We narrowed to 10,327 results for: Coli
-
Plasmid#234212PurposeHeterologous protein expression of Wufb25h12 in Escherichia coliDepositorAvailable SinceMarch 6, 2025AvailabilityIndustry, Academic Institutions, and Nonprofits
-
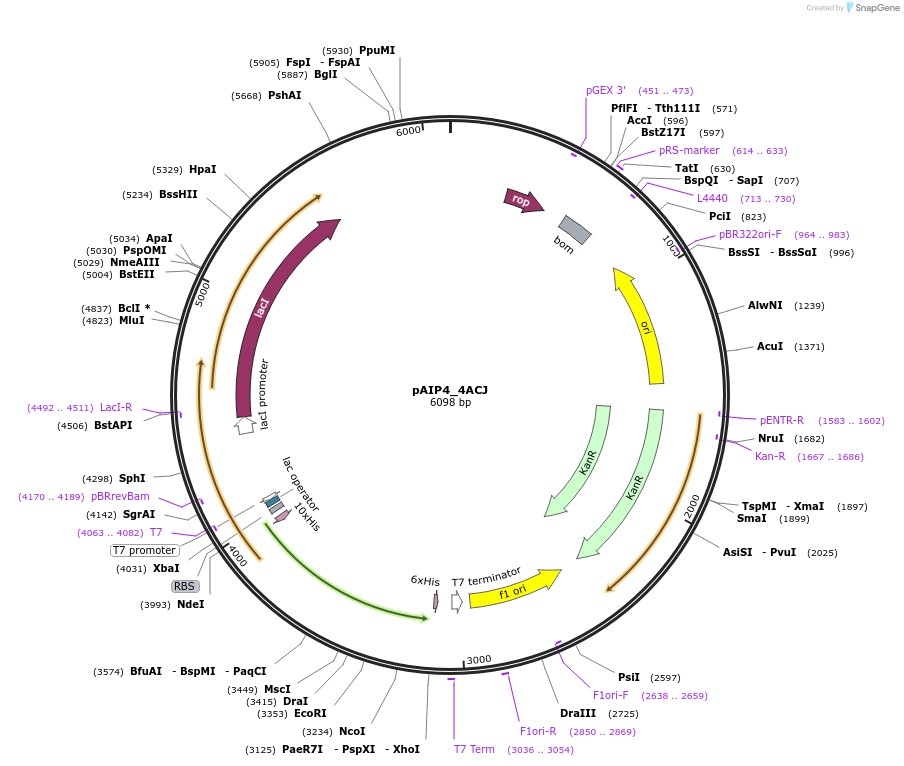
pAIP4_1EYH
Plasmid#234159PurposeHeterologous protein expression of epsin-1 (EPS-15-interacting protein 1) in Escherichia coliDepositorAvailable SinceMarch 6, 2025AvailabilityIndustry, Academic Institutions, and Nonprofits -
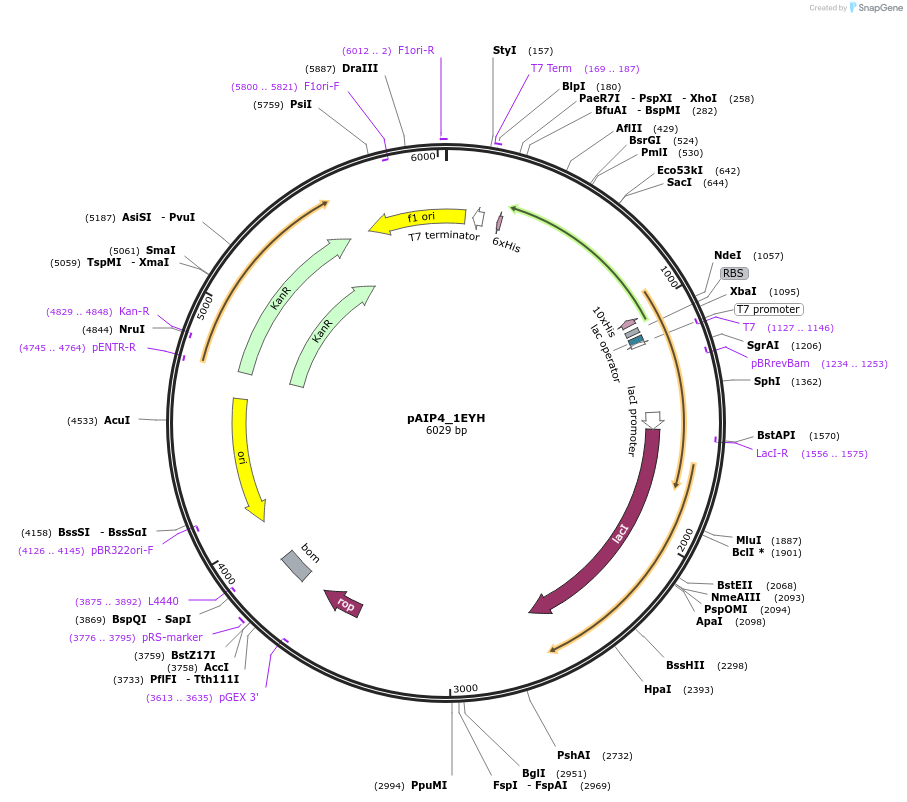
pET28a_HCK
Plasmid#224239PurposeExpression of human HCK Kinase in E. coliDepositorAvailable SinceJan. 24, 2025AvailabilityAcademic Institutions and Nonprofits only -
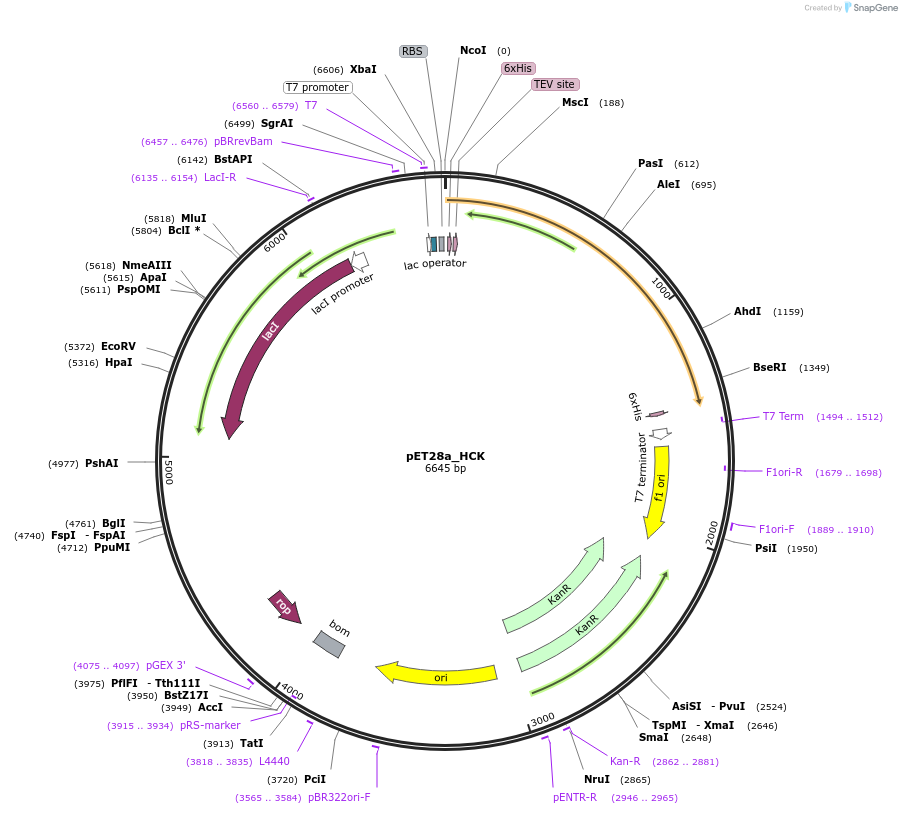
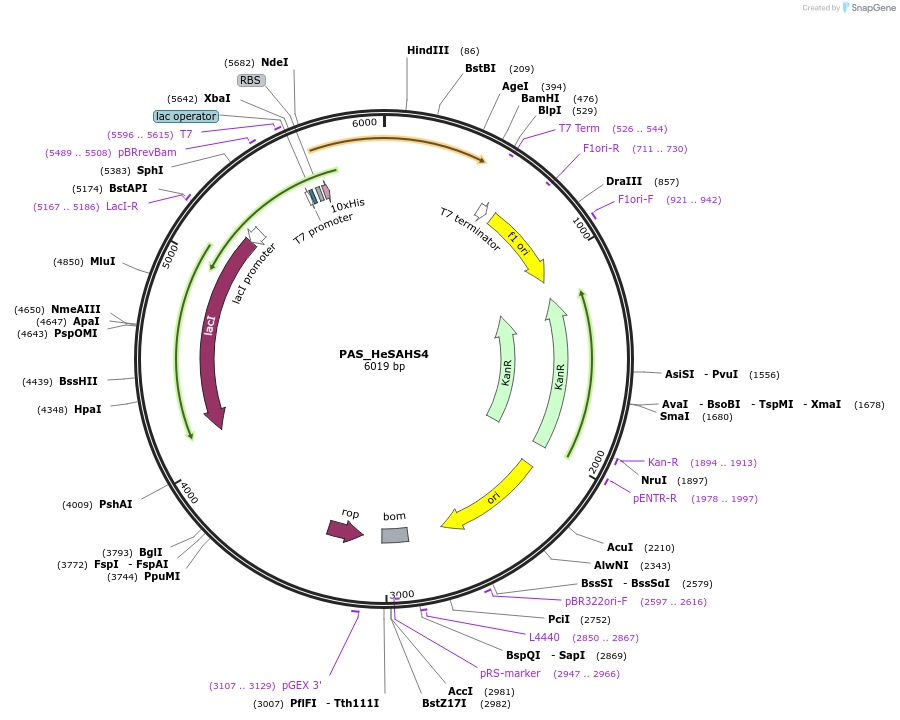
PAS_HeSAHS4
Plasmid#215648PurposeExpresses SUMO-tagged HeSAHS4 protein from tardigrade H. exemplaris in E. coliDepositorInsertSAHS4
TagsSUMOExpressionBacterialAvailable SinceMarch 19, 2024AvailabilityAcademic Institutions and Nonprofits only -
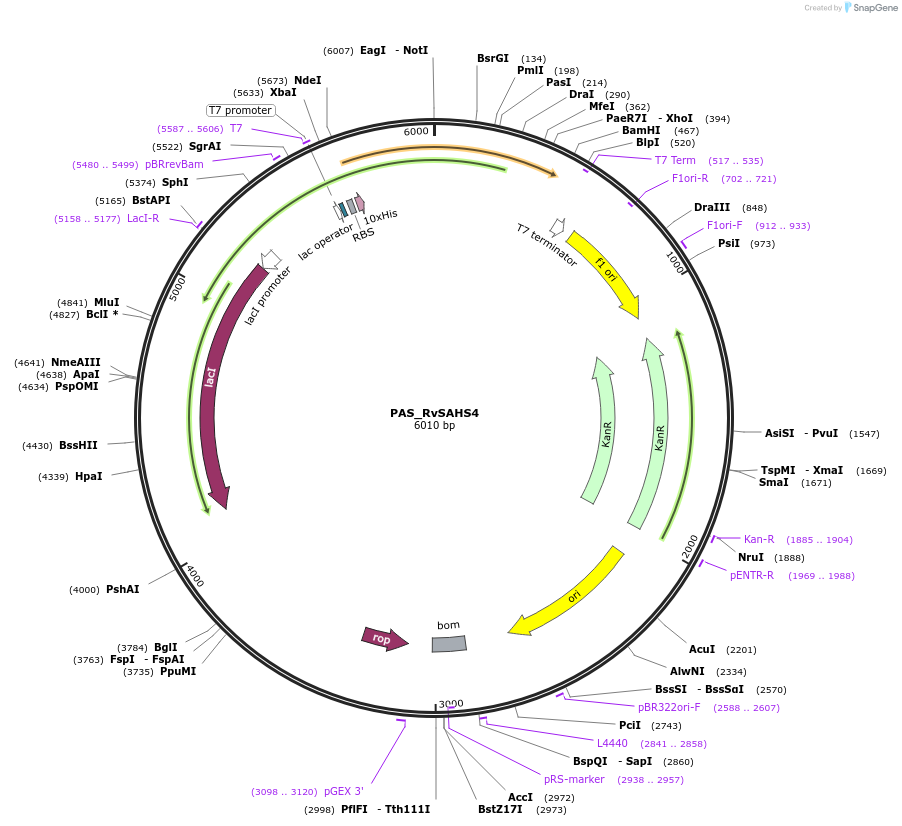
PAS_RvSAHS4
Plasmid#215646PurposeExpresses SUMO-tagged RvSAHS4 protein from tardigrade R. varieornatus in E. coliDepositorInsertSAHS4
TagsSUMOExpressionBacterialAvailable SinceMarch 19, 2024AvailabilityAcademic Institutions and Nonprofits only -
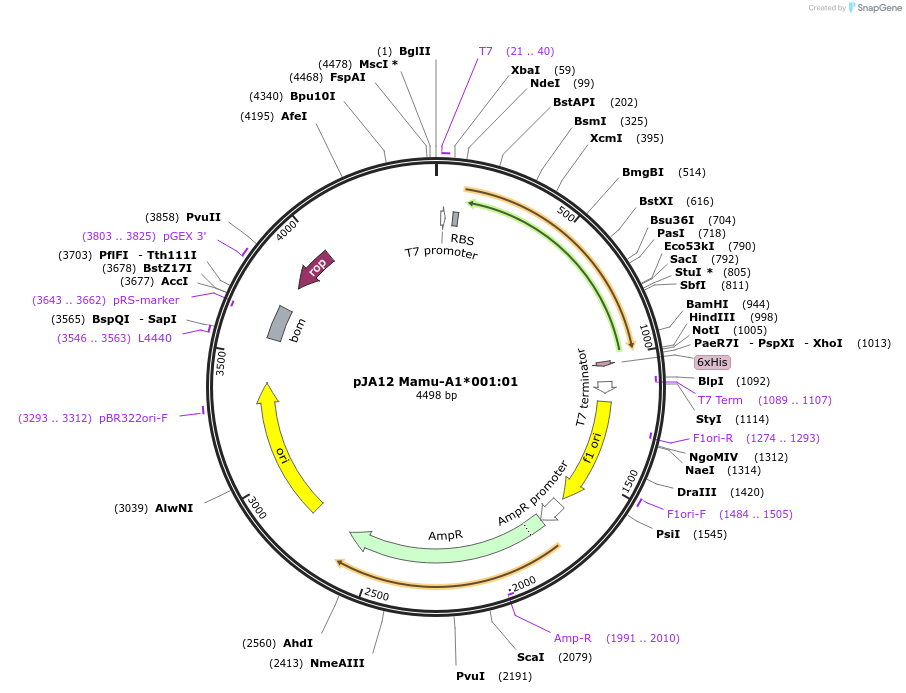
pJA12 Mamu-A1*001:01
Plasmid#180442PurposeE. coli expression of soluble MHC proteinDepositorInsertMamu-A1*001:01
TagsBSP41ExpressionBacterialMutationReplacement of transmembrane domain with enzymati…Available SinceApril 19, 2022AvailabilityAcademic Institutions and Nonprofits only -
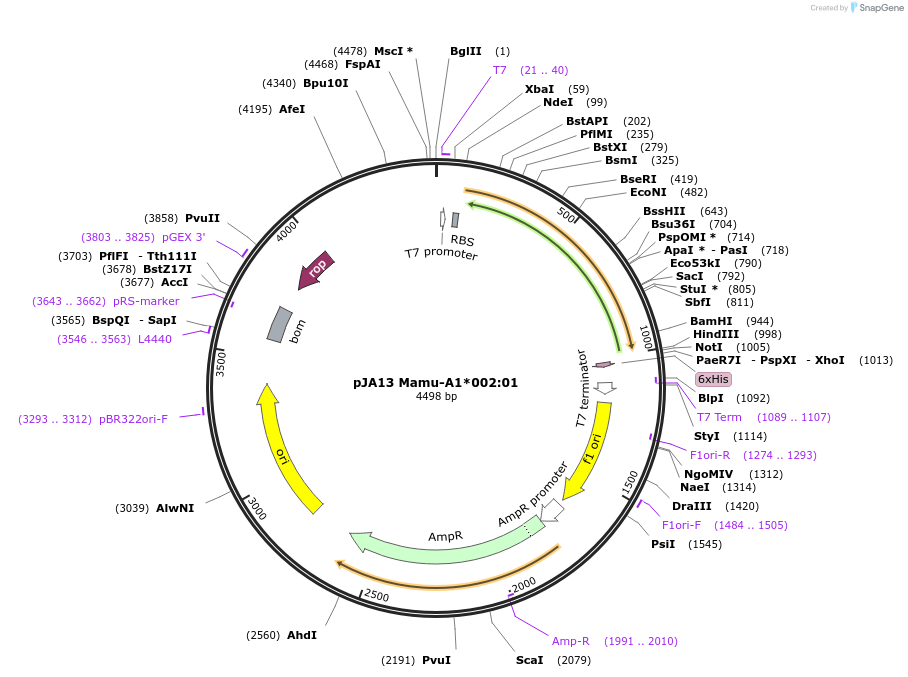
pJA13 Mamu-A1*002:01
Plasmid#180443PurposeE. coli expression of soluble MHC proteinDepositorInsertMamu-A1*002:01
TagsBSP41ExpressionBacterialMutationReplacement of transmembrane domain with enzymati…Available SinceMarch 8, 2022AvailabilityAcademic Institutions and Nonprofits only -
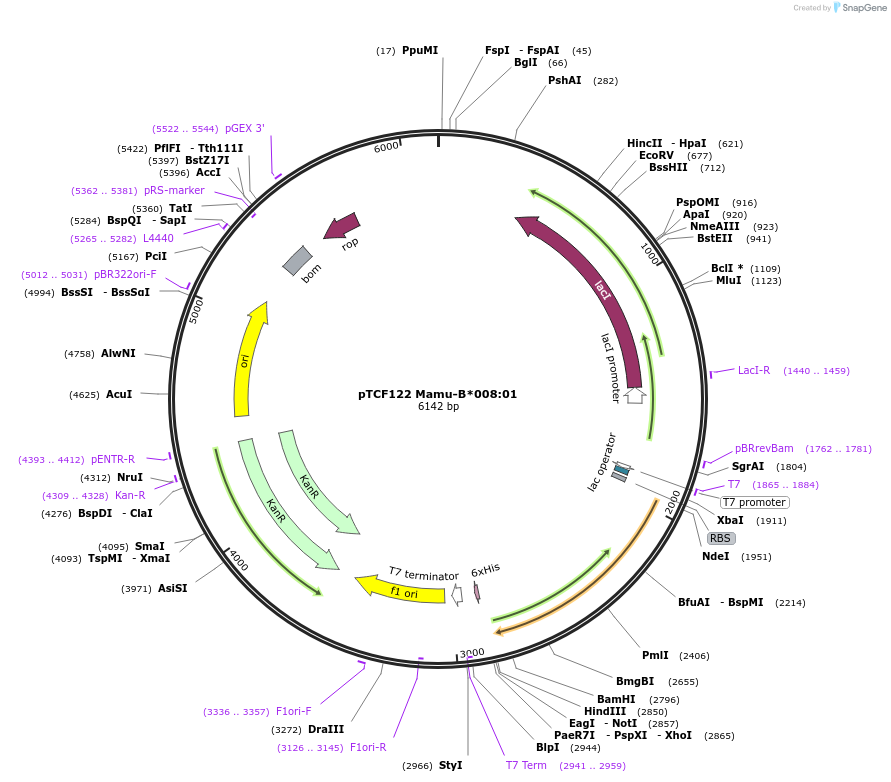
pTCF122 Mamu-B*008:01
Plasmid#180455PurposeE. coli expression of soluble MHC proteinDepositorInsertMamu-B*008:01
TagsBSP41ExpressionBacterialMutationReplacement of transmembrane domain with enzymati…Available SinceFeb. 23, 2022AvailabilityAcademic Institutions and Nonprofits only -
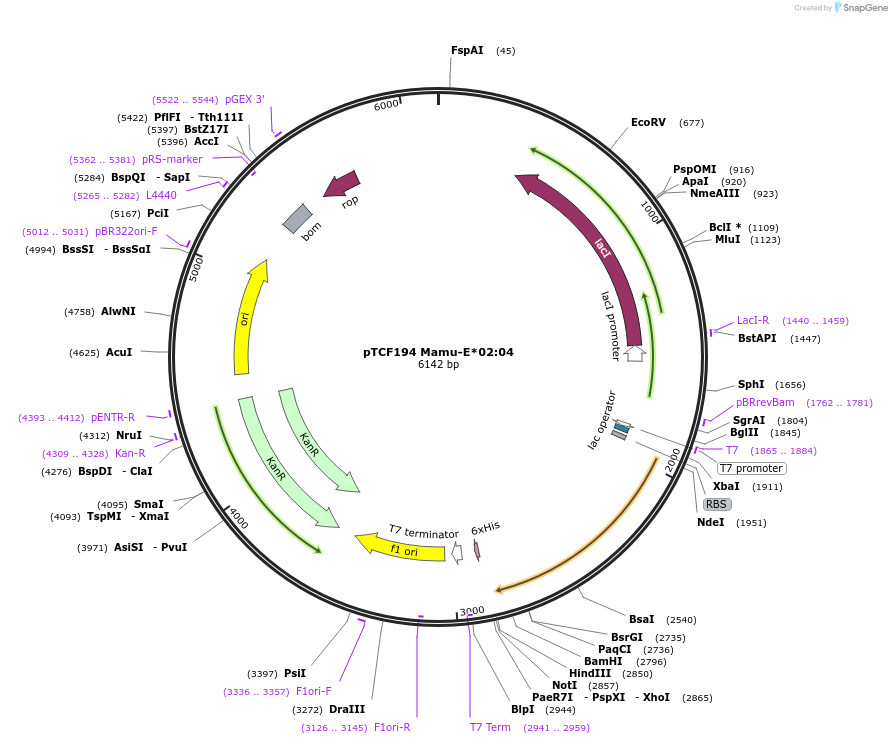
pTCF194 Mamu-E*02:04
Plasmid#180461PurposeE. coli expression of soluble MHC proteinDepositorInsertMamu-E*02:04
TagsBSP41ExpressionBacterialMutationReplacement of transmembrane domain with enzymati…Available SinceFeb. 23, 2022AvailabilityAcademic Institutions and Nonprofits only -
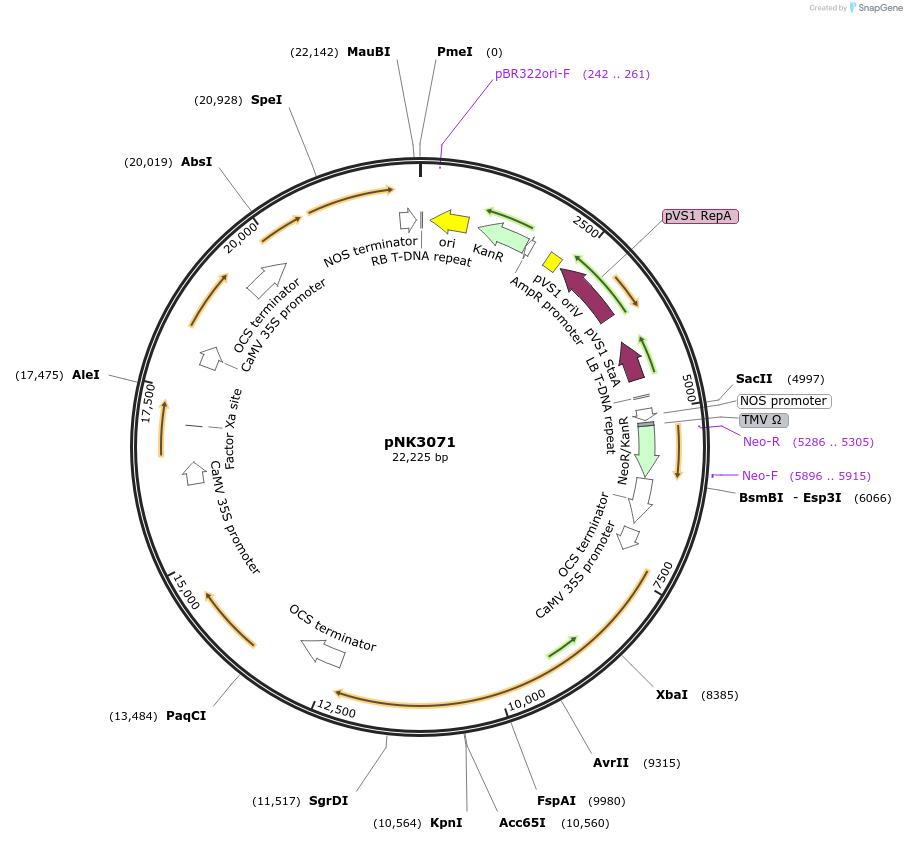
pNK3071
Plasmid#219755PurposeMoClo-compatible Level P vector for improved autonomous bioluminescence in plants encoding kanamycin resistance cassette, mcitHispS, NpgA, nnH3H_v2, nnLuz_v4 and nnCPH.DepositorInsertmcitHispS, NpgA, nnH3H_v2, nnLuz_v4 and nnCPH
ExpressionPlantAvailable SinceOct. 30, 2024AvailabilityAcademic Institutions and Nonprofits only -
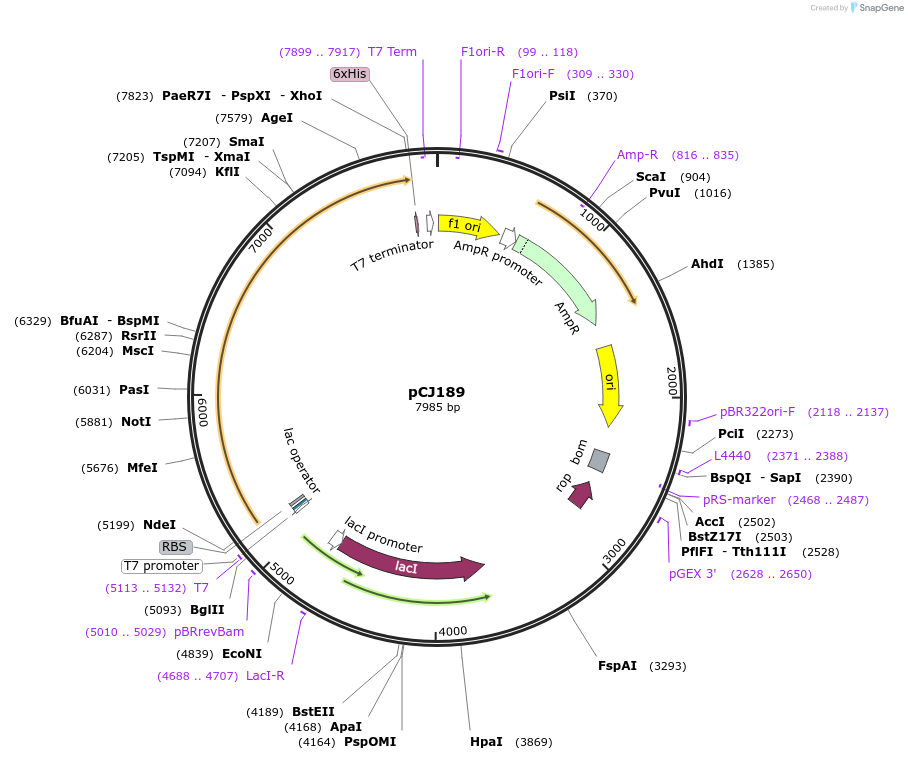
pCJ189
Plasmid#162666PurposepET-21b(+) based plasmid for expresion of MHETase (Genbank GAP38911.1) linked to PETase (Genbank GAP38373.1) from Ideonella sakaiensis by an 8 residue Gly/Ser linker; codon optimized for expression in E. coli K12, with C-terminal His tag.DepositorInsertMHETase (Genbank GAP38911.1) linked to PETase (Genbank GAP38373.1) from Ideonella sakaiensis by an 8 residue Gly/Ser linker
TagsHisExpressionBacterialMutationcodon optimized for expression in E. coli K12PromoterT7/lacAvailable SinceFeb. 3, 2021AvailabilityAcademic Institutions and Nonprofits only -
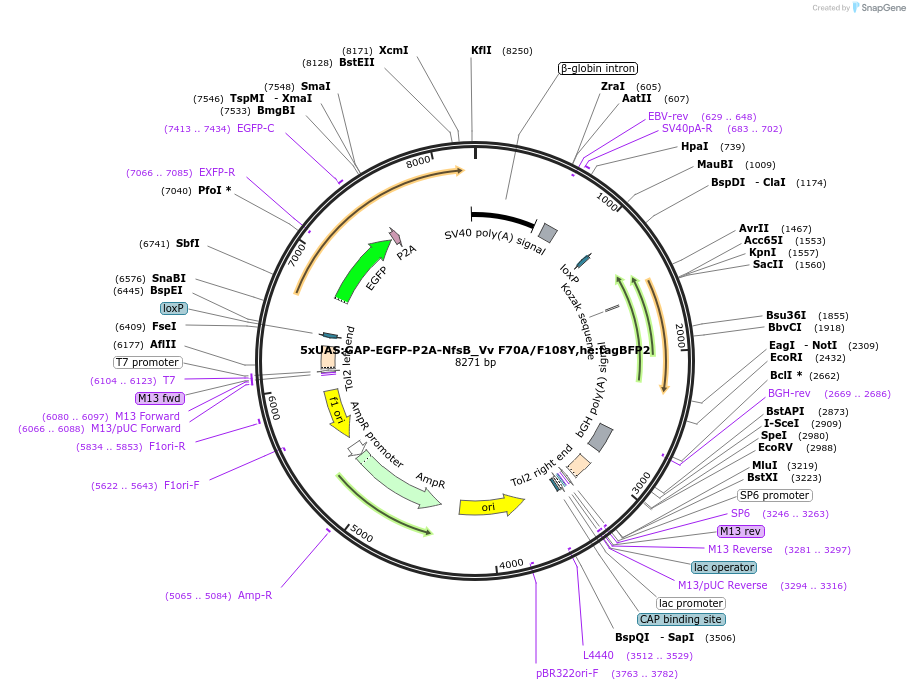
5xUAS:GAP-EGFP-P2A-NfsB_Vv F70A/F108Y,he:tagBFP2
Plasmid#158652Purposemaking transgenics expressing a UAS driven eGFP and NTR 2.0DepositorInsert5xUAS:GAP-eGFP-P2A-NfsB_Vv F70A/F108Y;he:BFP
UseTol2-based transgenic expression in zebrafish/oth…MutationF70A/F108YAvailable SinceSept. 14, 2020AvailabilityAcademic Institutions and Nonprofits only -
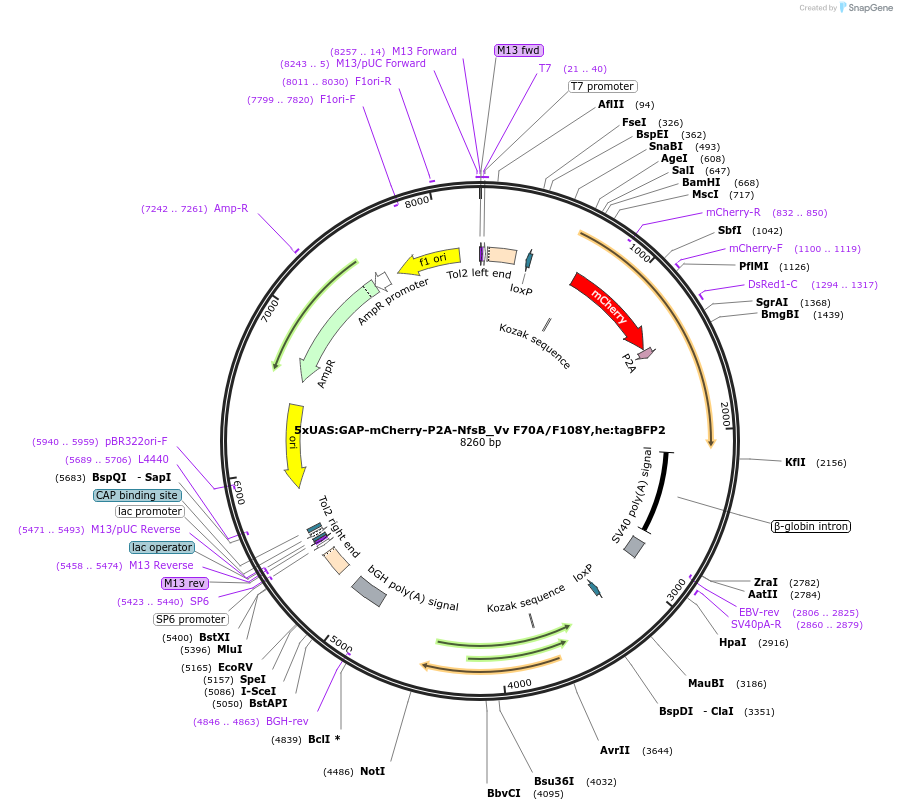
5xUAS:GAP-mCherry-P2A-NfsB_Vv F70A/F108Y,he:tagBFP2
Plasmid#158653Purposemaking transgenics expressing a UAS driven mCherry and NTR 2.0DepositorInsert5xUAS:GAP-mCherry-P2A-NfsB_Vv F70A/F108Y;he:BFP
UseTol2-based transgenic expression in zebrafish/oth…MutationF70A/F108YAvailable SinceSept. 17, 2020AvailabilityAcademic Institutions and Nonprofits only -
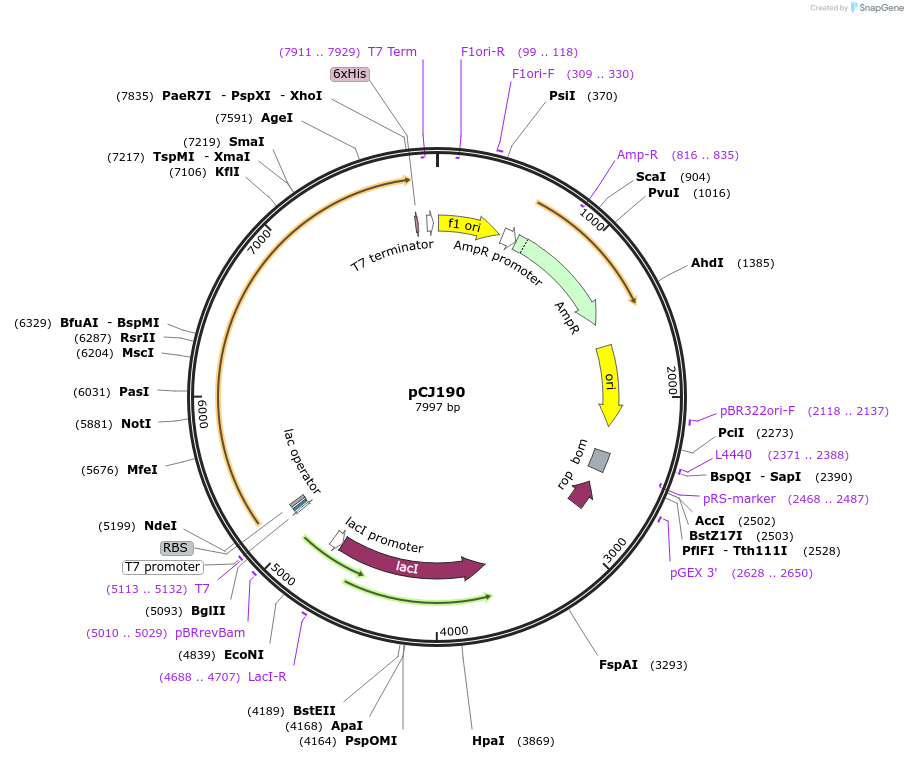
pCJ190
Plasmid#162667PurposepET-21b(+) based plasmid for expresion of MHETase (Genbank GAP38911.1) linked to PETase (Genbank GAP38373.1) from Ideonella sakaiensis by a 12 residue Gly/Ser linker; codon optimized for expression in E. coli K12, with C-terminal His tag.DepositorInsertMHETase (Genbank GAP38911.1) linked to PETase (Genbank GAP38373.1) from Ideonella sakaiensis by a 12 residue Gly/Ser linker
TagsHisExpressionBacterialMutationcodon optimized for expression in E. coli K12PromoterT7/lacAvailable SinceJan. 7, 2021AvailabilityAcademic Institutions and Nonprofits only -
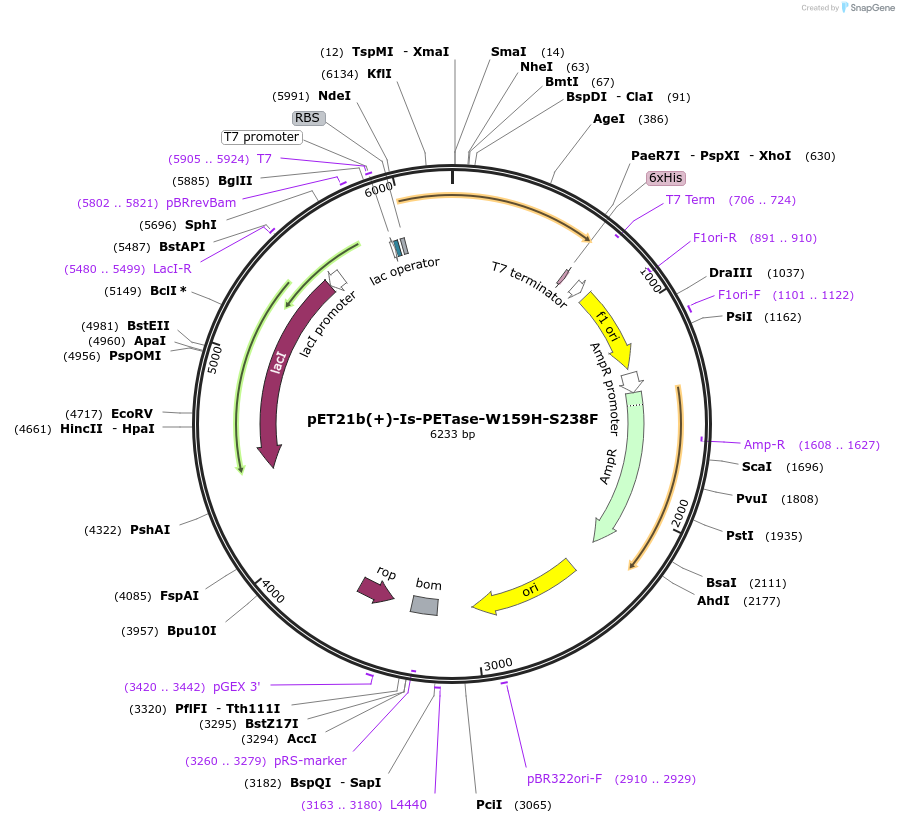
pET21b(+)-Is-PETase-W159H-S238F
Plasmid#112203PurposepET-21b(+) based plasmid for expression of PETase from Ideonella sakaiensis 201-F6 (Genbank GAP38373.1) with W159H and S238F mutations, codon optimized for expression in E. coli K12DepositorInsertPETase gene from Ideonella sakaiensis 201-F6 with W159H and S238F mutations, codon optimized for expression in E. coli K12
Tags6XHISExpressionBacterialMutationW159H, S238FPromoterN/AAvailable SinceJuly 30, 2018AvailabilityAcademic Institutions and Nonprofits only -
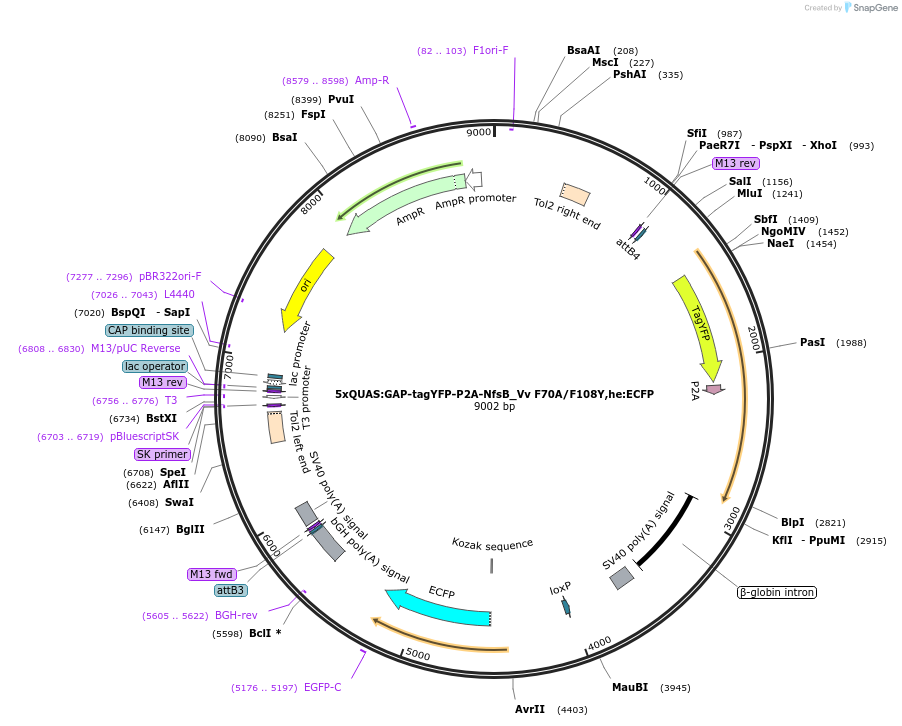
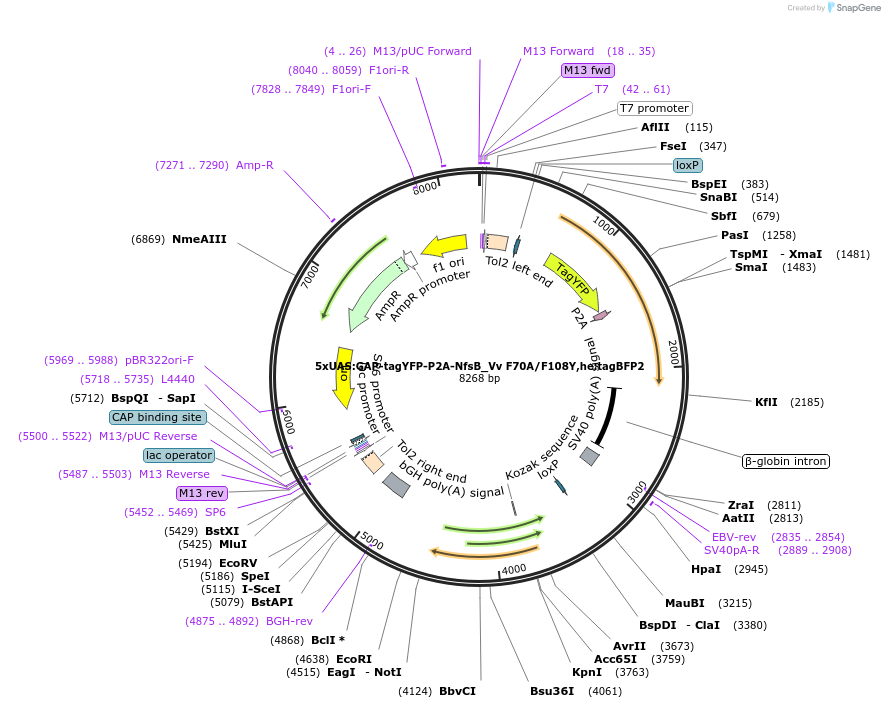
5xQUAS:GAP-tagYFP-P2A-NfsB_Vv F70A/F108Y,he:ECFP
Plasmid#158654Purposemaking transgenics expressing a QUAS driven YFP and NTR 2.0DepositorInsert5xQUAS:GAP-tagYFP-P2A-NfsB_Vv F70A/F108Y;he:eCFP
UseTol2-based transgenic expression in zebrafish/oth…MutationF70A/F108YAvailable SinceSept. 17, 2020AvailabilityAcademic Institutions and Nonprofits only -
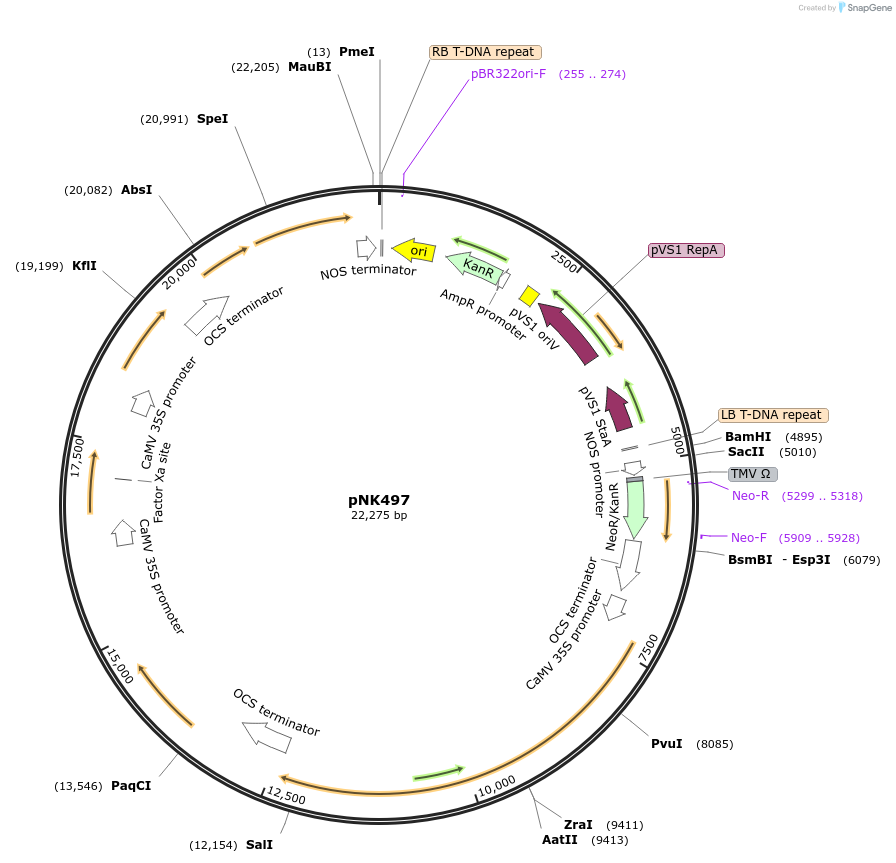
pNK497
Plasmid#219754PurposeMoClo-compatible Level P vector for improved autonomous bioluminescence in plants encoding kanamycin resistance cassette, nnHispS, NpgA, nnH3H_v2, nnLuz_v4 and nnCPH.DepositorInsertnnHispS, NpgA, nnH3H_v2, nnLuz_v4 and nnCPH
ExpressionPlantAvailable SinceJan. 21, 2025AvailabilityAcademic Institutions and Nonprofits only -
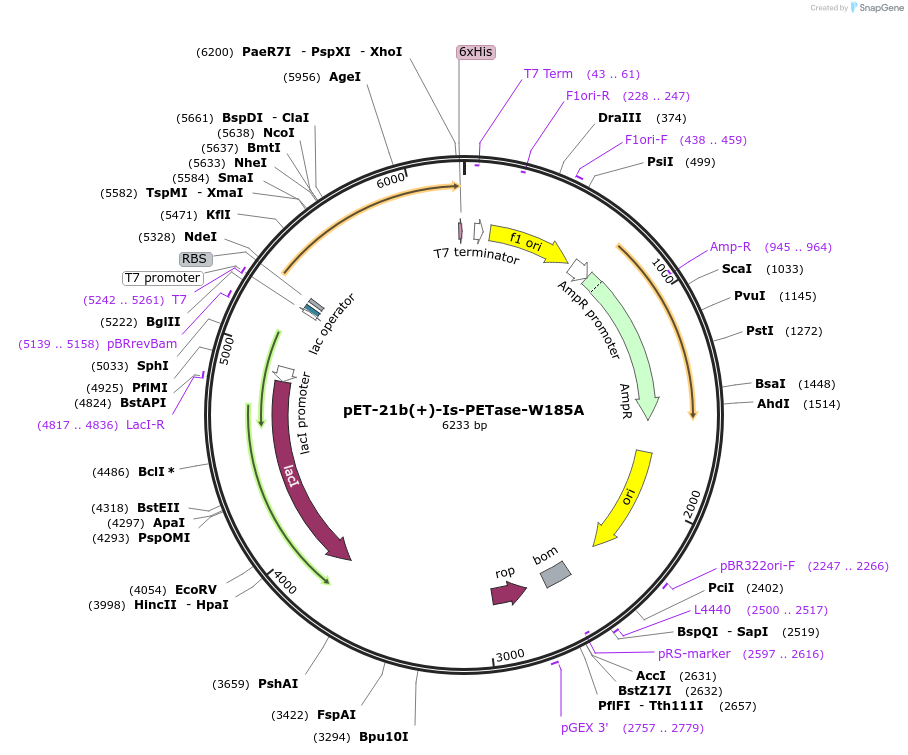
pET-21b(+)-Is-PETase-W185A
Plasmid#112204PurposepET-21b(+) based plasmid for expression of PETase from Ideonella sakaiensis 201-F6 (Genbank GAP38373.1) with W185A mutations, codon optimized for expression in E. coli K12DepositorInsertPETase gene from Ideonella sakaiensis 201-F6 with W185A mutation, codon optimized for expression in E. coli K12
Tags6XHISExpressionBacterialMutationW185APromoterN/AAvailable SinceJuly 18, 2018AvailabilityAcademic Institutions and Nonprofits only -
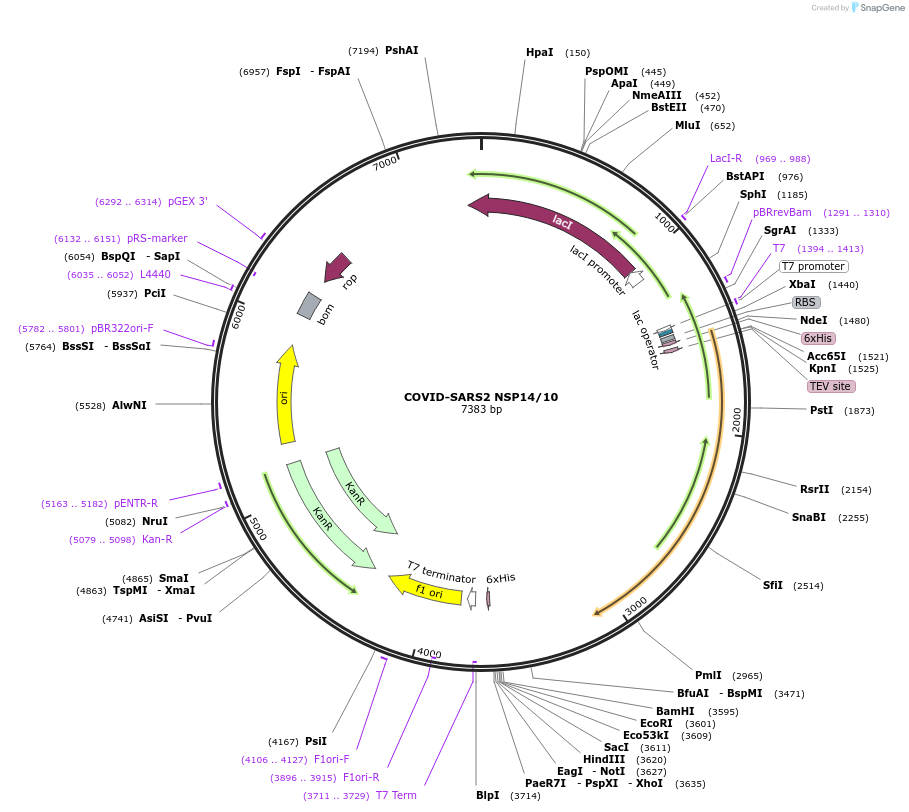
COVID-SARS2 NSP14/10
Plasmid#159613PurposeBacterial co-expression vector fo Covid-SARS2 NSP14 and NSP10DepositorInsertsTagsHis6, TEV cleavage site and noneExpressionBacterialMutationCodon-optimized for E. coli expressionPromoter(Bicistronic) and T7 - LacOAvailable SinceSept. 15, 2020AvailabilityAcademic Institutions and Nonprofits only -
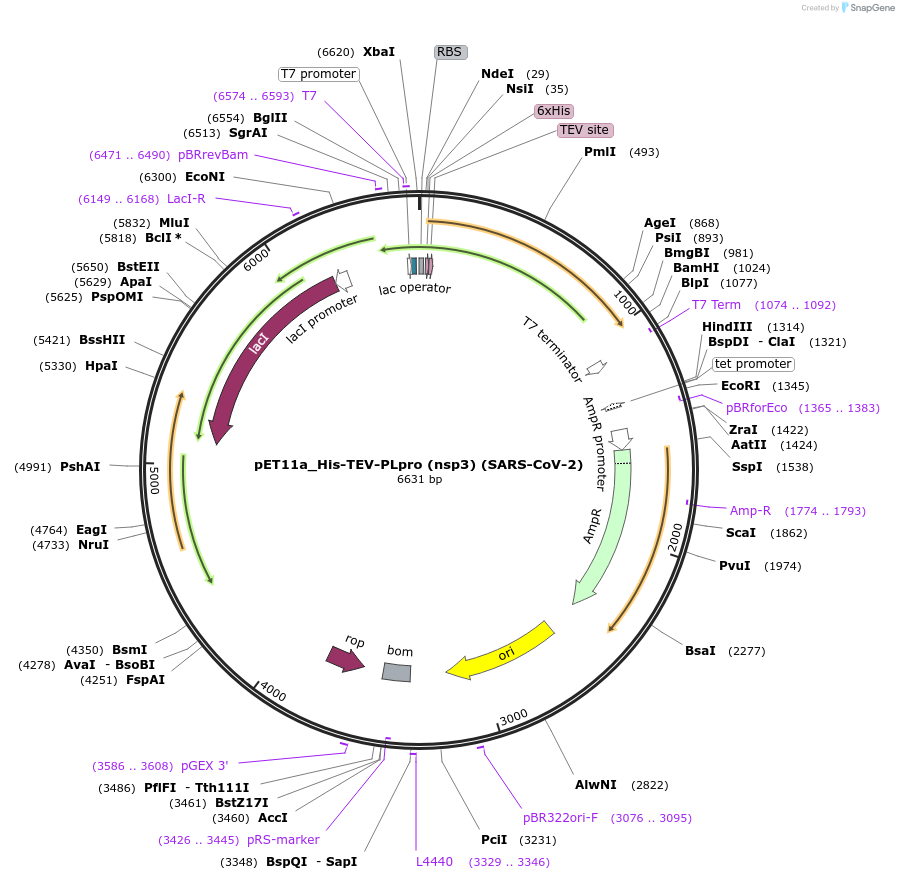
pET11a_His-TEV-PLpro (nsp3) (SARS-CoV-2)
Plasmid#169192PurposeTo express SARS-CoV-2 Papain-like protease in E. coliDepositorInsert6His-TEV-nsp3_E746-K1060 (pp1ab_E1564-K1878) (ORF1ab SARS-CoV-2, Synthetic)
UseUnspecifiedTags6His-TEVMutationCodon optimised for E. coliPromoterT7Available SinceJuly 6, 2021AvailabilityAcademic Institutions and Nonprofits only -
5xUAS:GAP-tagYFP-P2A-NfsB_Vv F70A/F108Y,he:tagBFP2
Plasmid#158651Purposemaking transgenics expressing a UAS driven YFP and NTR 2.0DepositorInsert5xUAS:GAP-tagYFP-P2A-NfsB_Vv F70A/F108Y;he:BFP
UseTol2-based transgenic expression in zebrafish/oth…MutationF70A/F108YAvailable SinceSept. 14, 2020AvailabilityAcademic Institutions and Nonprofits only -
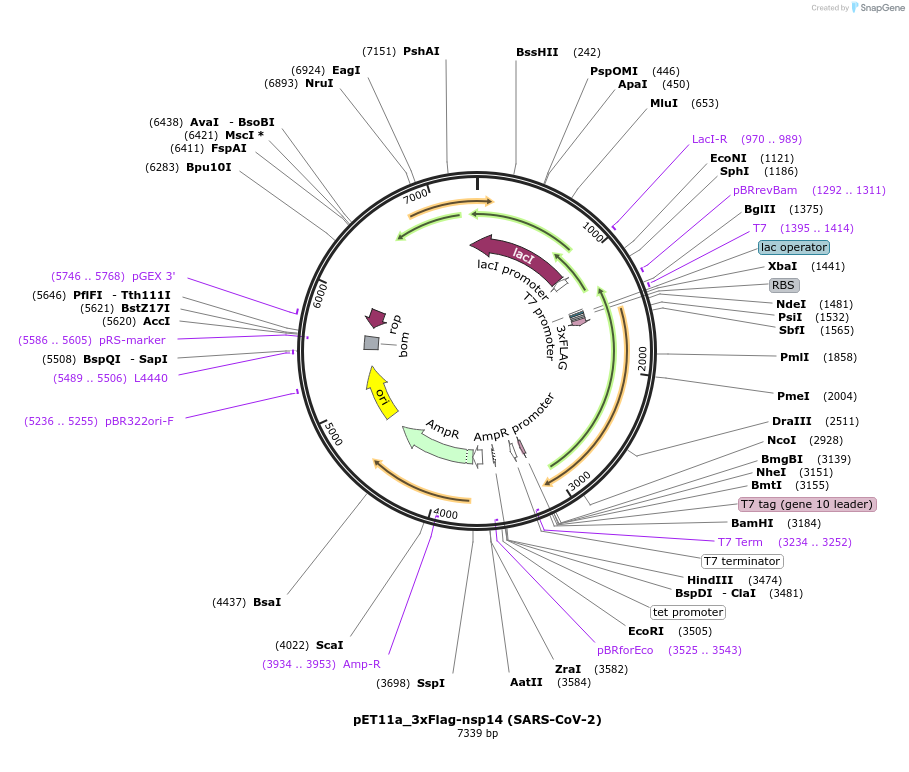
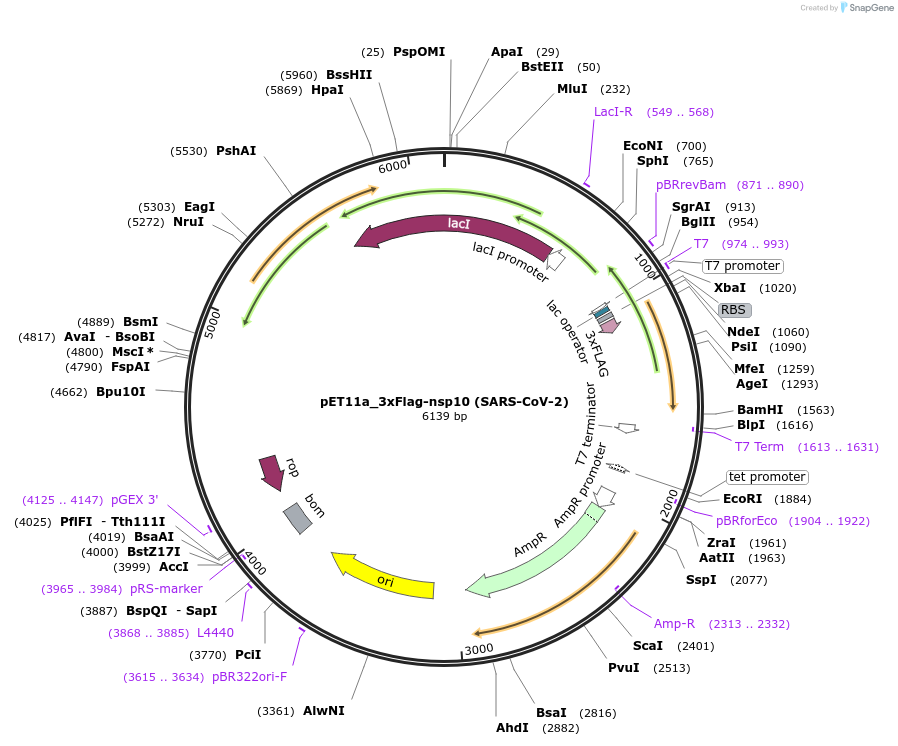
pET11a_3xFlag-nsp14 (SARS-CoV-2)
Plasmid#169159PurposeTo express SARS-CoV-2 nsp14 in E. coliDepositorInsert3xFlag-nsp5CS[VATLQ]-nsp14 (ORF1ab SARS-CoV-2)
Tags3xFlag and nsp5CS[VATLQ]: predicted nsp5 protease…ExpressionBacterialMutationCodon optimised for E. coliPromoterT7Available SinceJuly 6, 2021AvailabilityAcademic Institutions and Nonprofits only -
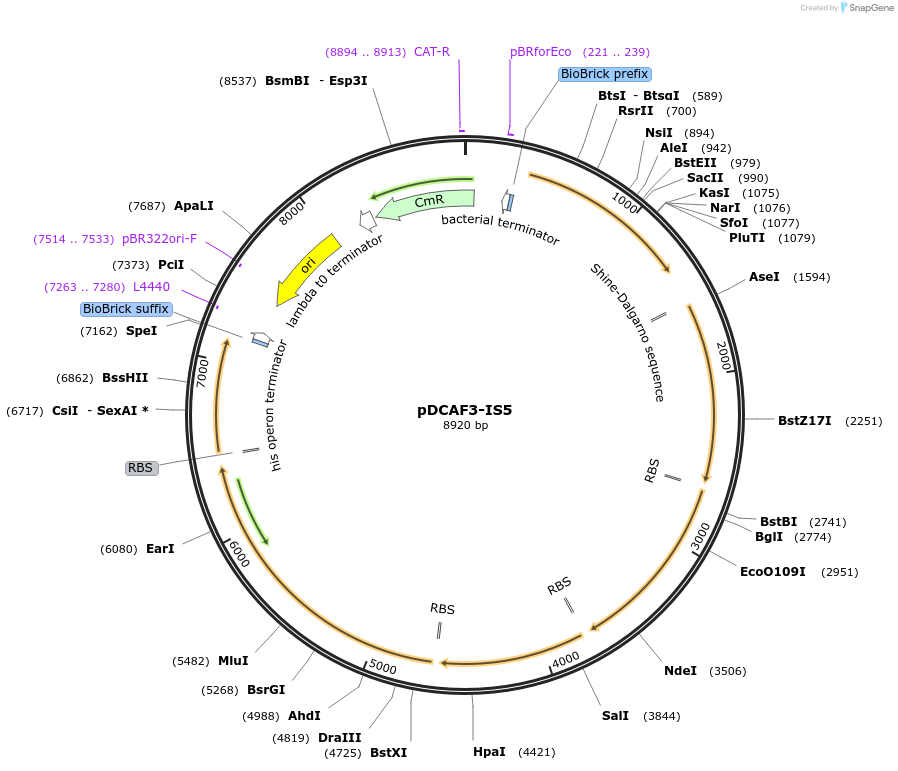
pDCAF3-IS5
Plasmid#65220PurposeExpresses N-demethylases for the degredation of caffeine and related methylxanthines. Can be used to measure caffeine content as described in the publication listed below.DepositorInsertsndmA
ndmB
ndmC
ndmD
gst9
chlR
UseSynthetic BiologyExpressionBacterialMutationChanged start codon from gtg to atg and Changed s…PromoterBBa_J23100Available SinceJan. 8, 2016AvailabilityIndustry, Academic Institutions, and Nonprofits -
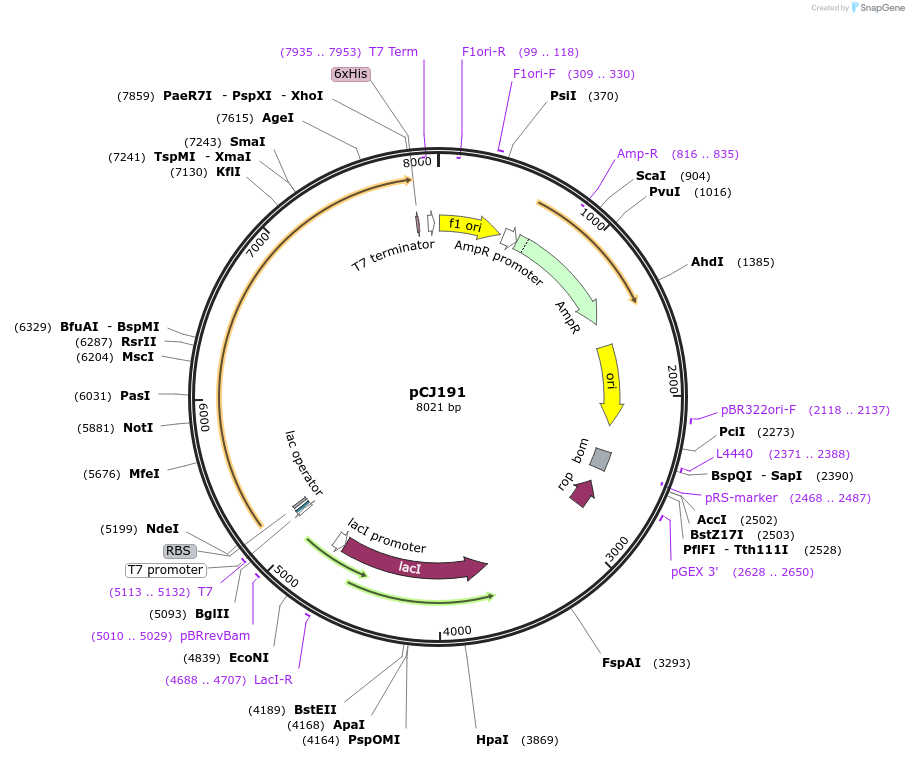
pCJ191
Plasmid#162668PurposepET-21b(+) based plasmid for expresion of MHETase (Genbank GAP38911.1) linked to PETase (Genbank GAP38373.1) from Ideonella sakaiensis by a 20 residue Gly/Ser linker; codon optimized for expression in E. coli K12, with C-terminal His tag.DepositorInsertMHETase (Genbank GAP38911.1) linked to PETase (Genbank GAP38373.1) from Ideonella sakaiensis by a 20 residue Gly/Ser linker
TagsHisExpressionBacterialMutationcodon optimized for expression in E. coli K12PromoterT7/lacAvailable SinceJan. 7, 2021AvailabilityAcademic Institutions and Nonprofits only -
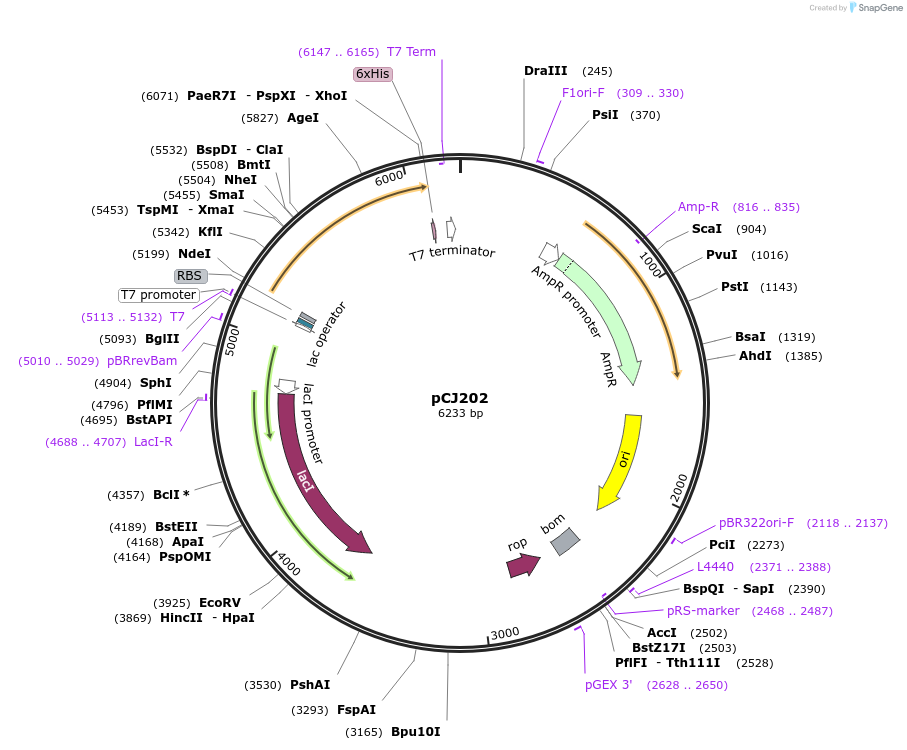
pCJ202
Plasmid#162675PurposepET-21b(+) based plasmid for expression of PETase from Ideonella sakaiensis 201-F6 (Genbank GAP38373.1), codon optimized for expression in E. coli K12, with C-terminal His tag, incorporating W159C and S238C mutations.DepositorInsertPETase from Ideonella sakaiensis 201-F6 (Genbank GAP38373.1)
TagsHisExpressionBacterialMutationW159C, S238C; codon optimized for expression in E…PromoterT7/lacAvailable SinceFeb. 3, 2021AvailabilityAcademic Institutions and Nonprofits only -
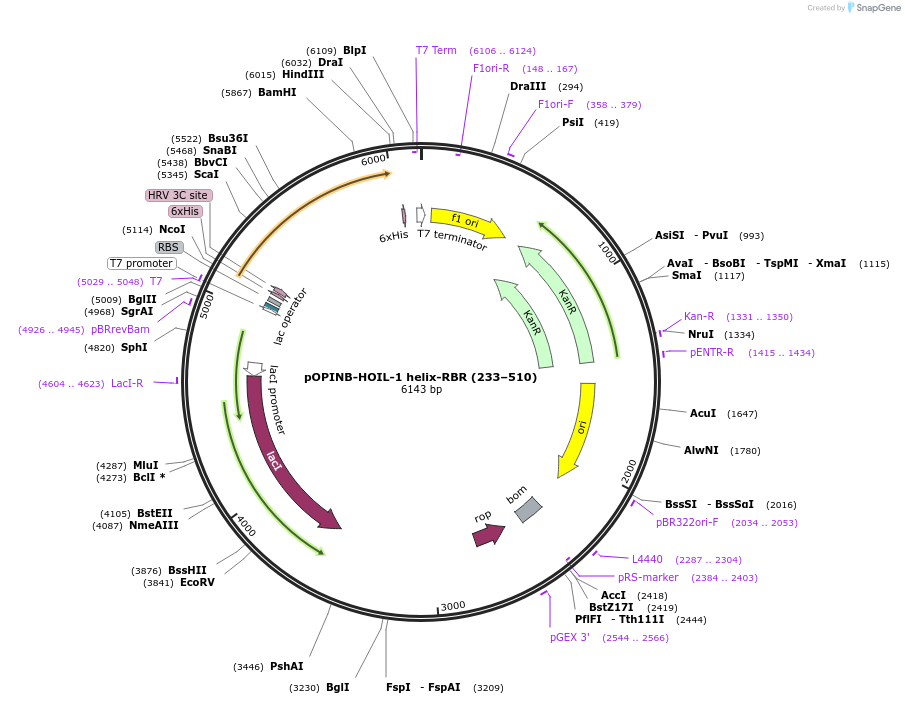
pOPINB-HOIL-1 helix-RBR (233–510)
Plasmid#193859PurposeExpression of human His-tagged HOIL-1 (RBCK1) helix-RBR (aa 233–510) codon optimized for E. coliDepositorInsertHOIL-1 (RBCK1 Human)
Tags3C protease cleavage site and 6x-His tagExpressionBacterialMutationResidues 233–510, codon optimised for expression …PromoterT7Available SinceJan. 12, 2023AvailabilityAcademic Institutions and Nonprofits only -
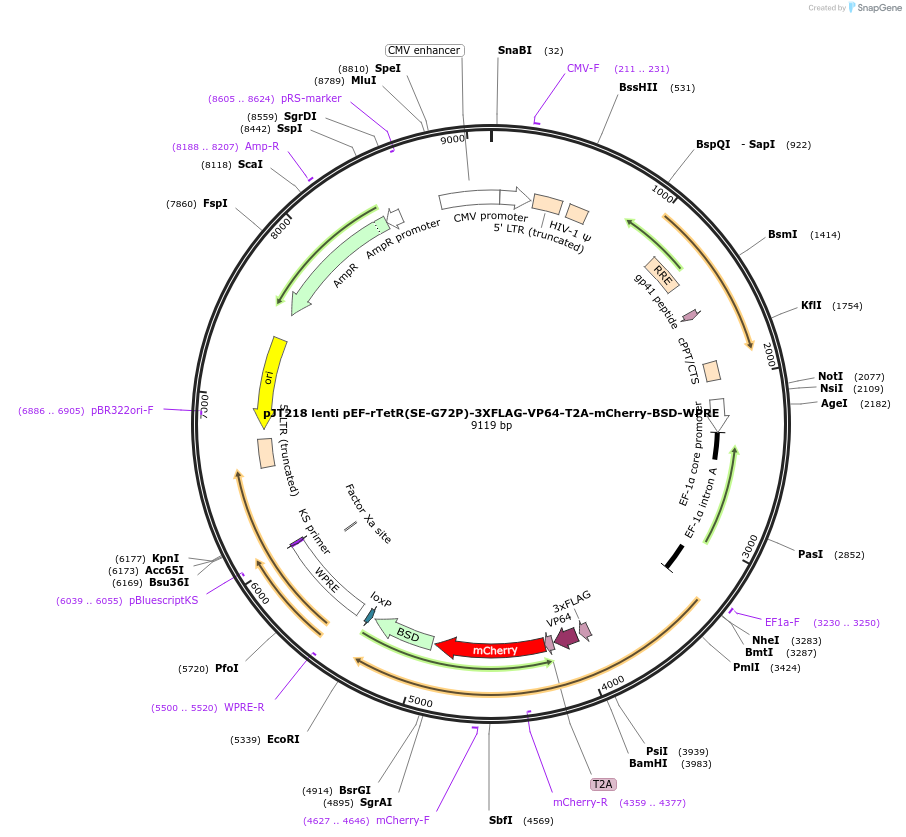
pJT218 lenti pEF-rTetR(SE-G72P)-3XFLAG-VP64-T2A-mCherry-BSD-WPRE
Plasmid#188758PurposeLentiviral backbone containing VP64 as a fusion with rTetR(SE-G72P)-3XFLAGDepositorInsertVP64
UseLentiviralTagsrTetR(SE-G72P)-3xFLAGExpressionMammalianPromoterpEF1aAvailable SinceMay 7, 2024AvailabilityAcademic Institutions and Nonprofits only -
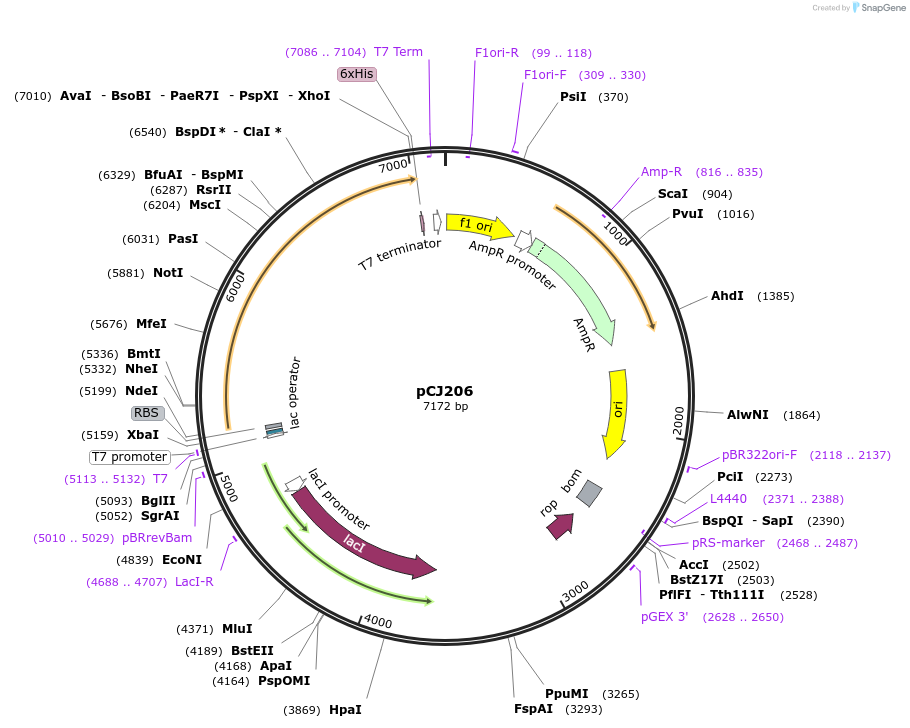
pCJ206
Plasmid#162679PurposepET-21b(+) based plasmid for expression of MHETase from Ideonella sakaiensis 201-F6 (Genbank GAP38911.1), codon optimized for expression in E. coli K12, with C-terminal His tag, incorporating the E226T mutation to the putative lipase box.DepositorInsertMHETase from Ideonella sakaiensis 201-F6 (Genbank GAP38911.1), incorporating the E226T mutation to the putative lipase box
TagsHisExpressionBacterialMutationE226T; codon optimized for expression in E. coli …PromoterT7/lacAvailable SinceJan. 7, 2021AvailabilityAcademic Institutions and Nonprofits only -
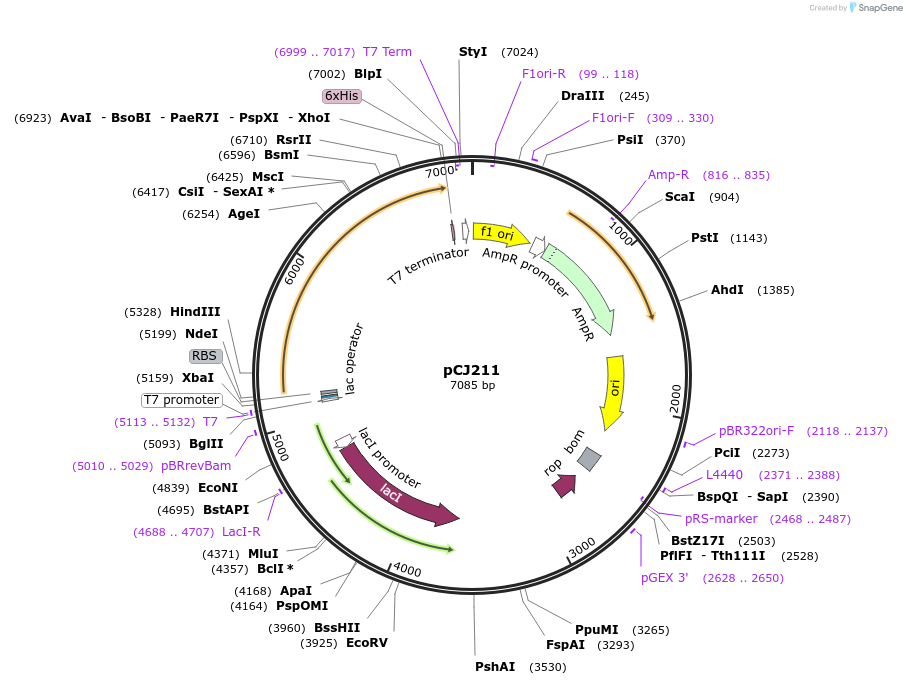
pCJ211
Plasmid#162684PurposepET-21b(+) based plasmid for expression of the putative MHETase from Hydrogenophaga sp. PML113 (Genbank WP_083293388.1) with deletion of the predicted signal peptide (Lys2:Gly19), with C-terminal His tag, codon optimized for expression in E. coli K12.DepositorInsertPutative MHETase from Hydrogenophaga sp. PML113 (Genbank WP_083293388.1) without 19 residue signal peptide
TagsHisExpressionBacterialMutationdeltaK2:G19; codon optimized for expression in E.…PromoterT7/lacAvailable SinceJan. 7, 2021AvailabilityAcademic Institutions and Nonprofits only -
pET11a_3xFlag-nsp10 (SARS-CoV-2)
Plasmid#169157PurposeTo express SARS-CoV-2 nsp10 in E. coliDepositorInsert3xFlag-nsp5CS[VRLQ]-nsp10 (ORF1ab SARS-CoV-2, Synthetic)
Tags3xFlag and nsp5CS[VRLQ]: predicted nsp5 protease …ExpressionBacterialMutationCodon optimised for E. coliPromoterT7Available SinceJuly 6, 2021AvailabilityAcademic Institutions and Nonprofits only -
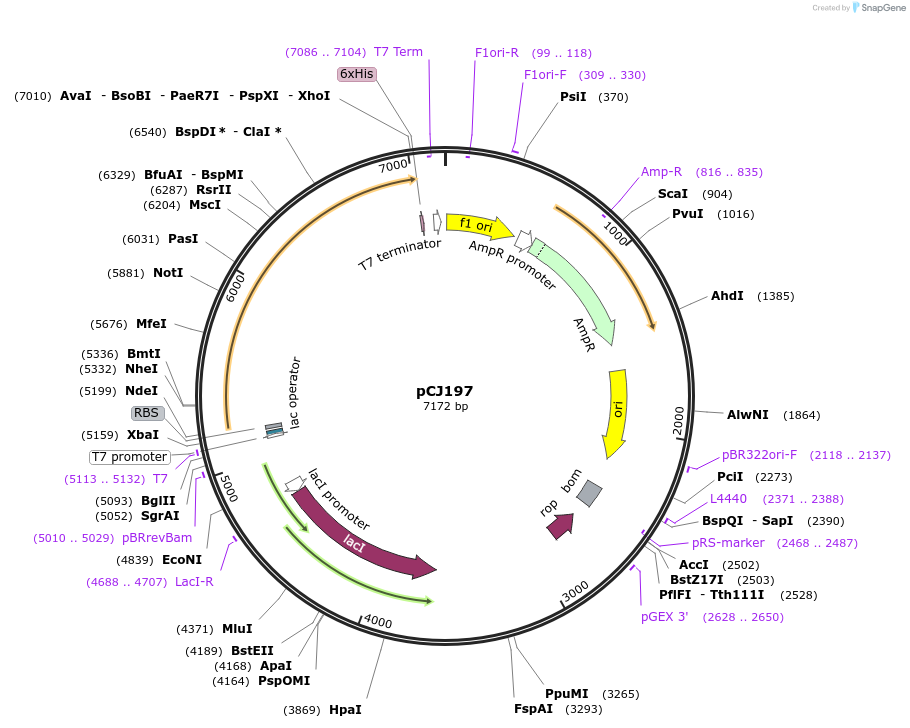
pCJ197
Plasmid#162670PurposepET-21b(+) based plasmid for expression of MHETase from Ideonella sakaiensis 201-F6 (Genbank GAP38911.1), codon optimized for expression in E. coli K12, with C-terminal His tag, incorporating S131G mutation.DepositorInsertMHETase from Ideonella sakaiensis 201-F6 (Genbank GAP38911.1
TagsHisExpressionBacterialMutationS131G; codon optimized for expression in E. coli …PromoterT7/lacAvailable SinceJan. 7, 2021AvailabilityAcademic Institutions and Nonprofits only -
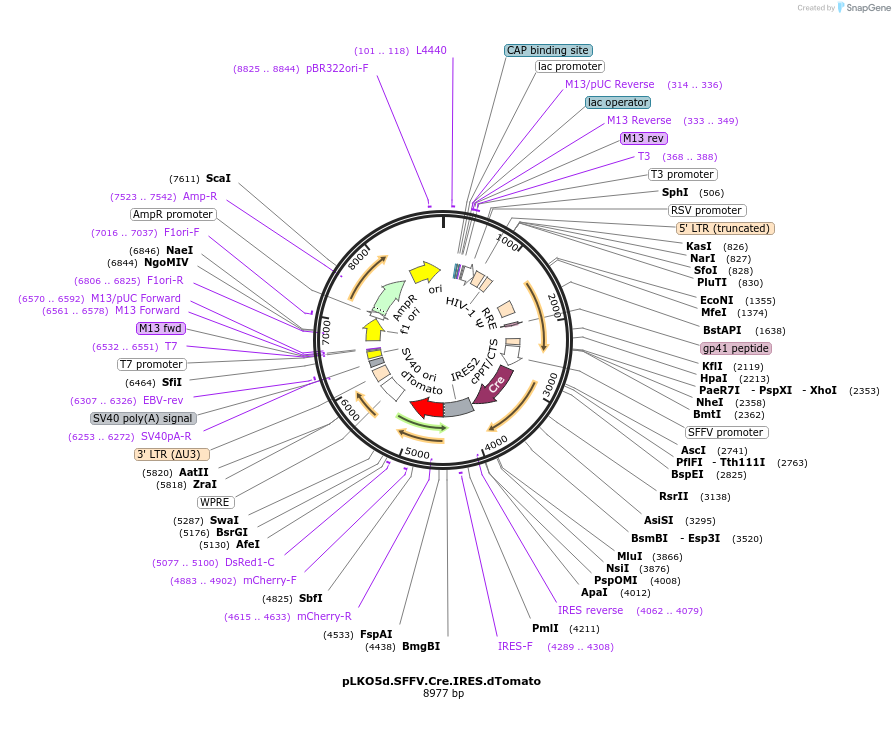
pLKO5d.SFFV.Cre.IRES.dTomato
Plasmid#187192PurposeLentiviral overexpression of Cre recombinaseDepositorInsertCre recombinase (cre Escherichia phage P1 (isolate: mod749::IS5 c1.100 mutant, nat-host: Escherichia coli))
UseLentiviralAvailable SinceFeb. 2, 2026AvailabilityAcademic Institutions and Nonprofits only -
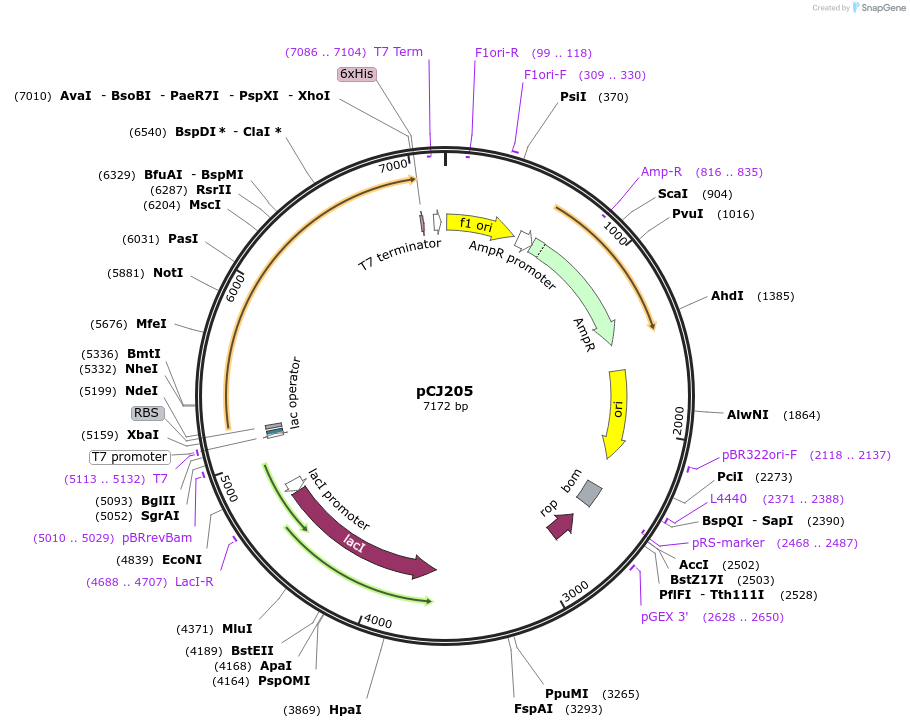
pCJ205
Plasmid#162676PurposepET-21b(+) based plasmid for expression of MHETase from Ideonella sakaiensis 201-F6 (Genbank GAP38911.1), codon optimized for expression in E. coli K12, with C-terminal His tag, incorporating C224A anDepositorInsertMHETase from Ideonella sakaiensis 201-F6 (Genbank GAP38911.1)
TagsHisExpressionBacterialMutationC224A, C529A; codon optimized for expression in E…PromoterT7/lacAvailable SinceFeb. 3, 2021AvailabilityAcademic Institutions and Nonprofits only -
pCJ198
Plasmid#162671PurposepET-21b(+) based plasmid for expression of MHETase from Ideonella sakaiensis 201-F6 (Genbank GAP38911.1), codon optimized for expression in E. coli K12, with C-terminal His tag, incorporating F495I mutation.DepositorInsertMHETase from Ideonella sakaiensis 201-F6 (Genbank GAP38911.1)
TagsHisExpressionBacterialMutationF495I; codon optimized for expression in E. coli …PromoterT7/lacAvailable SinceJan. 7, 2021AvailabilityAcademic Institutions and Nonprofits only -
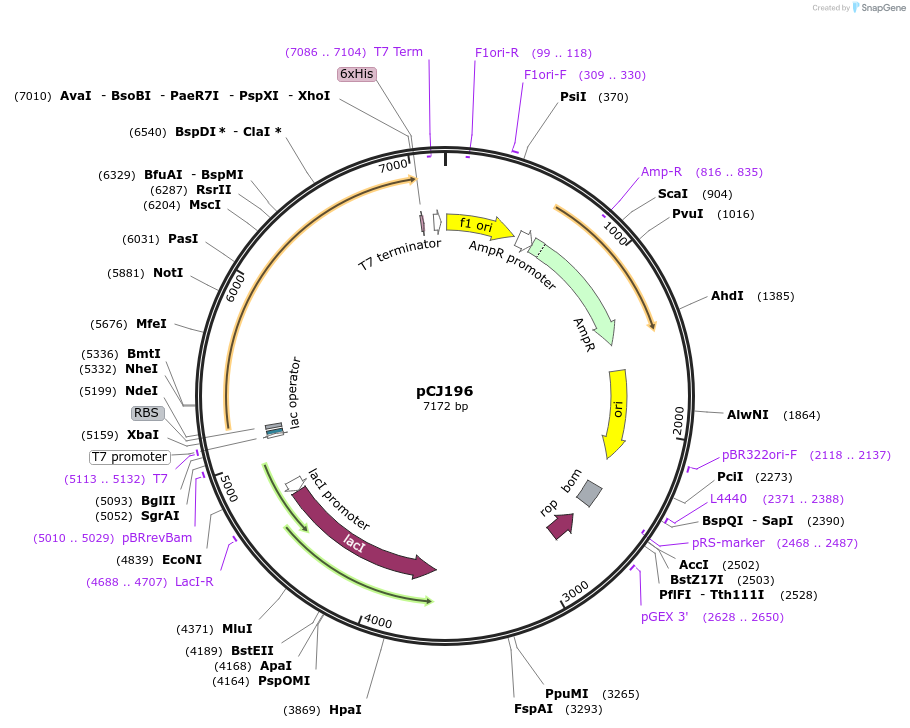
pCJ196
Plasmid#162669PurposepET-21b(+) based plasmid for expression of MHETase from Ideonella sakaiensis 201-F6 (Genbank GAP38911.1), codon optimized for expression in E. coli K12, with C-terminal His tag, incorporating catalytic mutation, S225ADepositorInsertMHETase from Ideonella sakaiensis 201-F6 (Genbank GAP38911.1)
TagsHisExpressionBacterialMutationS225A; codon optimized for expression in E. coli …PromoterT7/lacAvailable SinceJan. 7, 2021AvailabilityAcademic Institutions and Nonprofits only -
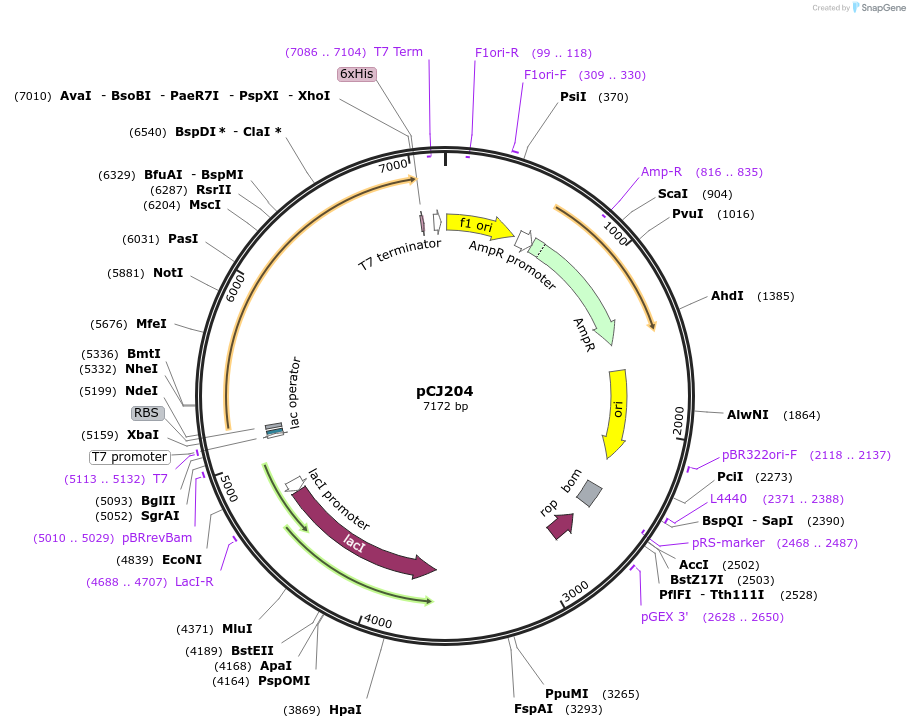
pCJ204
Plasmid#162677PurposepET-21b(+) based plasmid for expression of MHETase from Ideonella sakaiensis 201-F6 (Genbank GAP38911.1), codon optimized for expression in E. coli K12, with C-terminal His tag, incorporating C224H and C529F mutations.DepositorInsertMHETase from Ideonella sakaiensis 201-F6 (Genbank GAP38911.1)
TagsHisExpressionBacterialMutationC224H, C529F; codon optimized for expression in E…PromoterT7/lacAvailable SinceJan. 7, 2021AvailabilityAcademic Institutions and Nonprofits only -
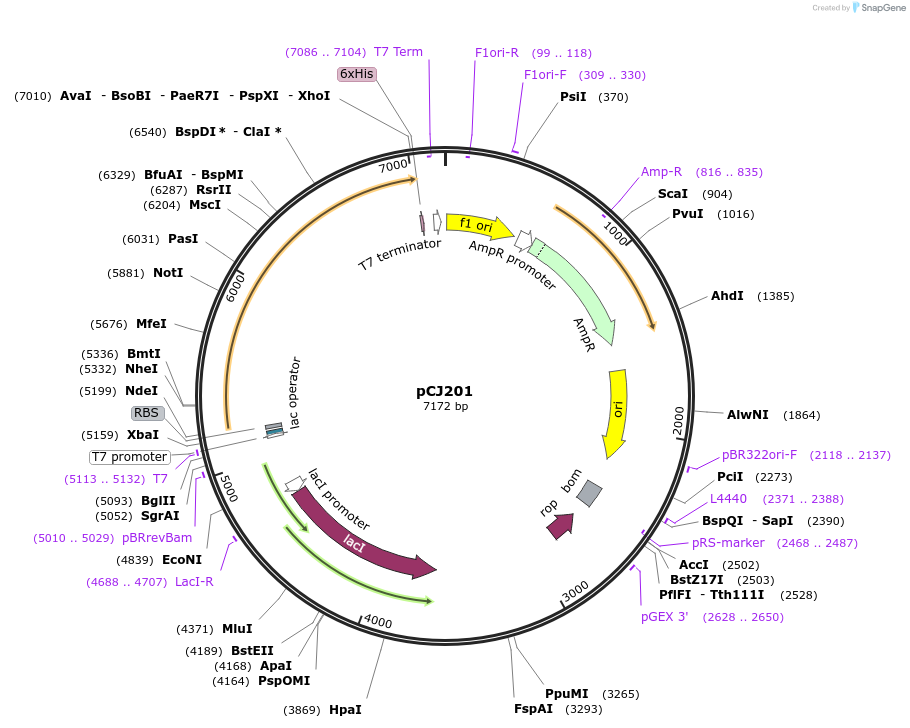
pCJ201
Plasmid#162674PurposepET-21b(+) based plasmid for expression of MHETase from Ideonella sakaiensis 201-F6 (Genbank GAP38911.1), codon optimized for expression in E. coli K12, with C-terminal His tag, incorporating C224W and C529SS mutations.DepositorInsertMHETase from Ideonella sakaiensis 201-F6 (Genbank GAP38911.1)
TagsHisExpressionBacterialMutationC224W, C529SS; codon optimized for expression in …PromoterT7/lacAvailable SinceJan. 6, 2021AvailabilityAcademic Institutions and Nonprofits only -
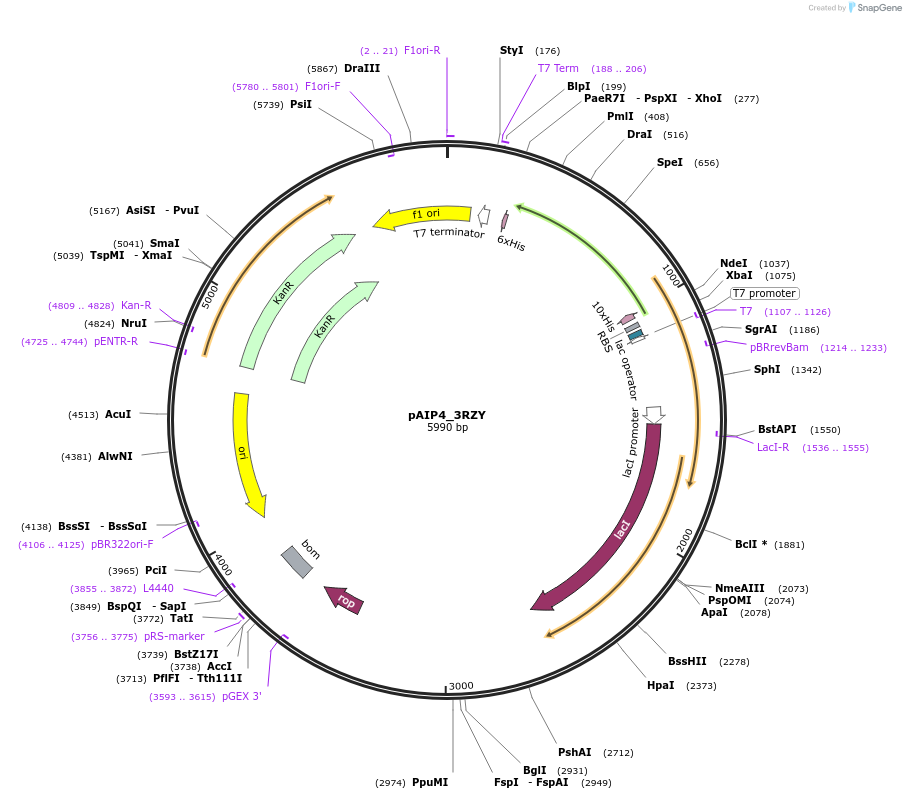
pAIP4_3RZY
Plasmid#234156PurposeHeterologous protein expression of fatty acid-binding protein, adipocyte (ALBP) in Escherichia coliDepositorAvailable SinceMarch 6, 2025AvailabilityIndustry, Academic Institutions, and Nonprofits -
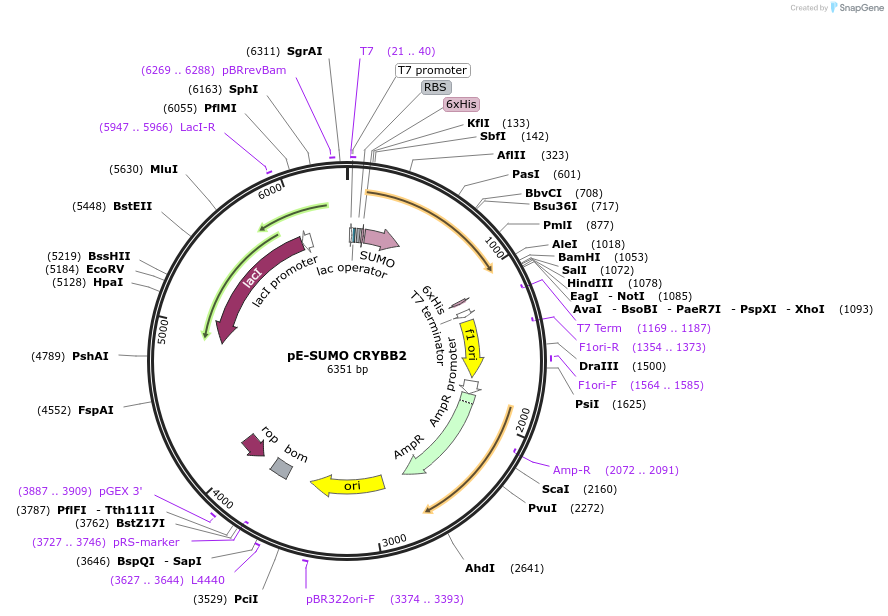
pE-SUMO CRYBB2
Plasmid#80749PurposeExpression of human beta B2-crystallin in E. coliDepositorAvailable SinceMarch 9, 2017AvailabilityAcademic Institutions and Nonprofits only -
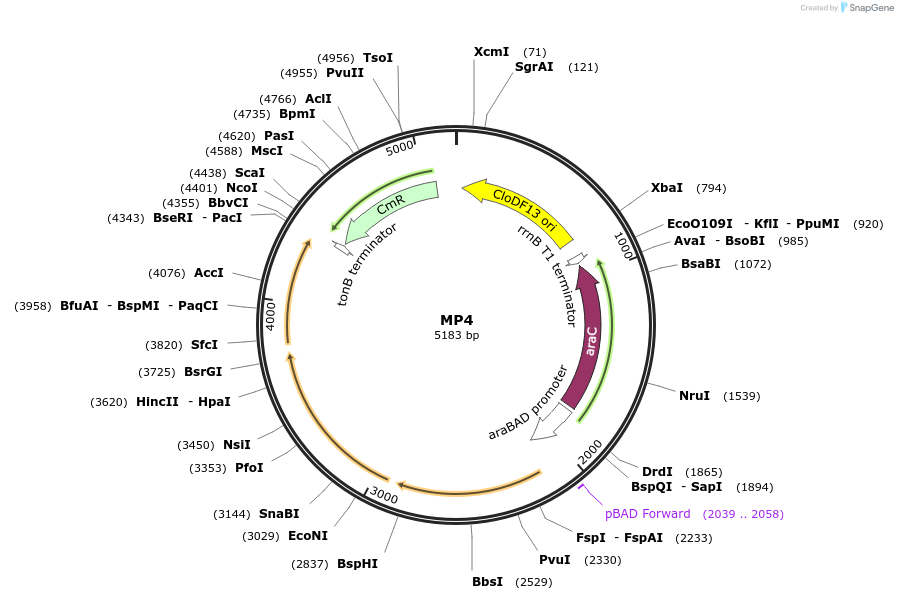
MP4
Plasmid#69652PurposeUntargetted Bacterial MutagenesisDepositorInsertdnaQ926 dam seqA
ExpressionBacterialPromoterE. coli PBADAvailable SinceOct. 8, 2015AvailabilityAcademic Institutions and Nonprofits only