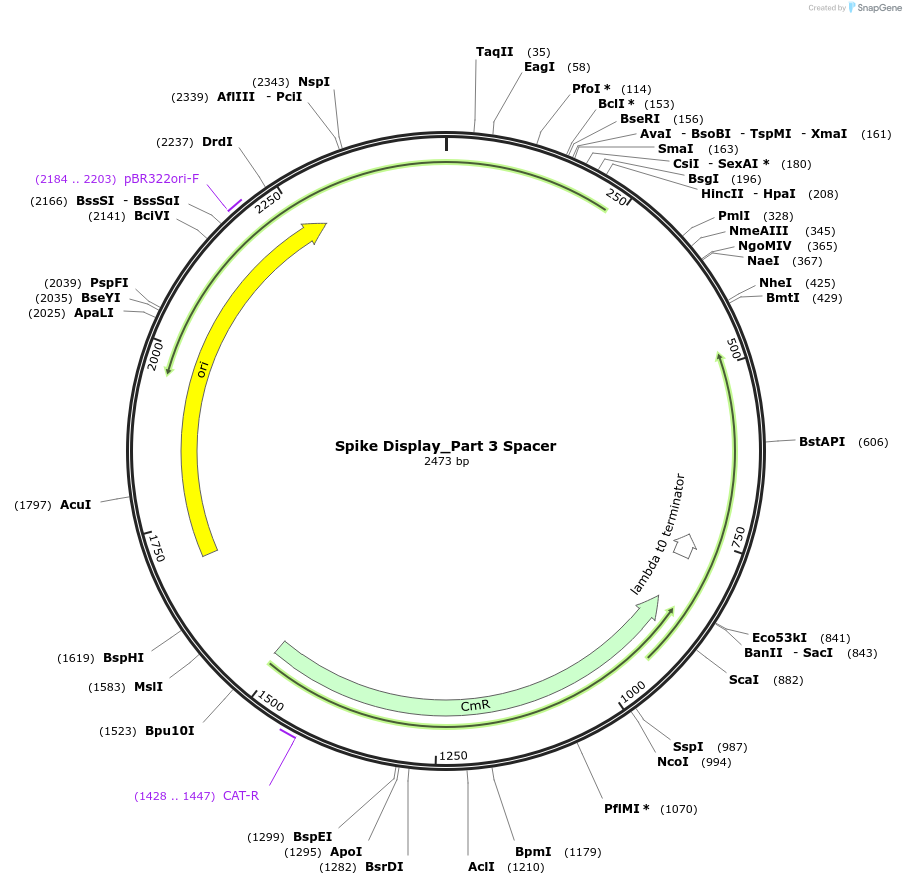
Spike Display_Part 3 Spacer
Plasmid#172731PurposeEncodes Part 3 of HexaPro-D614G spike to be used in the Spike Display systemDepositorInsertSARS-CoV-2 S HexaPro-D614G (S Severe acute respiratory syndrome coronavirus 2)
UseSynthetic BiologyMutationEctodomain only (AAs 1-1208); 682-685 (furin site…Available SinceAug. 17, 2021AvailabilityIndustry, Academic Institutions, and Nonprofits -
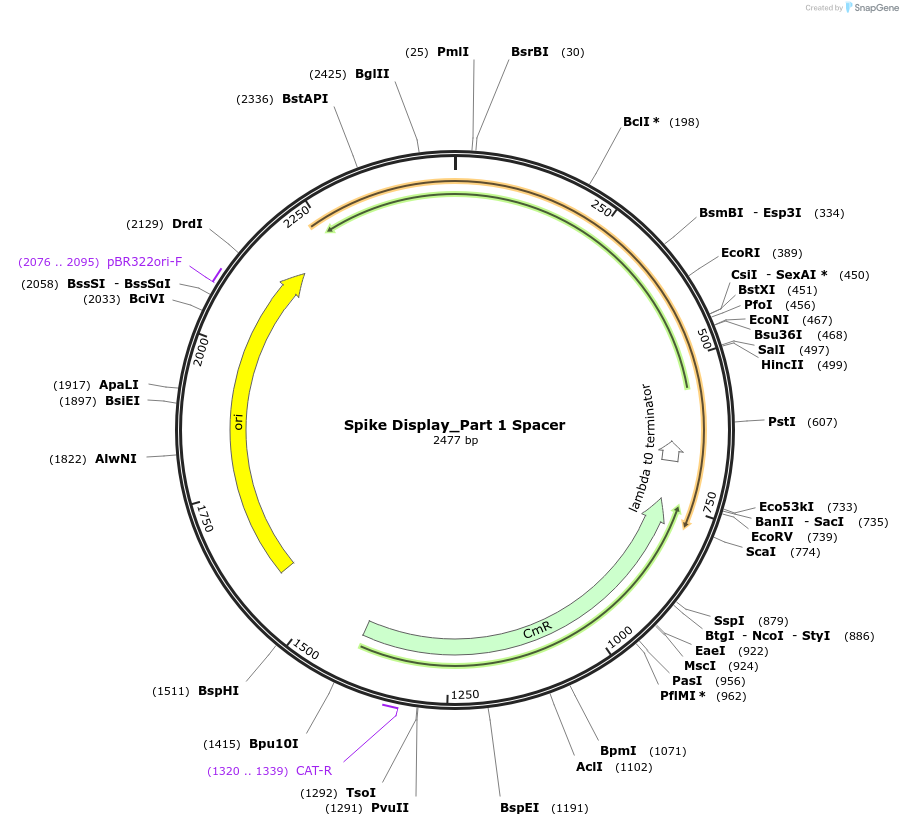
Spike Display_Part 1 Spacer
Plasmid#172729PurposeEncodes Part 1 of HexaPro-D614G spike to be used in the Spike Display systemDepositorInsertSARS-CoV-2 S HexaPro-D614G (S Severe acute respiratory syndrome coronavirus 2)
UseSynthetic BiologyMutationEctodomain only (AAs 1-1208); 682-685 (furin site…Available SinceAug. 17, 2021AvailabilityIndustry, Academic Institutions, and Nonprofits -
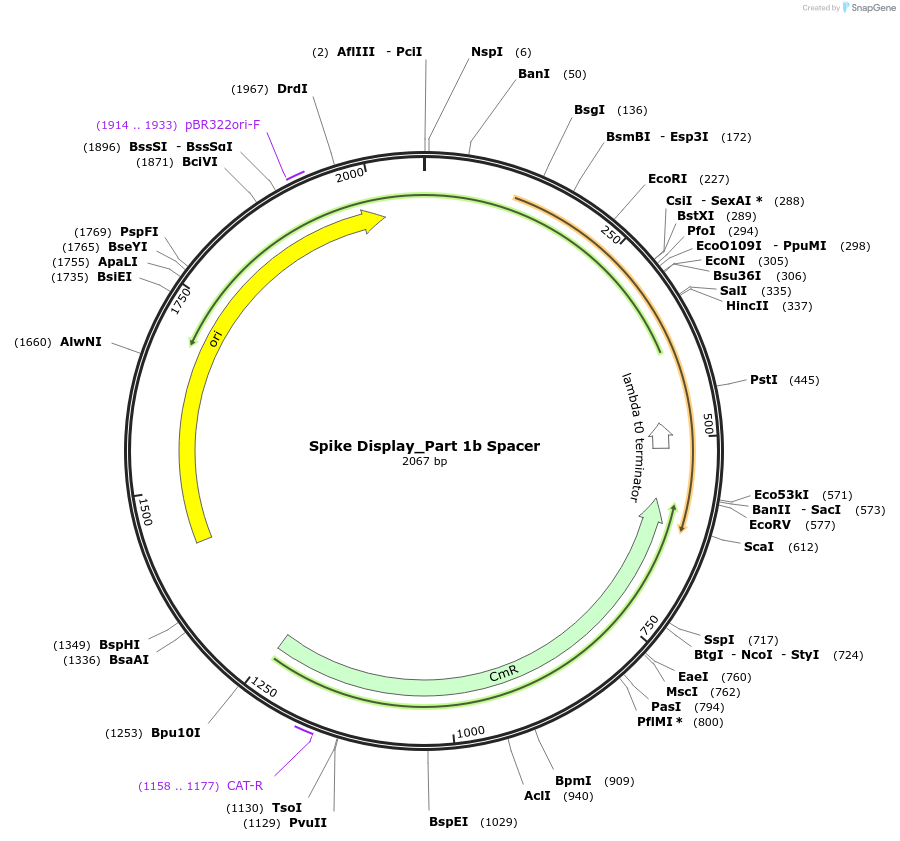
Spike Display_Part 1b Spacer
Plasmid#172728PurposeEncodes Part 1b of HexaPro-D614G spike to be used in the Spike Display systemDepositorInsertSARS-CoV-2 S HexaPro-D614G (S Severe acute respiratory syndrome coronavirus 2)
UseSynthetic BiologyMutationEctodomain only (AAs 1-1208); 682-685 (furin site…Available SinceAug. 17, 2021AvailabilityIndustry, Academic Institutions, and Nonprofits -
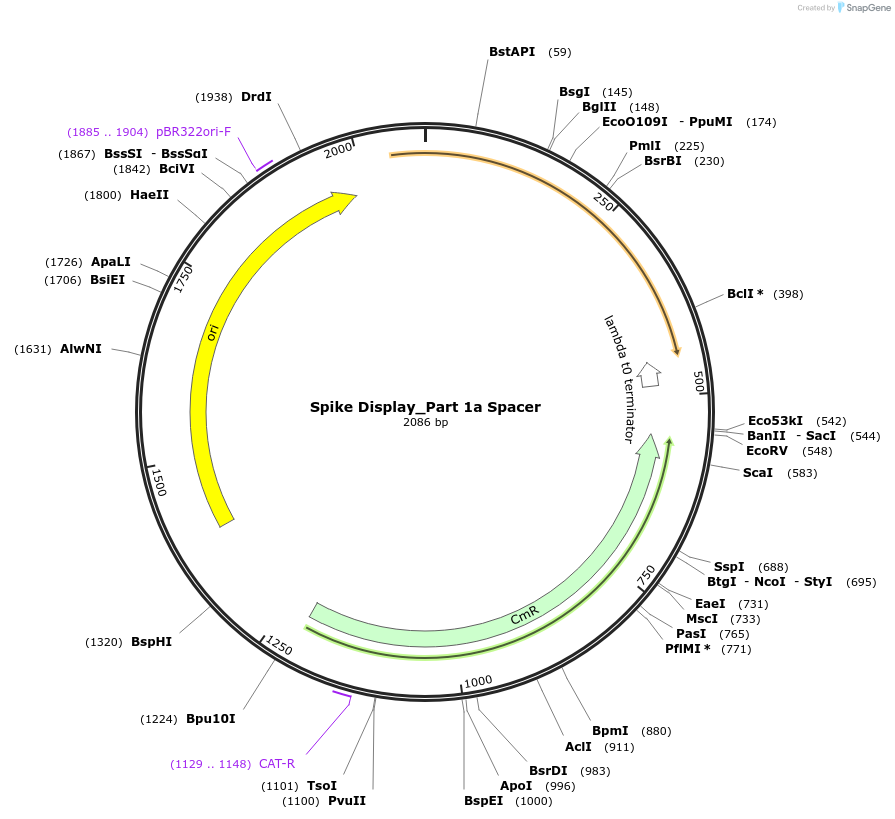
Spike Display_Part 1a Spacer
Plasmid#172727PurposeEncodes Part 1a of HexaPro-D614G spike to be used in the Spike Display systemDepositorInsertSARS-CoV-2 S HexaPro-D614G (S Severe acute respiratory syndrome coronavirus 2)
UseSynthetic BiologyMutationEctodomain only (AAs 1-1208); 682-685 (furin site…Available SinceAug. 17, 2021AvailabilityIndustry, Academic Institutions, and Nonprofits -
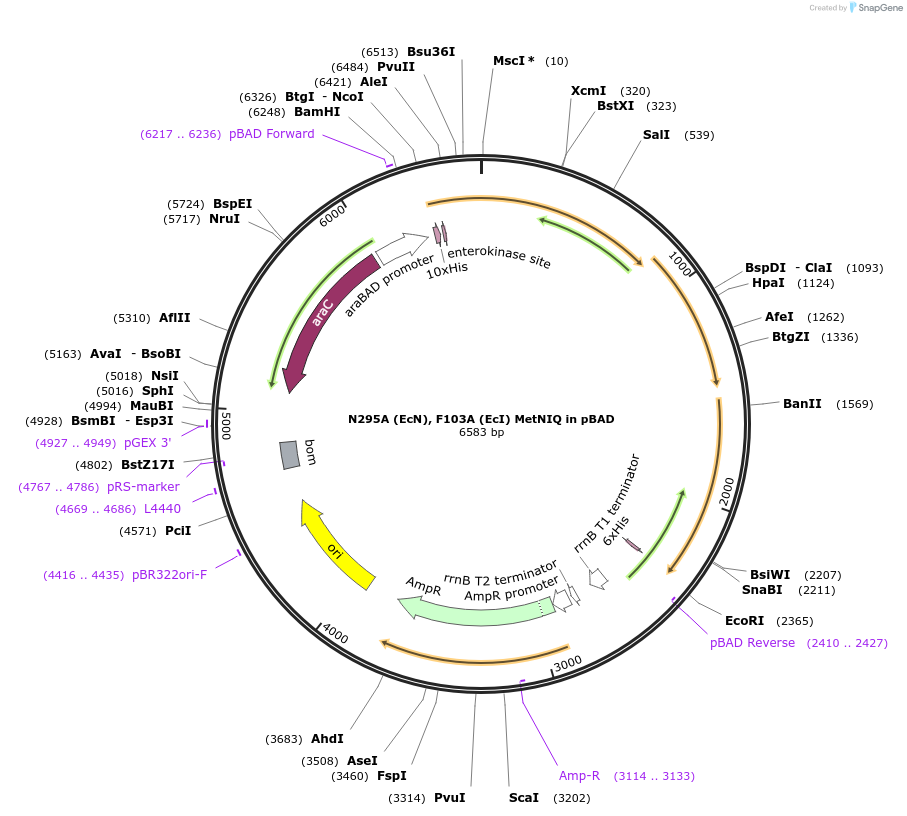
N295A (EcN), F103A (EcI) MetNIQ in pBAD
Plasmid#118258PurposeN295A Mutation of MetN and F103A in MetI in Methionine ABC Transporter and Wild Type Methionine Binding Protein for Transport Activity AssayDepositorTags10x HIs Tag, 6x His Tag, and Enterokinase Cleavag…ExpressionBacterialMutationChanged Glutamic Acid (Glu) 295 to Alanine (Ala) …PromoterNone and araBAD PromoterAvailable SinceNov. 20, 2018AvailabilityAcademic Institutions and Nonprofits only -
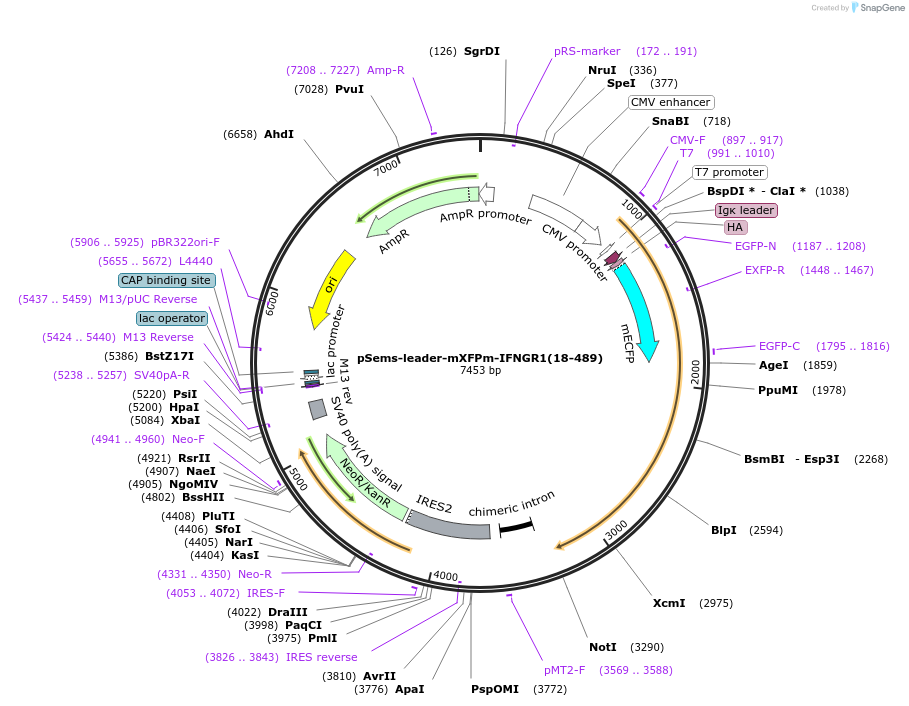
pSems-leader-mXFPm-IFNGR1(18-489)
Plasmid#192786PurposeExpression of mXFPm-tagged IFNGR1 at the plasma membrane, e.g. for single molecule fluorescence microscopyDepositorInsertIFNGR1 (IFNGR1 Human)
TagsIg k-chain leader sequence and mXFPmExpressionMammalianMutationmXFPm: tryptophan 66 to phenylalanine, gluatmic a…PromoterCMVAvailable SinceDec. 21, 2022AvailabilityAcademic Institutions and Nonprofits only -
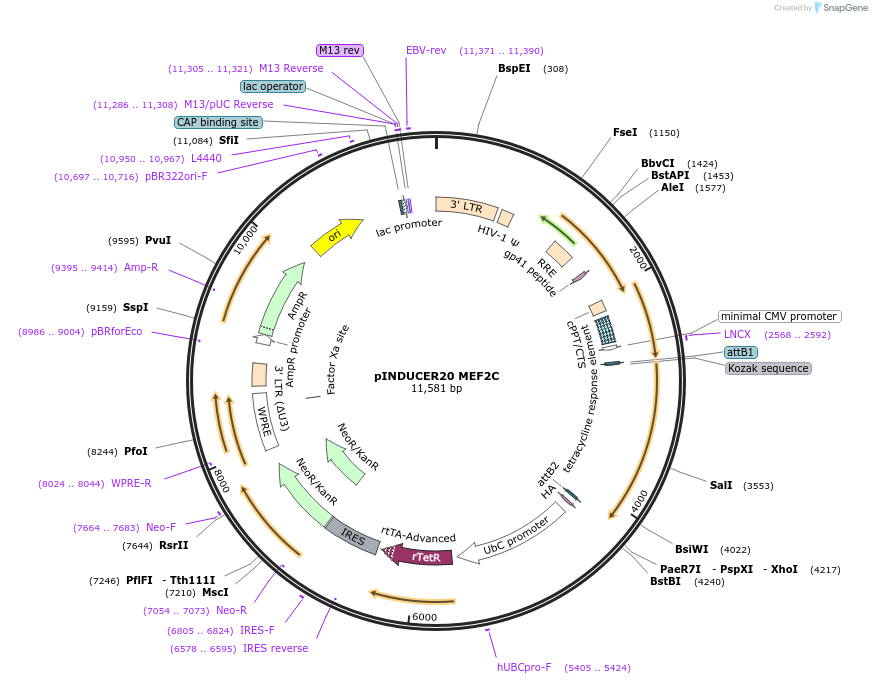
pINDUCER20 MEF2C
Plasmid#89717PurposeLentiviral gene expression vector for doxycycline inducible MEF2C expressionDepositorAvailable SinceMay 23, 2017AvailabilityAcademic Institutions and Nonprofits only -
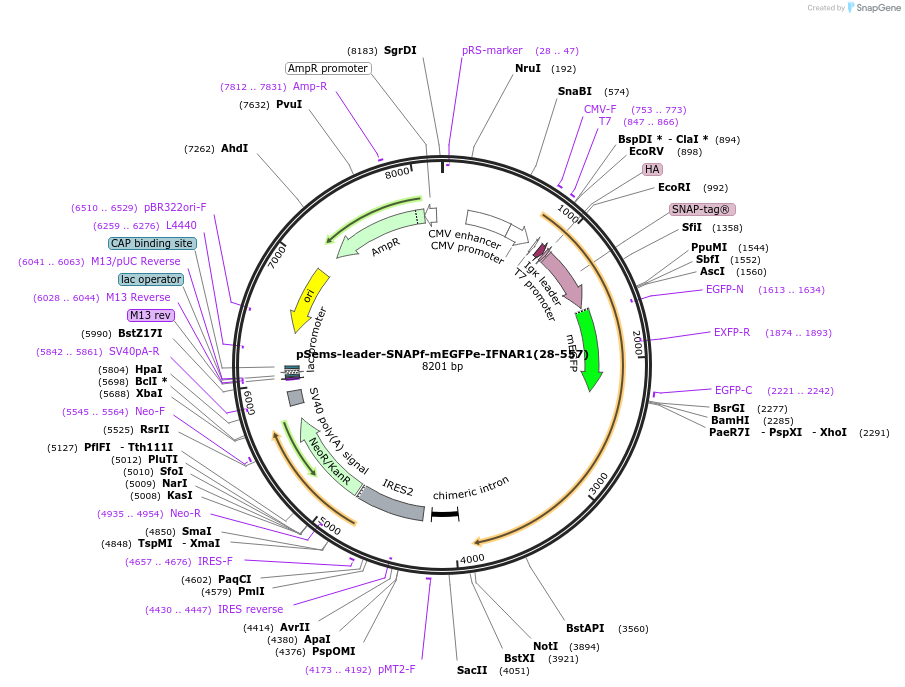
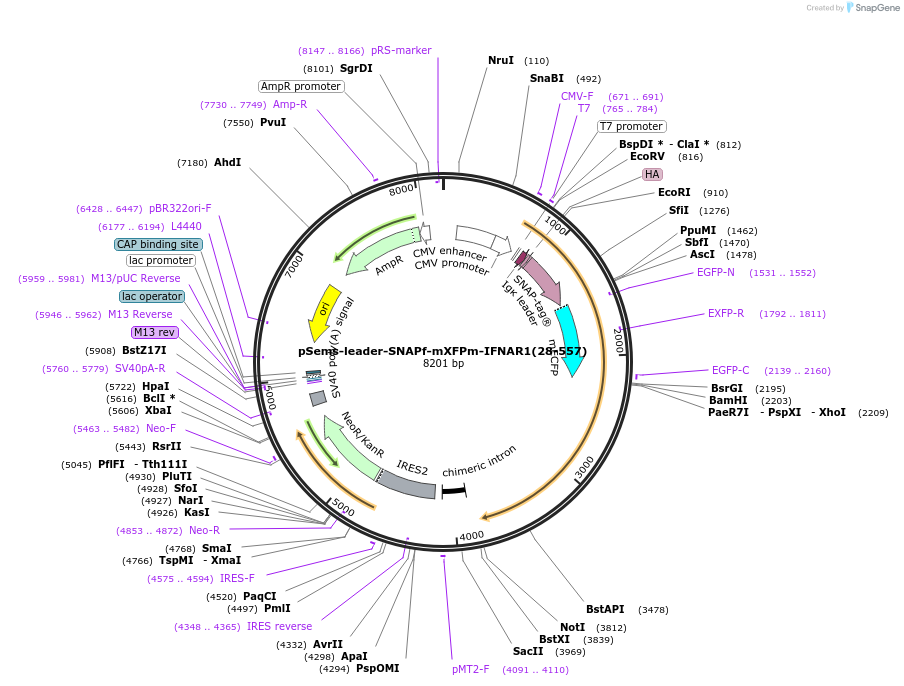
pSems-leader-SNAPf-mEGFPe-IFNAR1(28-557)
Plasmid#187844PurposeExpression of SNAPf and mEGFPe-tagged IFNAR1 at the plasma membrane, e.g. for single molecule fluorescence microscopyDepositorInsertIFNAR1 (IFNAR1 Human)
TagsIg k-chain leader sequence, SNAPf-tag, and meGFPe…ExpressionMammalianMutationmeGFPe: asparagine 198 to aspartic acid and tyros…PromoterCMVAvailable SinceDec. 21, 2022AvailabilityAcademic Institutions and Nonprofits only -
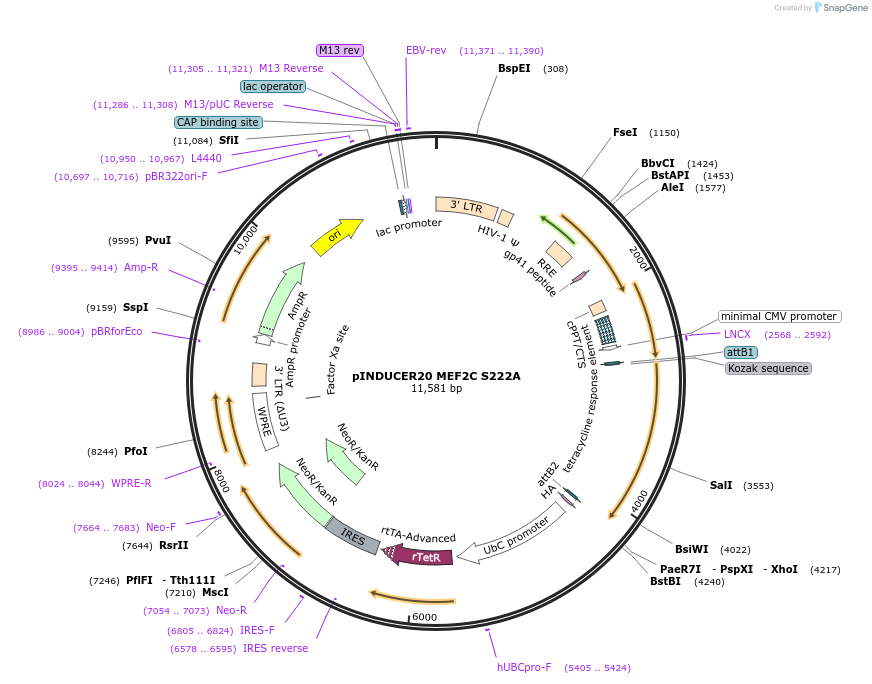
pINDUCER20 MEF2C S222A
Plasmid#89718PurposeLentiviral gene expression vector for doxycycline inducible MEF2C-S222A expressionDepositorInsertMEF2C (MEF2C Human)
UseLentiviralExpressionMammalianMutationChanged serine 222 to alanineAvailable SinceMay 23, 2017AvailabilityAcademic Institutions and Nonprofits only -
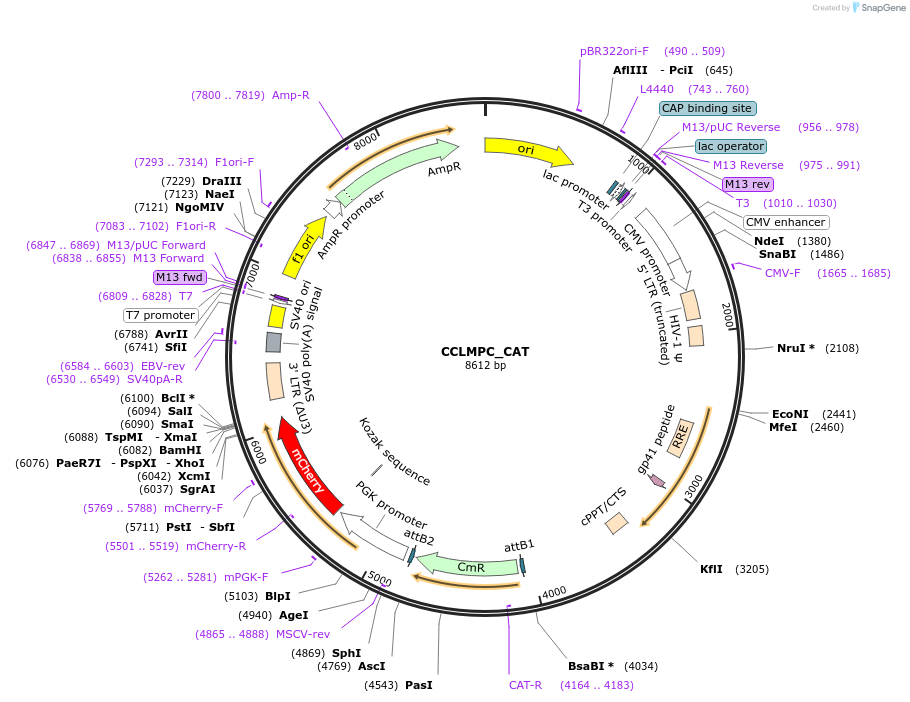
CCLMPC_CAT
Plasmid#237500PurposeDual promoter lentiviral expression control vectorDepositorInsertChloramphenicol acetyl transferase fusion protein
UseLentiviralExpressionMammalianPromoterMND/PGKAvailable SinceFeb. 2, 2026AvailabilityAcademic Institutions and Nonprofits only -
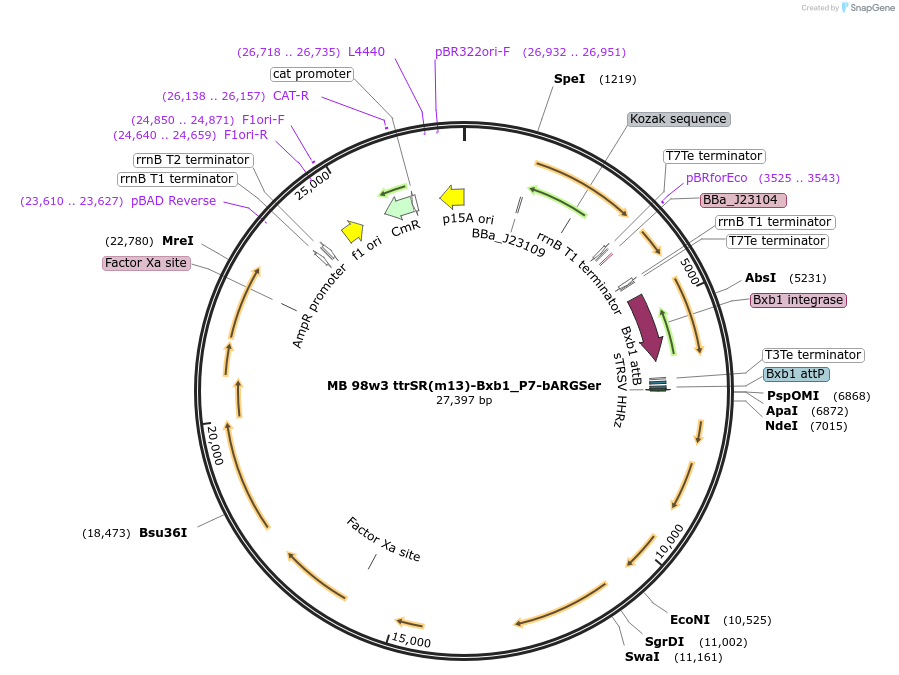
MB 98w3 ttrSR(m13)-Bxb1_P7-bARGSer
Plasmid#232473PurposeOptimized tetrathionate sensor with recombinase switch and acoustic reporter genes (bARGSer)DepositorInsertsttrS
ttrR
PttrB185-269
bARGSer
AxeTxe
Bxb1 integrase
TagsssrA degradation tagExpressionBacterialMutationthe gene Ser39006_001280 was deleted from the clu…PromoterConstitutive (TTGATAGCTAGCTCAGTCCTAGGTATTGTGCTAGC…Available SinceSept. 17, 2025AvailabilityAcademic Institutions and Nonprofits only -
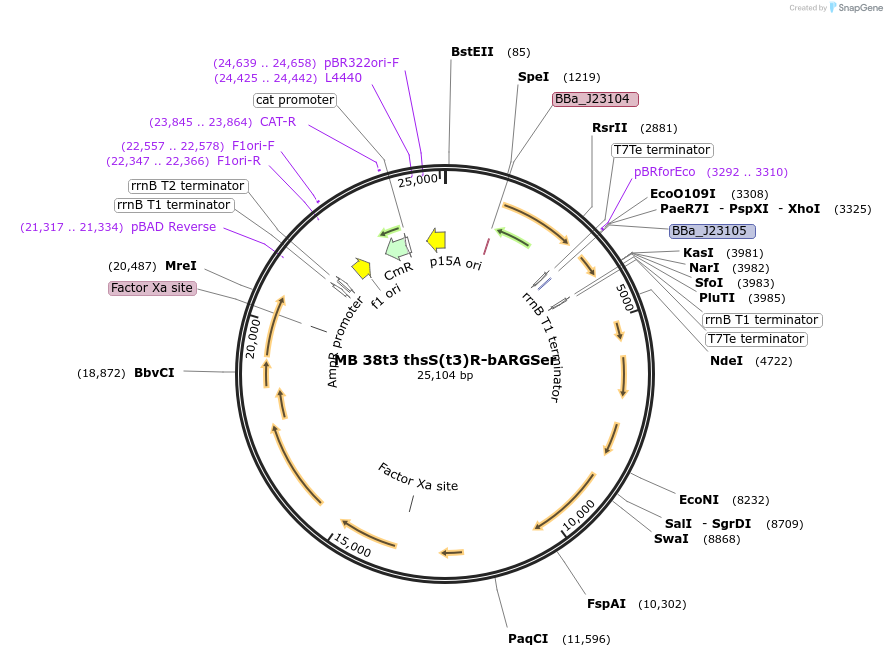
MB 38t3 thsS(t3)R-bARGSer
Plasmid#232468PurposeOptimized thiosulfate sensor with acoustic reporter genes (bARGSer)DepositorInsertsthsS(t3)
thsR
PphsA342
bARGSer
AxeTxe
ExpressionBacterialMutationContains the following mutations K286Q, Q350H, I…PromoterJ23104 (ttgacagctagctcagtcctaggtattgtgctagc), J23…Available SinceSept. 17, 2025AvailabilityAcademic Institutions and Nonprofits only -
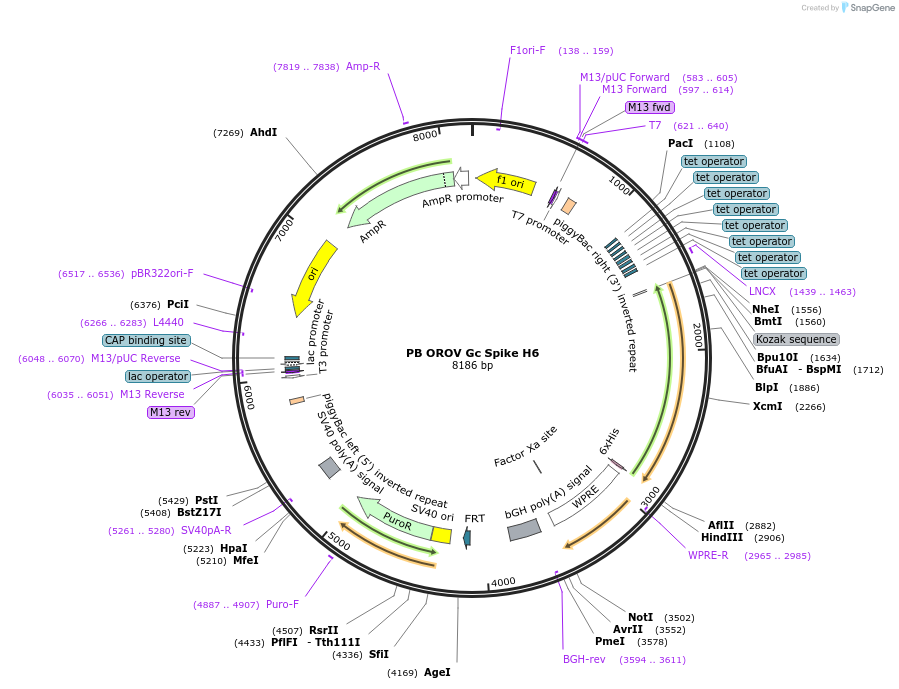
PB OROV Gc Spike H6
Plasmid#229662PurposeMake PiggyBac stable cell line expressing Oropouche virus BeAn 19991 Gc spike region (residues 482–894 of M polyprotein) with C-terminal His6 tagDepositorInsertOROV Gc spike
UsePiggybacTags6xHisExpressionMammalianMutationcontains residues 482-894 onlyPromoterTREAvailable SinceAug. 19, 2025AvailabilityAcademic Institutions and Nonprofits only -
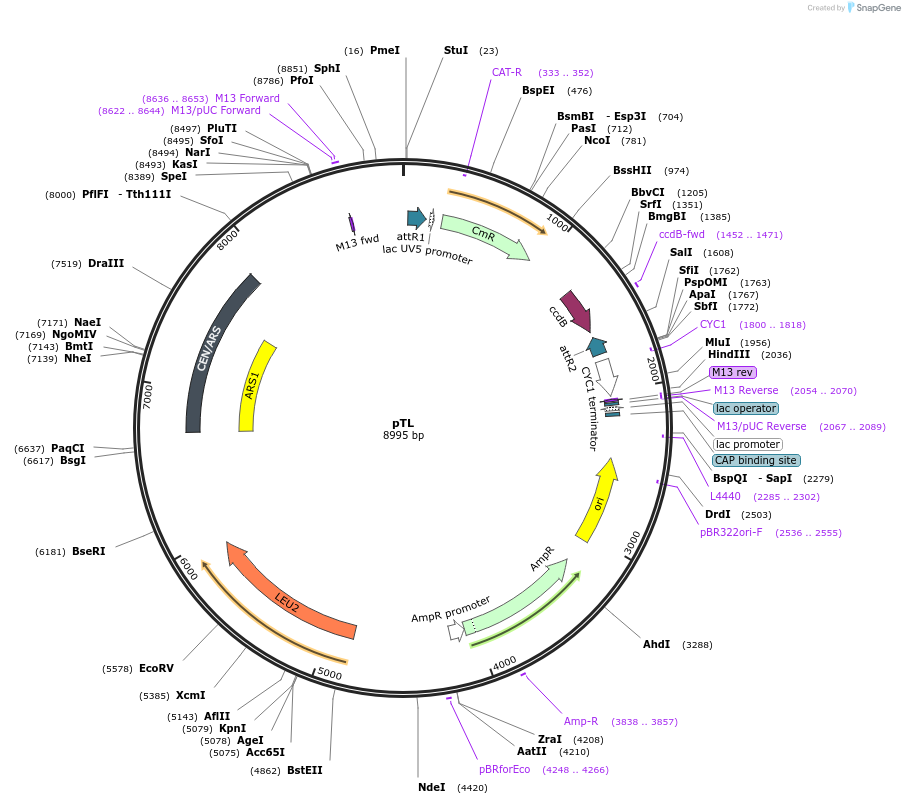
pTL
Plasmid#203176PurposeCentromeric yeast expression vector, leucine selectionDepositorTypeEmpty backboneExpressionYeastAvailable SinceAug. 29, 2023AvailabilityAcademic Institutions and Nonprofits only -
pSems-leader-SNAPf-mXFPm-IFNAR1(28-557)
Plasmid#192784PurposeExpression of SNAPf and mXFPm-tagged IFNAR1 at the plasma membrane, e.g. for single molecule fluorescence microscopyDepositorInsertIFNAR1 (IFNAR1 Human)
TagsIg k-chain leader sequence, SNAPf-tag, and mXFPmExpressionMammalianMutationmXFPm: tryptophan 66 to phenylalanine, gluatmic a…PromoterCMVAvailable SinceDec. 21, 2022AvailabilityAcademic Institutions and Nonprofits only -
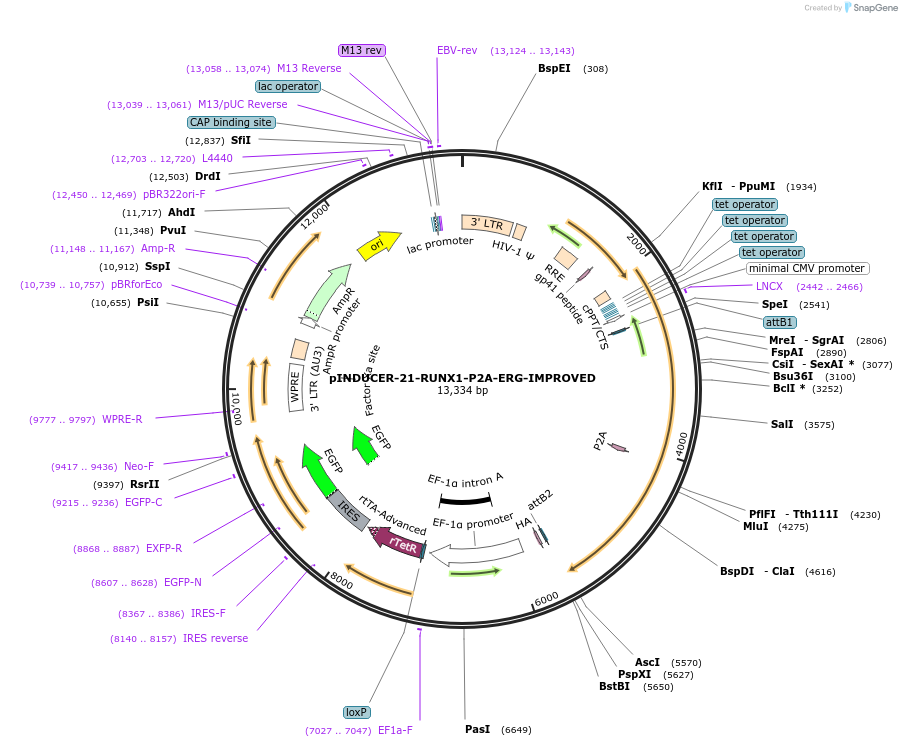
pINDUCER-21-RUNX1-P2A-ERG-IMPROVED
Plasmid#154089PurposeImproved version to express RUNX1, P2A, ERG in mammalian cells.DepositorUseLentiviralExpressionMammalianPromoterTet onAvailable SinceMay 28, 2021AvailabilityAcademic Institutions and Nonprofits only -
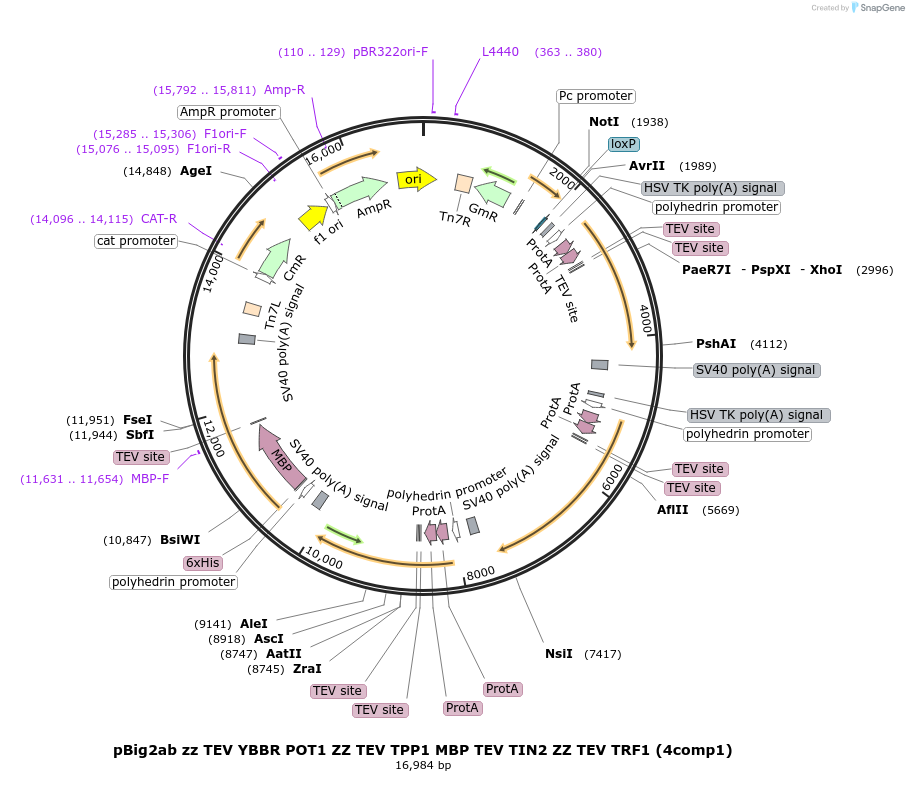
pBig2ab zz TEV YBBR POT1 ZZ TEV TPP1 MBP TEV TIN2 ZZ TEV TRF1 (4comp1)
Plasmid#185447PurposeCoexpresses human POT1 with a zz affinity tag, YBBR site, and TEV site, human TRF1 and TPP1 each with a zz tag and TEV site, and human TIN2 with an MBP affinity tag and TEV site in insect cellsDepositorTagsMBP, YBBR, and ZZExpressionInsectAvailable SinceJune 22, 2022AvailabilityAcademic Institutions and Nonprofits only -
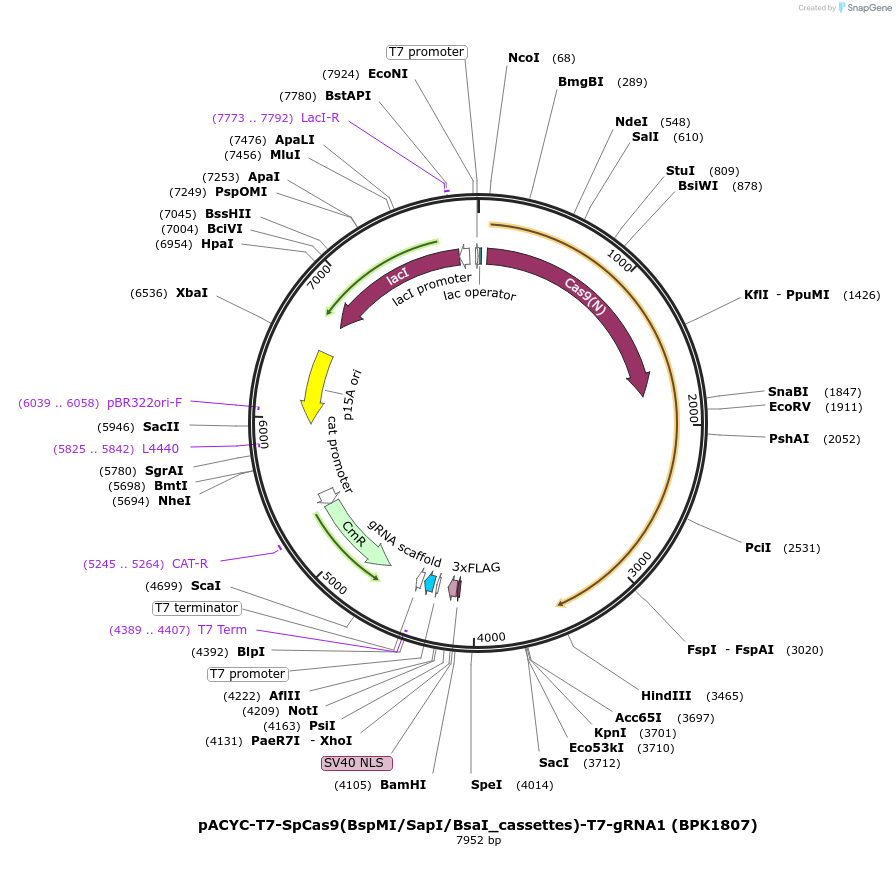
pACYC-T7-SpCas9(BspMI/SapI/BsaI_cassettes)-T7-gRNA1 (BPK1807)
Plasmid#223065PurposeDual pT7 entry plasmid for SpCas9(6AA) library; bacterial expression plasmid with type IIS RE cassettes around D1135/S1136, G1218/E1219, R1335/T1337 (precursor to MMW94). Expresses a gRNA.DepositorInsertentry vector for human codon opt. bacterial expr. plasmid for SpCas9(6AA_NNS) library, with gRNA targeting EGFP site 1
UseCRISPRTagsNLS(SV40)-3xFLAGExpressionBacterialMutationThree regions of SpCas9 encode type IIS restricti…PromoterDual T7Available SinceApril 23, 2025AvailabilityAcademic Institutions and Nonprofits only -
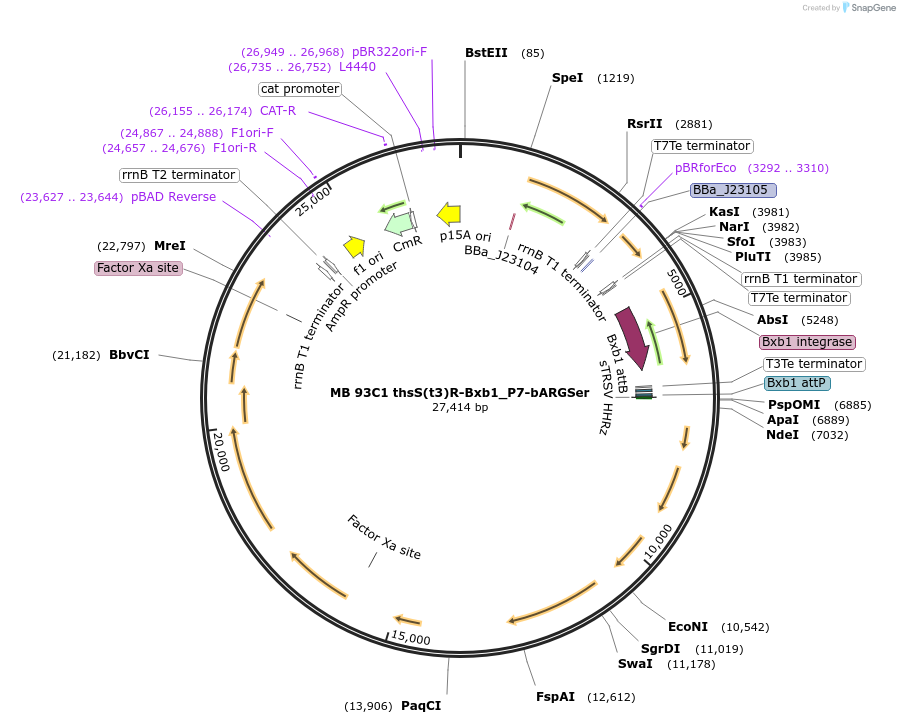
MB 93C1 thsS(t3)R-Bxb1_P7-bARGSer
Plasmid#232469PurposeOptimized thiosulfate sensor with recombinase switch and acoustic reporter genes (bARGSer)DepositorInsertsthsS(t3)
thsR
PphsA342
bARGSer
AxeTxe
Bxb1 integrase
TagsssrA degradation tagExpressionBacterialMutationContains the following mutations K286Q, Q350H, I…PromoterJ23104 (ttgacagctagctcagtcctaggtattgtgctagc), J23…Available SinceSept. 17, 2025AvailabilityAcademic Institutions and Nonprofits only -
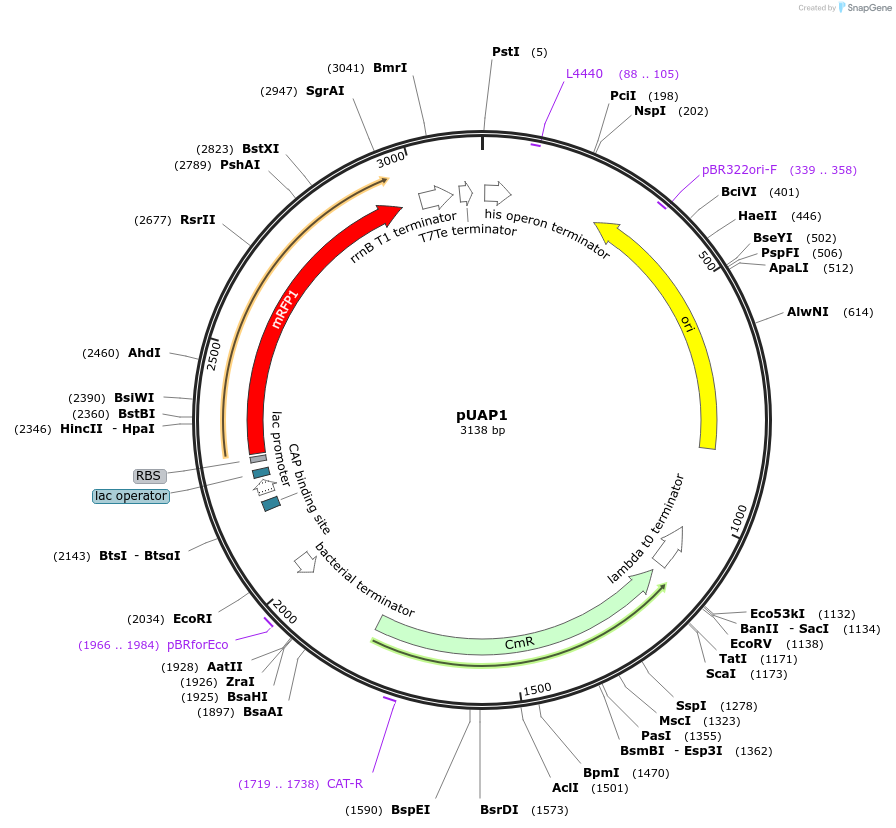
pUAP1
Plasmid#63674PurposeAccepts DNA sequences via BpiI cloning sites, resulting in Level 0 standard parts for Golden Gate cloning. Parts can be released with a user-defined 4-base-pair, 5 prime overhangs using BsaI.DepositorInsertRFP cloning selection cassette
UseSynthetic BiologyAvailable SinceApril 24, 2015AvailabilityAcademic Institutions and Nonprofits only -
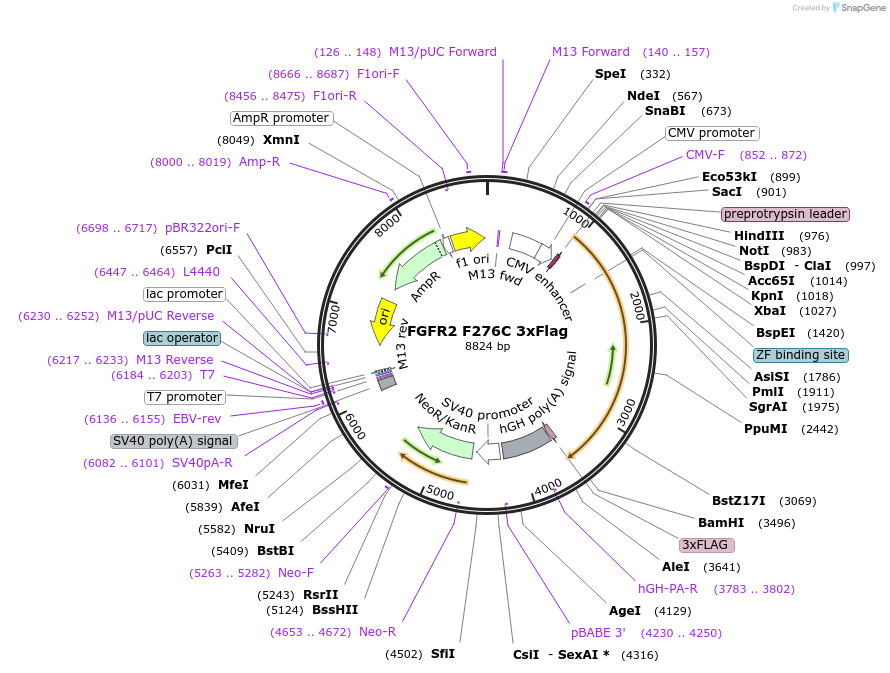
FGFR2 F276C 3xFlag
Plasmid#110064PurposeFGFR2 F276C expression in mammalian cells (JCOPO paper)DepositorInsertfibroblast growth factor receptor 2 (FGFR2 Human)
TagsFlagExpressionMammalianMutationmutated Phenylalanine 276 to Cysteine (F276C)PromoterCMVAvailable SinceMay 3, 2018AvailabilityAcademic Institutions and Nonprofits only -
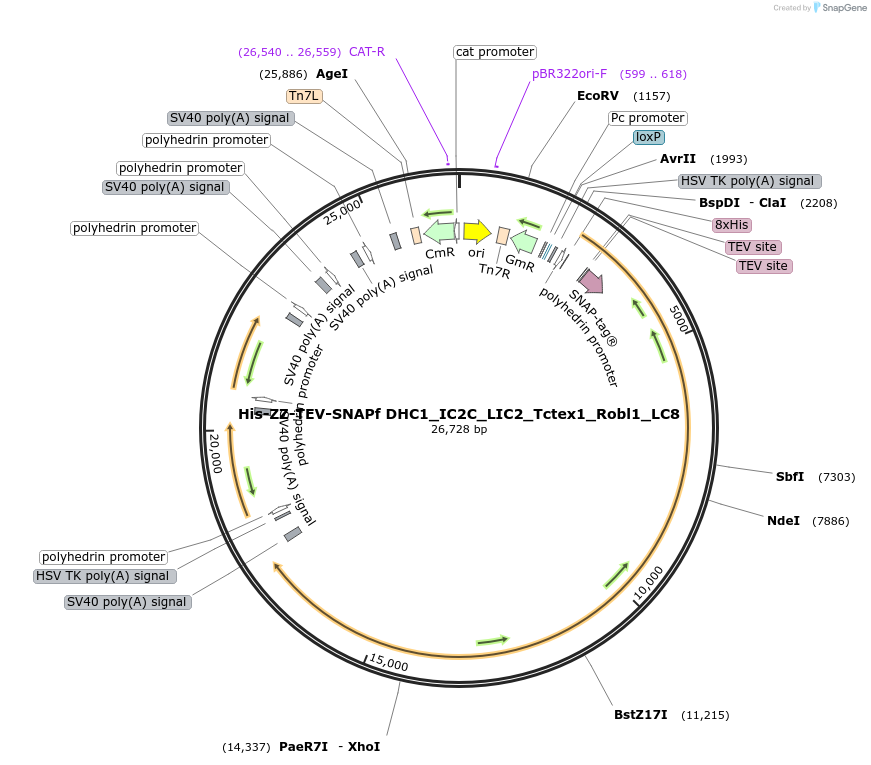
His-ZZ-TEV-SNAPf DHC1_IC2C_LIC2_Tctex1_Robl1_LC8
Plasmid#111903PurposeFull length human cytoplasmic dynein 1 complex, optimised for Sf9 expression. 8xHis-ZZ tag followed by TEV cleavage site and SNAPf tag on the N-terminus of dynein heavy chain 1.DepositorInsertsTags8xHis, SNAPf-tag, TEV site, and ZZ-tagExpressionInsectMutationOptimised for expression in Sf9 cells.Available SinceJuly 30, 2018AvailabilityAcademic Institutions and Nonprofits only -
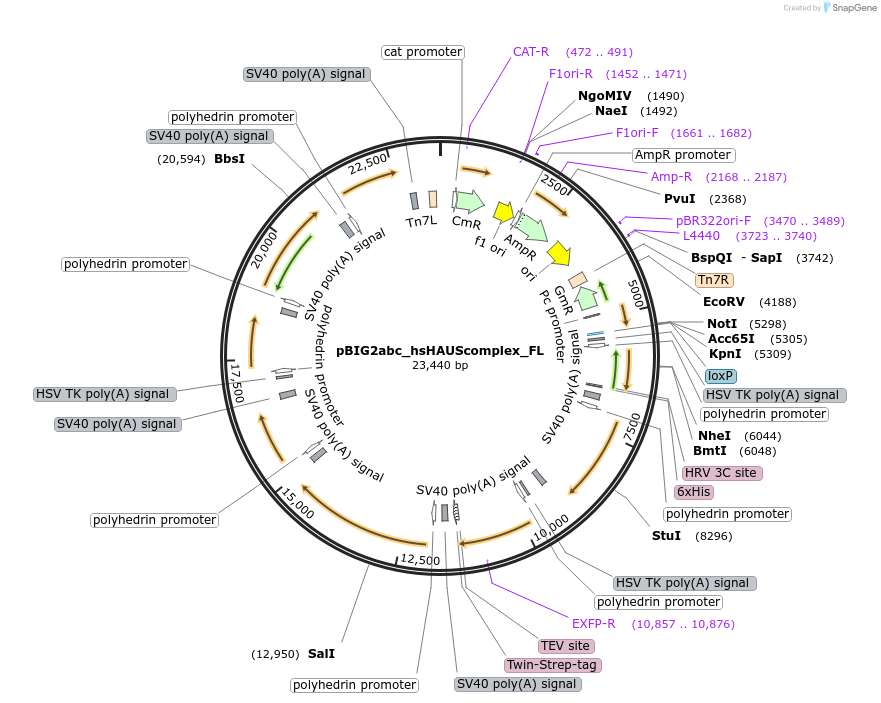
pBIG2abc_hsHAUScomplex_FL
Plasmid#202203PurposeBaculoviral transfer vector to co-express 8 subunits of human augmin complexDepositorInsertsHAUS1 (HAUS1 codon-optimized form for expression in Trichoplusia ni cells, Human)
HAUS3 (HAUS3 codon-optimized form for expression in Trichoplusia ni cells, Human)
HAUS2 (HAUS2 codon-optimized form for expression in Trichoplusia ni cells, Human)
HAUS6 (HAUS6 codon-optimized form for expression in Trichoplusia ni cells, Human)
HAUS7 (HAUS7 codon-optimized form for expression in Trichoplusia ni cells, Human)
HAUS4 (HAUS4 codon-optimized form for expression in Trichoplusia ni cells, Human)
HAUS5 (HAUS5 codon-optimized form for expression in Trichoplusia ni cells, Human)
HAUS8 (HAUS8 codon-optimized form for expression in Trichoplusia ni cells, Human)
Tags-mGFP-TEV-TwinStrep and His-tag (6x)ExpressionInsectPromoterpolyhedrin promoterAvailable SinceSept. 19, 2023AvailabilityAcademic Institutions and Nonprofits only -
MK01
Bacterial Strain#195090PurposeImproved E. coli strain created by knocking out the endogenous lacI gene with chloramphenicol acetyltransferase (cat) selection cassette in the parent strain BW27783 that overcome the all-or-none response of araBAD promoter (PBAD).DepositorBacterial ResistanceChloramphenicolAvailable SinceFeb. 6, 2023AvailabilityAcademic Institutions and Nonprofits only -
HL 1255
Bacterial Strain#52941PurposeMG1655 + fimS (off orientation)::gfp::Asp terminator::CamR at fimADepositorBacterial ResistanceChloramphenicolSpeciesEscherichia coliAvailable SinceApril 15, 2014AvailabilityAcademic Institutions and Nonprofits only -
C321.∆A.opt
Bacterial Strain#87359PurposeThis is C321.∆A with additional mutations engineered by Kuznetsov et al. to improve doubling time.DepositorBacterial ResistanceGentamicinSpeciesSyntheticAvailable SinceMay 1, 2017AvailabilityAcademic Institutions and Nonprofits only -
UQ5853
Bacterial Strain#37159DepositorBacterial ResistanceChloramphenicolAvailable SinceJuly 30, 2012AvailabilityAcademic Institutions and Nonprofits only -
EcNR2 Strain
Bacterial Strain#26931DepositorBacterial ResistanceChloramphenicol and AmpicillinAvailable SinceJan. 28, 2011AvailabilityAcademic Institutions and Nonprofits only -
HL 932
Bacterial Strain#52940PurposeMG1655 + ΔlacIDepositorBacterial ResistanceNoneSpeciesEscherichia coliAvailable SinceApril 15, 2014AvailabilityAcademic Institutions and Nonprofits only -
CY007
Bacterial Strain#72337PurposeBacterial strain: E. coli BW25113 ΔlacI ΔaraC ΔsdiADepositorBacterial ResistanceChloramphenicolAvailable SinceMarch 10, 2016AvailabilityAcademic Institutions and Nonprofits only -
HL 4233
Bacterial Strain#52944PurposeΔfimB + ΔfimE + ΔfimADepositorBacterial ResistanceNoneSpeciesEscherichia coliAvailable SinceApril 15, 2014AvailabilityAcademic Institutions and Nonprofits only -
HL 1285
Bacterial Strain#52942PurposeMG1655 + fimS (off orientation)::gfp::Asp terminator at fimADepositorBacterial ResistanceNoneSpeciesEscherichia coliAvailable SinceApril 15, 2014AvailabilityAcademic Institutions and Nonprofits only -
HL 4195
Bacterial Strain#52943PurposeMG1655 + fimS (off orientation)::gfp::Asp terminator at fimA + ΔlacI + KanR replaces fimB and fimE(partial)DepositorBacterial ResistanceKanamycinSpeciesEscherichia coliAvailable SinceApril 15, 2014AvailabilityAcademic Institutions and Nonprofits only -
HL 6139
Bacterial Strain#52946PurposeMG1655 + fimS (off orientation)::gfp::Asp terminator at fimA + ΔlacI + KanR replaces fimB and fimE(stop codon)DepositorBacterial ResistanceKanamycinSpeciesEscherichia coliAvailable SinceApril 15, 2014AvailabilityAcademic Institutions and Nonprofits only -
HL 6279
Bacterial Strain#52949PurposeMG1655 + fimS (off orientation)::gfpAAV tag::Asp terminator::CamR at fimA + ΔlacI + KanR replaces fimB and fimE(partial)DepositorBacterial ResistanceKanamycinSpeciesEscherichia coliAvailable SinceApril 15, 2014AvailabilityAcademic Institutions and Nonprofits only -
HL 6166
Bacterial Strain#52947PurposeMG1655 + fimS (off orientation)::gfp::Asp terminator at fimA + ΔlacI + KanR replaces fimB and fimE(IRL)DepositorBacterial ResistanceKanamycinSpeciesEscherichia coliAvailable SinceApril 15, 2014AvailabilityAcademic Institutions and Nonprofits only