We narrowed to 30,644 results for: PLE
-
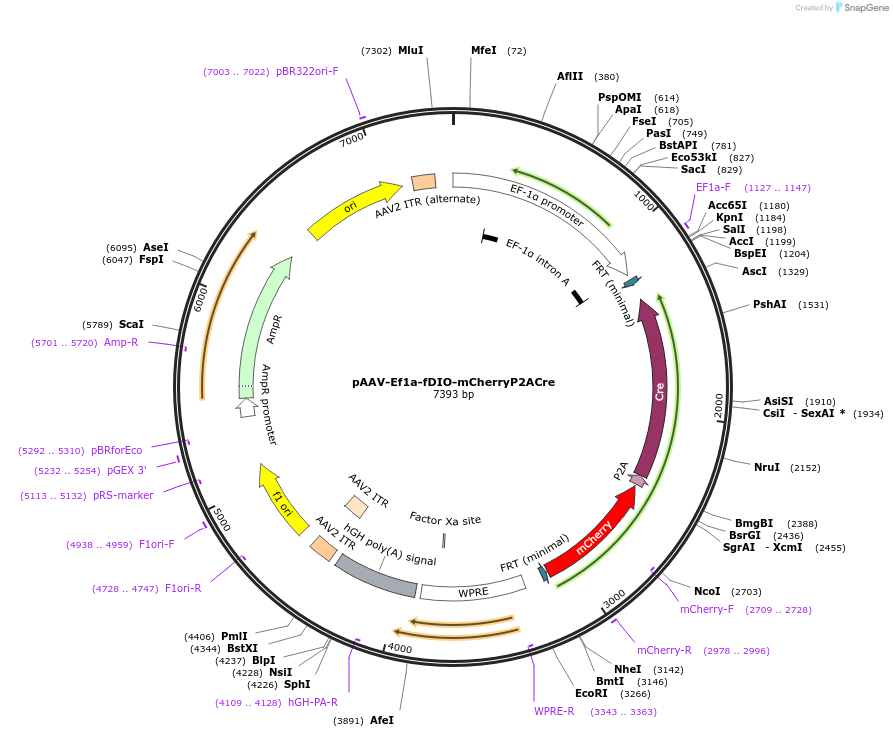
Plasmid#161773PurposeFlp-Dependent mCherry and Cre expressionDepositorInsertmCherry-P2A-Cre
UseAAVExpressionMammalianPromoterEf1aAvailable SinceNov. 20, 2020AvailabilityAcademic Institutions and Nonprofits only -
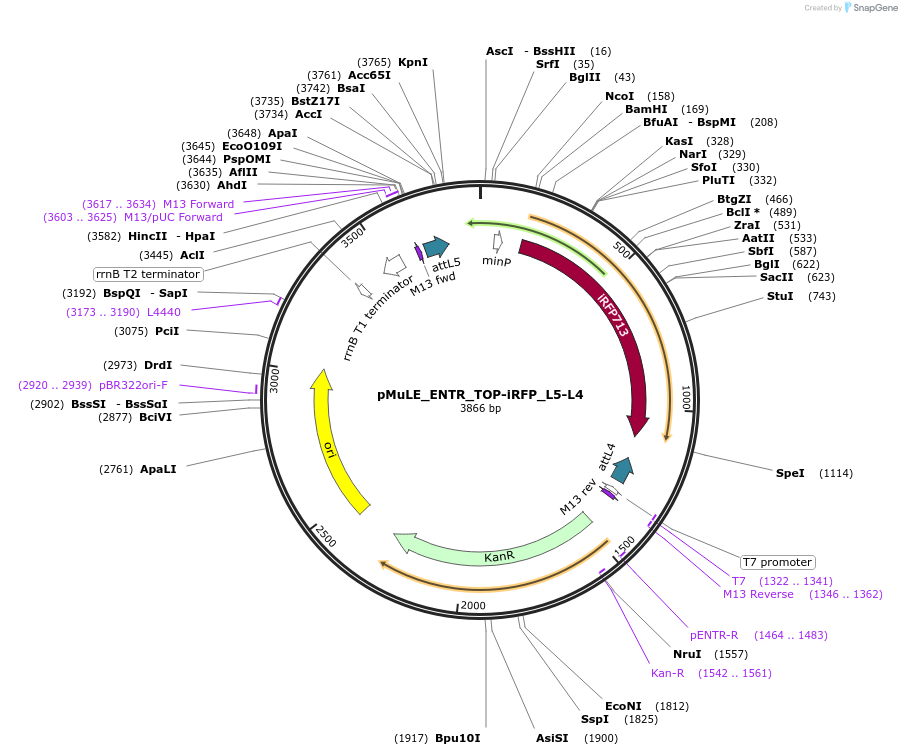
pMuLE_ENTR_TOP-iRFP_L5-L4
Plasmid#113866PurposeSingle pathway reporter; wnt promoter-driven expression of iRFP; MultiSite Gateway entry vector.DepositorInsertsTOP (wnt dependent promoter)
iRFP713
UseMule gateway entry vectorExpressionMammalianAvailable SinceJan. 22, 2020AvailabilityAcademic Institutions and Nonprofits only -
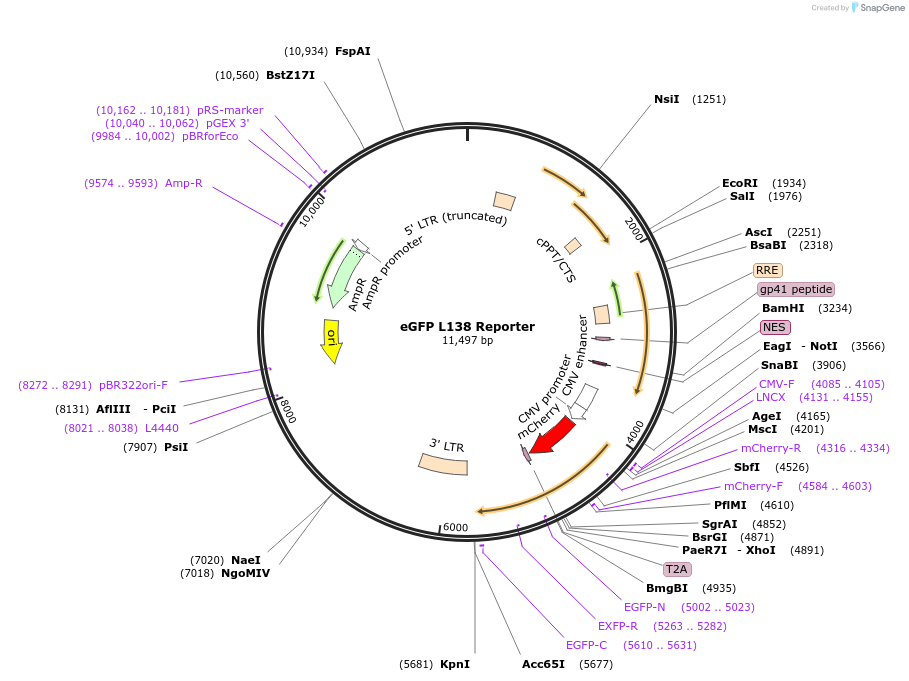
eGFP L138 Reporter
Plasmid#119130PurposeActs as a reporter for APOBEC-medated single-base editing. mCherry acts as a transfection control and eGFP as a reporter.DepositorInsertmCherry and eGFP
UseLentiviralMutationeGFP L138SPromoterCMVAvailable SinceJune 10, 2019AvailabilityAcademic Institutions and Nonprofits only -
pCRISPomyces-1
Plasmid#61736PurposeStreptomyces expression of codon-optimized Cas9, tracrRNA, and custom crRNADepositorInsertssSpCas9
tracrRNA
crRNA cassette
UseCRISPRExpressionBacterialMutationBbsI-flanked lacZ cassette inserted in place of s…Promotergapdhp(EL), rpsLp(CF), and rpsLp(XC)-BbsIAvailable SinceJan. 27, 2015AvailabilityAcademic Institutions and Nonprofits only -
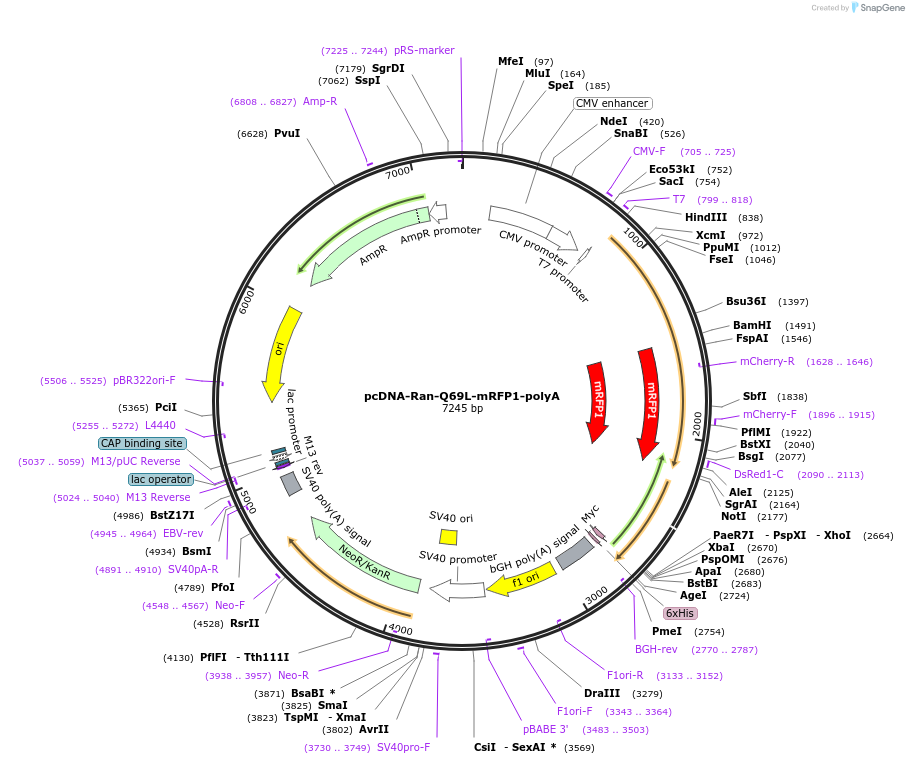
pcDNA-Ran-Q69L-mRFP1-polyA
Plasmid#104560PurposeExpress Ran Q69L mutant in mammalian cells (CMV promoter). Also suitable for in vitro transcription (T7 promoter) for injecting mRNA into oocytes.DepositorAvailable SinceDec. 21, 2017AvailabilityAcademic Institutions and Nonprofits only -
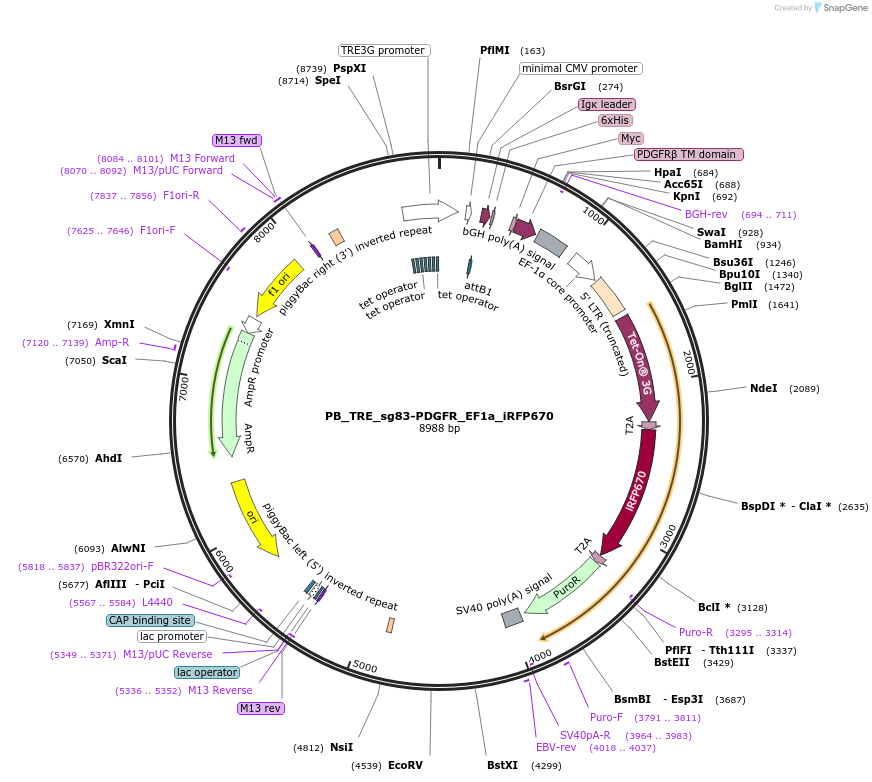
PB_TRE_sg83-PDGFR_EF1a_iRFP670
Plasmid#186311PurposePlasmid for inducible mammalian cell expression of the sg83 helixCAM and constitutive expression of an identifying fluorescent protein (iRFP670). Flanked with PiggyBac sites for easy line constructionDepositorInsertssg83-PDGFR
iRFP670
TagsHis, mycExpressionMammalianAvailable SinceAug. 10, 2022AvailabilityAcademic Institutions and Nonprofits only -
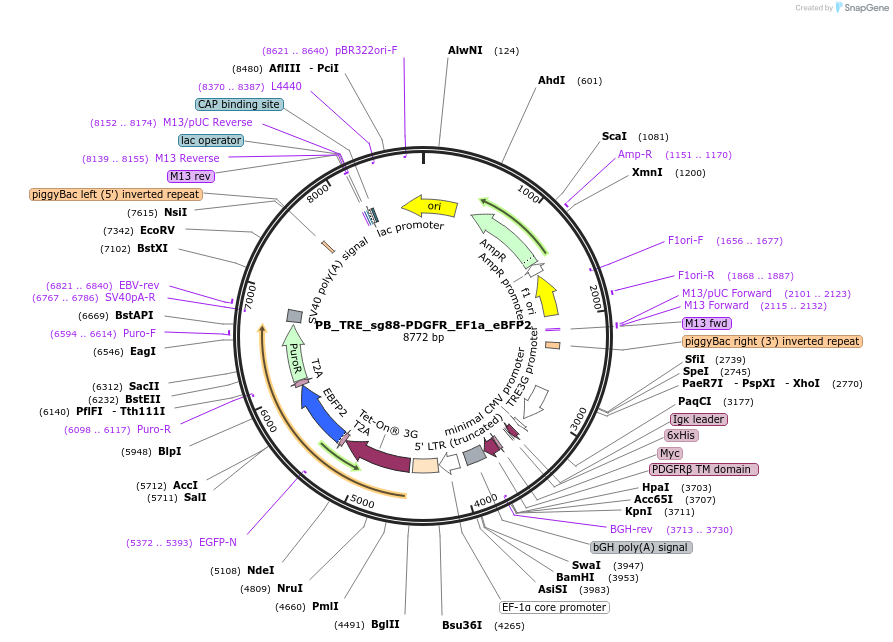
PB_TRE_sg88-PDGFR_EF1a_eBFP2
Plasmid#186312PurposePlasmid for inducible mammalian cell expression of the sg88 helixCAM and constitutive expression of an identifying fluorescent protein (eBFP2). Flanked with PiggyBac sites for easy line construction.DepositorInsertssg88-PDGFR
eBFP2
TagsHis, mycExpressionMammalianAvailable SinceAug. 10, 2022AvailabilityAcademic Institutions and Nonprofits only -
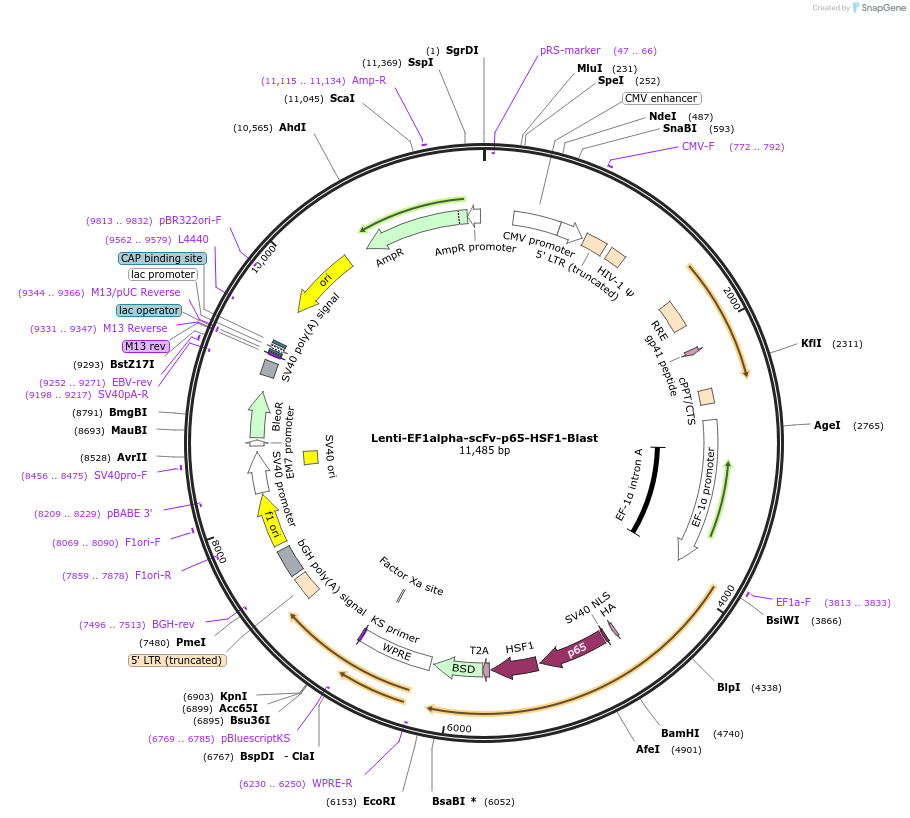
Lenti-EF1alpha-scFv-p65-HSF1-Blast
Plasmid#192652Purpose3rd generation lenti vector encoding scFV-p65-HSF1 with 2A Blast resistance markerDepositorInsertscFv-p65-HSF1
UseCRISPR and LentiviralExpressionMammalianPromoterEF1aAvailable SinceDec. 14, 2022AvailabilityAcademic Institutions and Nonprofits only -
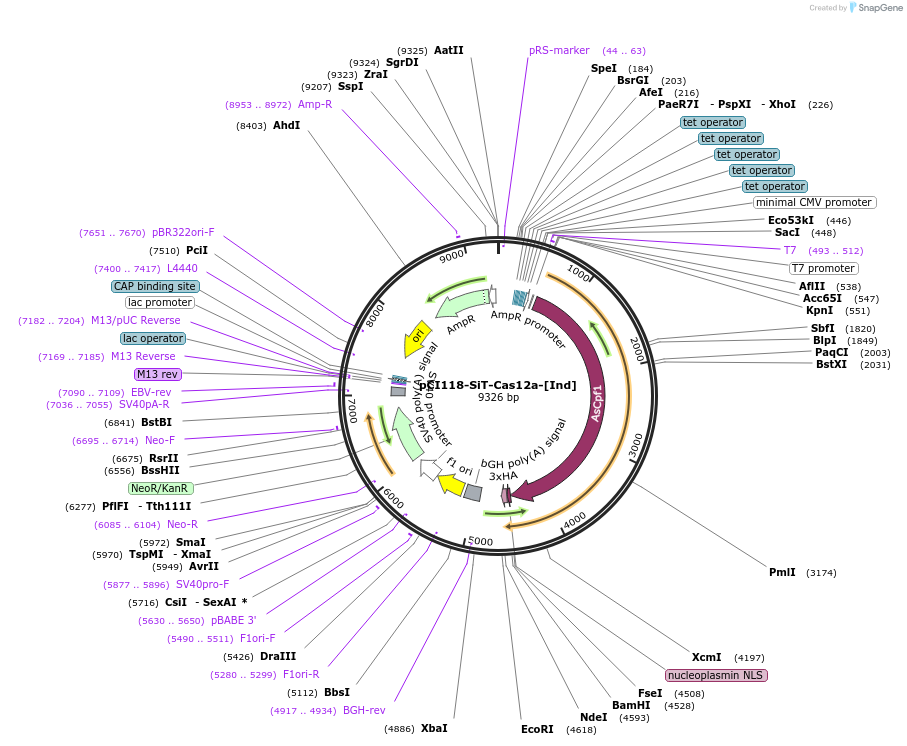
pCI118-SiT-Cas12a-[Ind]
Plasmid#128406PurposeInducible expression of both AsCas12a and CRISPR arrays on single transcriptsDepositorInsertAsCas12a-Triplex
UseAAVExpressionMammalianAvailable SinceOct. 1, 2019AvailabilityAcademic Institutions and Nonprofits only -
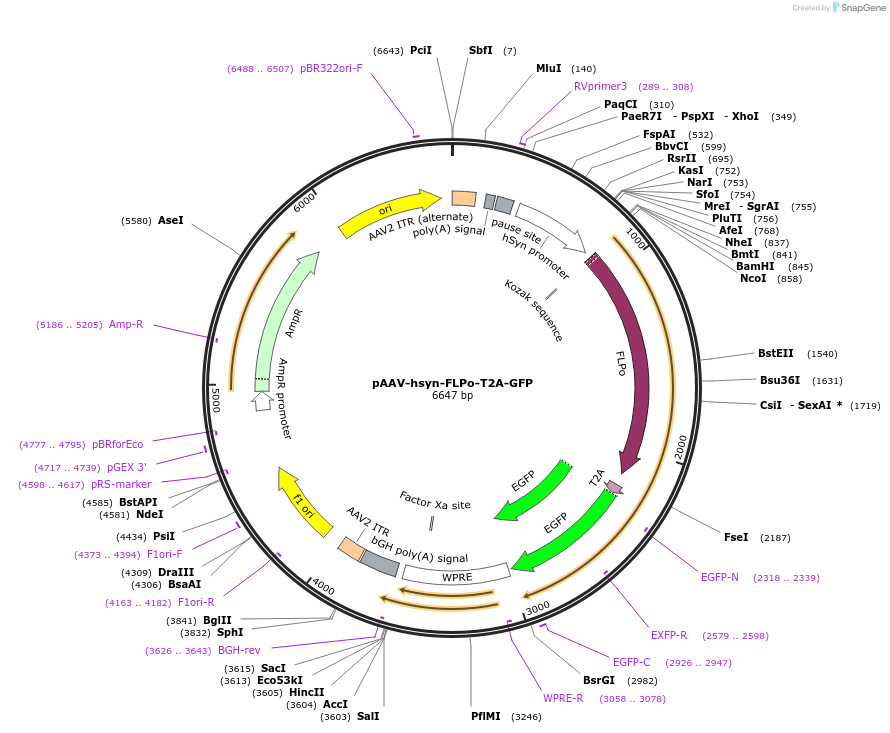
pAAV-hsyn-FLPo-T2A-GFP
Plasmid#161766Purposeexpress FLPo and GFP under the control of human synapsin promoterDepositorInsertsFLPo
T2AEGFP
UseAAVExpressionMammalianPromoterhSynapsin1Available SinceNov. 20, 2020AvailabilityAcademic Institutions and Nonprofits only -
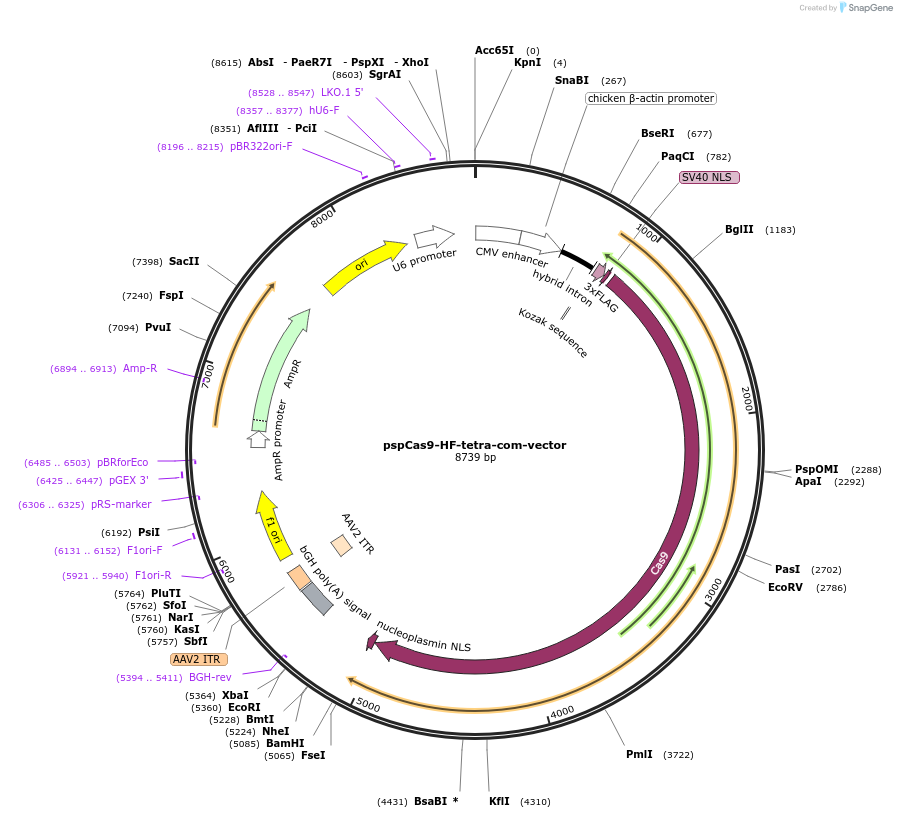
pspCas9-HF-tetra-com-vector
Plasmid#132555PurposeVector plasmid expressing high fidelity spCas9 (R691A) and sgRNA scaffold with com replacing the Tetraloop.DepositorInsertsHigh fidelity Cas9
sgRNA scaffold with com replacing the Tetraloop
ExpressionMammalianAvailable SinceMarch 22, 2021AvailabilityAcademic Institutions and Nonprofits only -
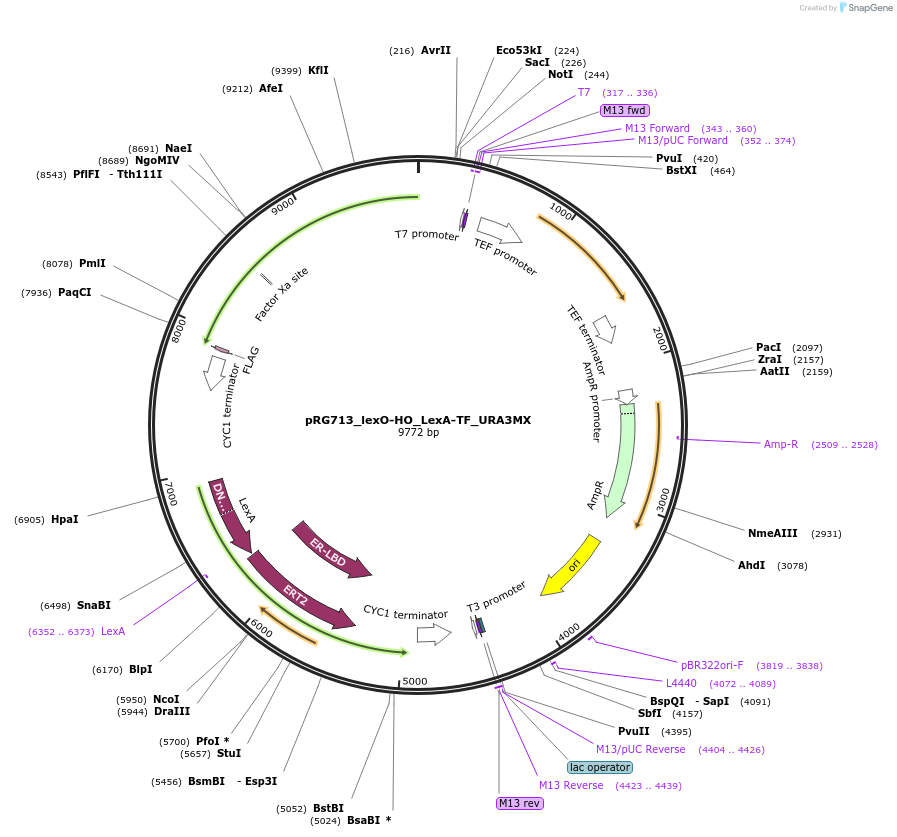
pRG713_lexO-HO_LexA-TF_URA3MX
Plasmid#154821PurposeInducible expression of HO endonuclease in S. cerevisiaeDepositorInsertsTagsFLAGExpressionYeastPromoterACT1 and lexO_4-CYC1_coreAvailable SinceOct. 26, 2020AvailabilityAcademic Institutions and Nonprofits only -
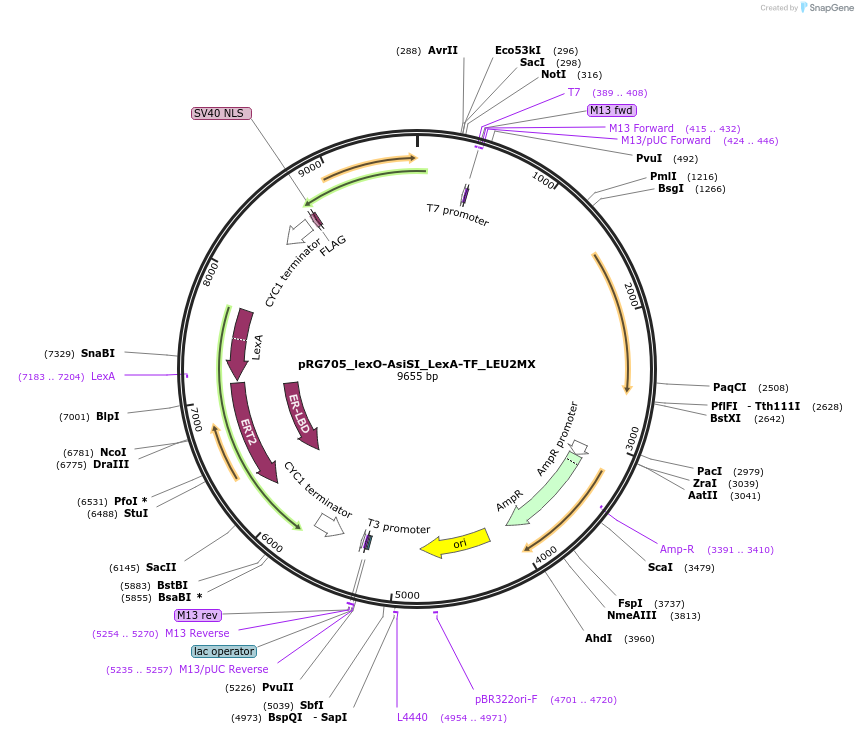
pRG705_lexO-AsiSI_LexA-TF_LEU2MX
Plasmid#154817PurposeInducible expression of AsiSI endonuclease in S. cerevisiaeDepositorInsertsAsiSI endonuclease
LexA-ER-B112
TagsFLAG and NLSExpressionYeastPromoterACT1 and lexO_4-CYC1_coreAvailable SinceOct. 26, 2020AvailabilityAcademic Institutions and Nonprofits only -
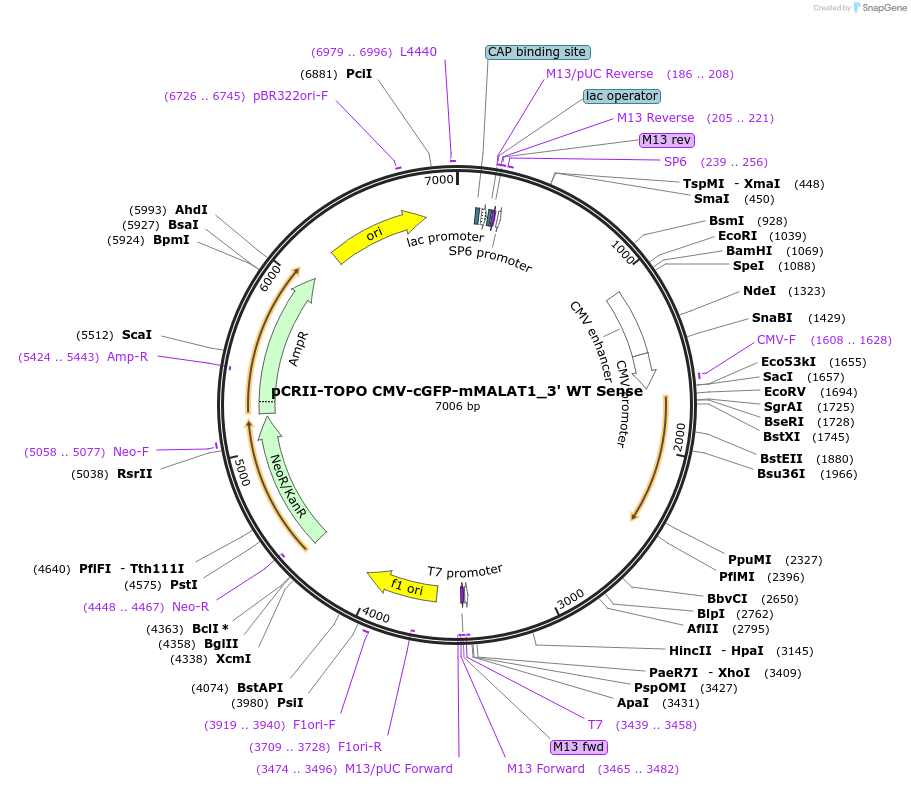
pCRII-TOPO CMV-cGFP-mMALAT1_3' WT Sense
Plasmid#46834PurposeExpress cGFP transcript ending in the wildtype MALAT1 triple helixDepositorInsertCoral Green Fluorescent Protein (cGFP)
ExpressionMammalianPromoterCMVAvailable SinceSept. 25, 2013AvailabilityAcademic Institutions and Nonprofits only -
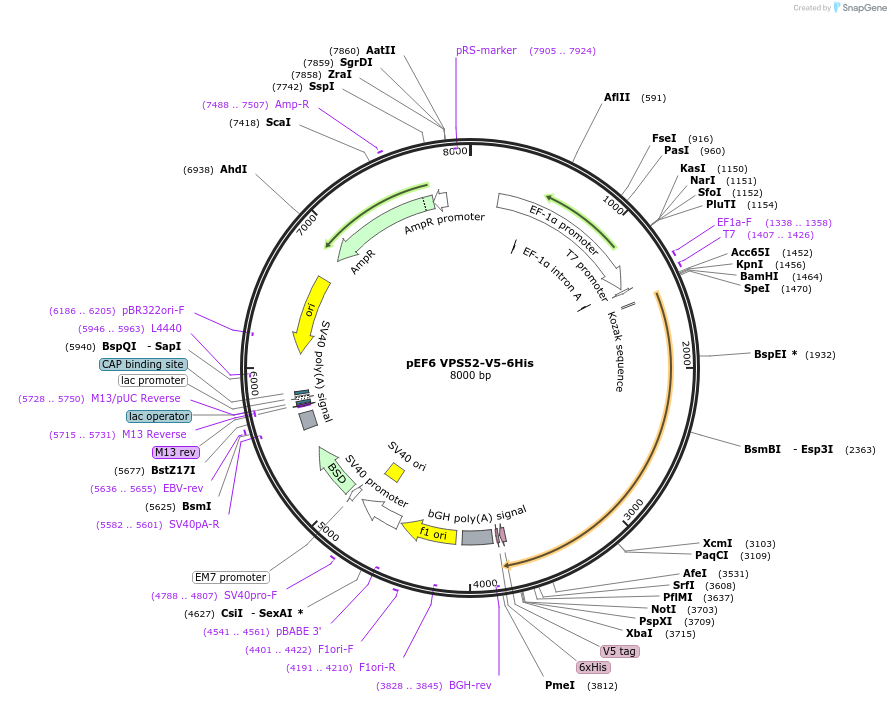
pEF6 VPS52-V5-6His
Plasmid#175096PurposeV5 tagged VPS52 is expressed in mammalian cellsDepositorInsertVPS52 (VPS52 Human)
TagsV5ExpressionMammalianMutationN666D- please see depositor commentAvailable SinceDec. 1, 2021AvailabilityAcademic Institutions and Nonprofits only -
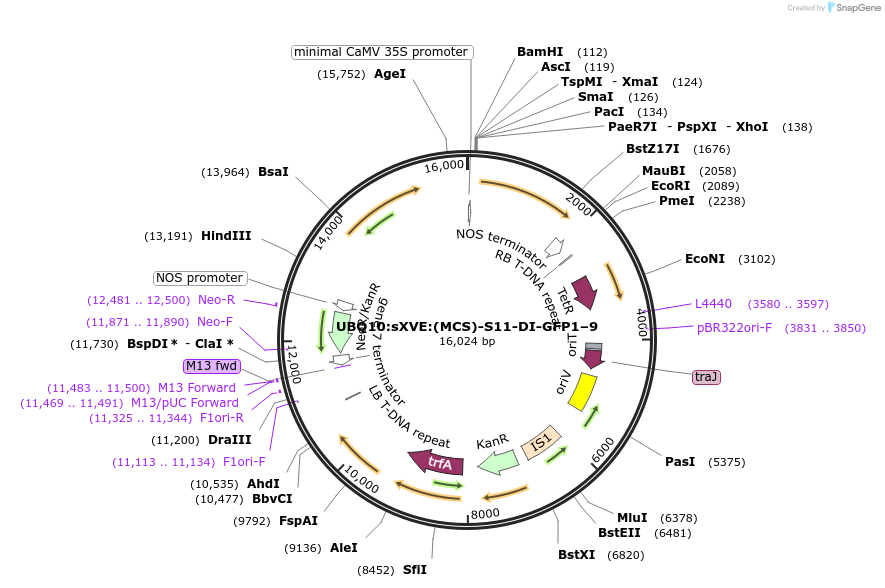
UBQ10:sXVE:(MCS)-S11-DI-GFP1–9
Plasmid#108260PurposeBinary vectors used for the inducible expression of dual intein-coupled tripartite split-sfGFP components in plantsDepositorTypeEmpty backboneUseSynthetic BiologyExpressionPlantPromoterUBQ10:sXVE:Available SinceApril 18, 2018AvailabilityAcademic Institutions and Nonprofits only -
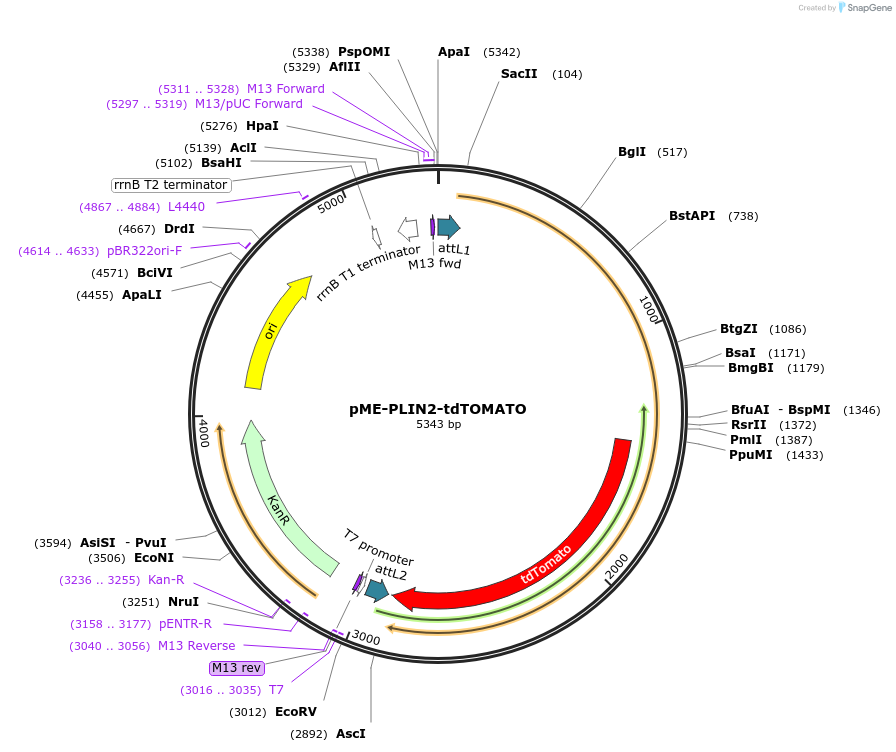
pME-PLIN2-tdTOMATO
Plasmid#172533PurposePLIN2-tdTOMATO fusion (zebrafish lipid droplet reporter) in middle entry vector for Gateway cloningDepositorInsertplin2-tdtomato
MutationS80R, A81S, E144D, I148F, and R415H mutations in …Available SinceSept. 20, 2021AvailabilityAcademic Institutions and Nonprofits only -
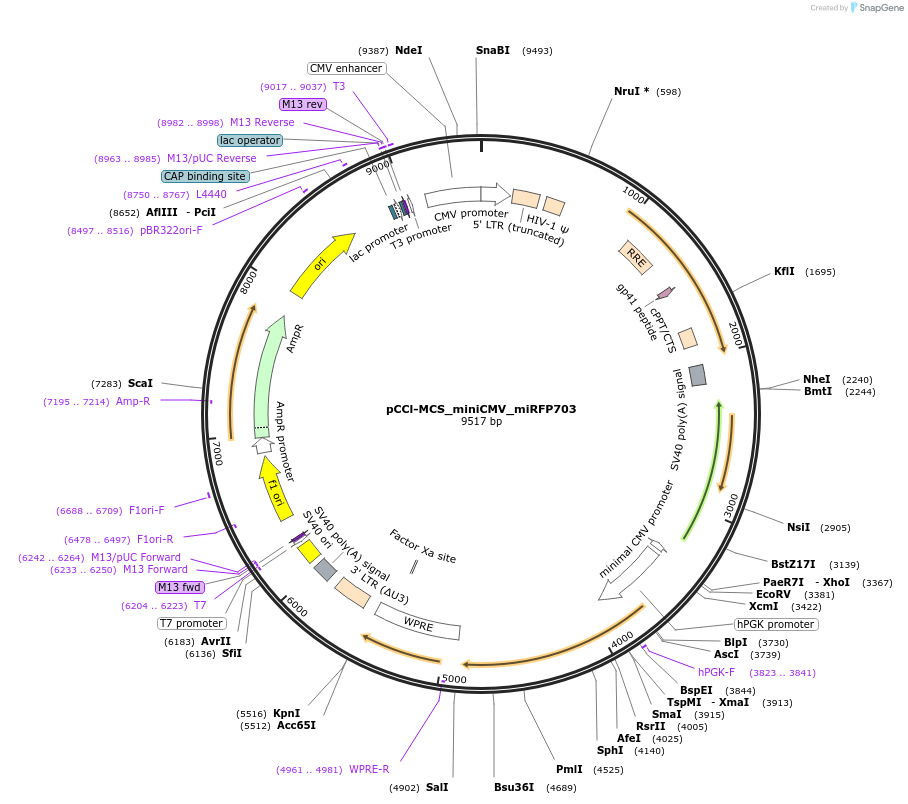
pCCl-MCS_miniCMV_miRFP703
Plasmid#134985PurposeThe plasmid includes a miRFP703 gene driven by a minimal CMV promoter. Cis-regulatory elements can be cloned into the single EcoRV site. This plasmid can be used for generating a lentiviral vectorDepositorInsertmiRFP703
UseLentiviralPromoterminCMVAvailable SinceJuly 15, 2020AvailabilityAcademic Institutions and Nonprofits only -
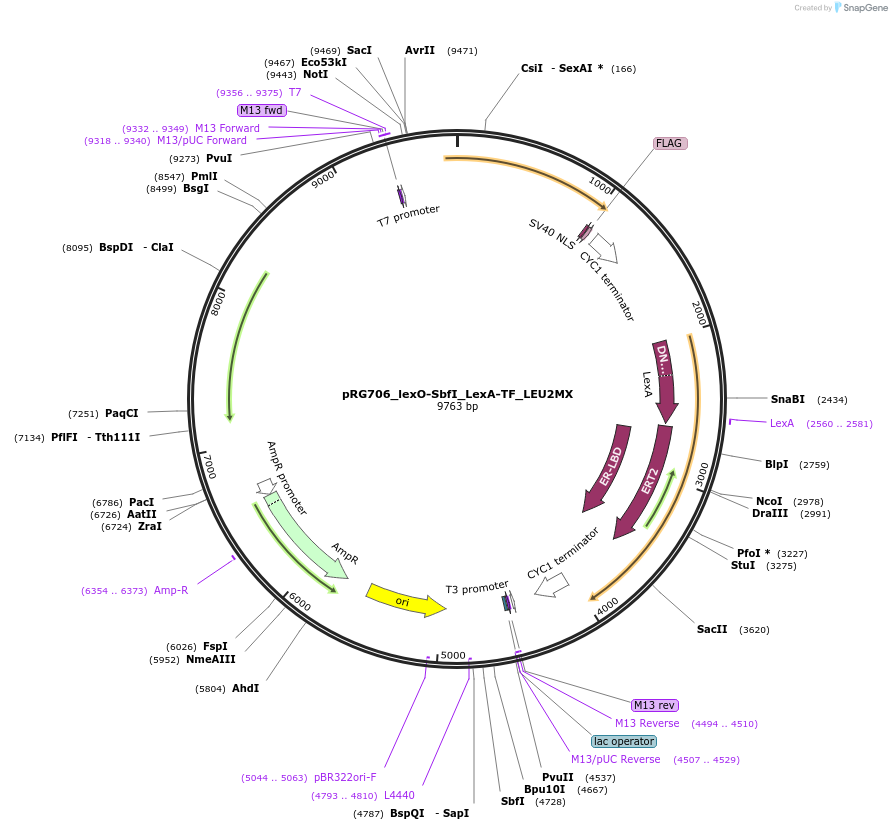
pRG706_lexO-SbfI_LexA-TF_LEU2MX
Plasmid#154818PurposeInducible expression of SbfI endonuclease in S. cerevisiaeDepositorInsertsSbfI endonuclease
LexA-ER-B112
TagsFLAG and NLSExpressionYeastPromoterACT1 and lexO_4-CYC1_coreAvailable SinceOct. 26, 2020AvailabilityAcademic Institutions and Nonprofits only -
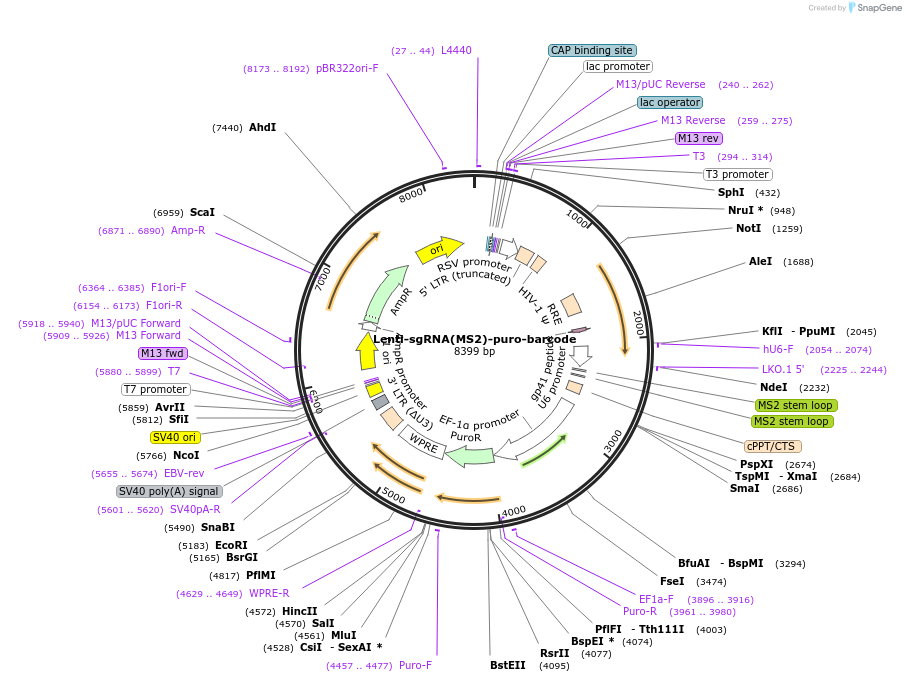
Lenti-sgRNA(MS2)-puro-barcode
Plasmid#89493PurposeCloning backbone for sgRNA, also has a barcode that allows detection from scRNA-seqDepositorInsertsgRNA (with MS2 loop)
UseLentiviralPromoterU6Available SinceJune 30, 2017AvailabilityAcademic Institutions and Nonprofits only -
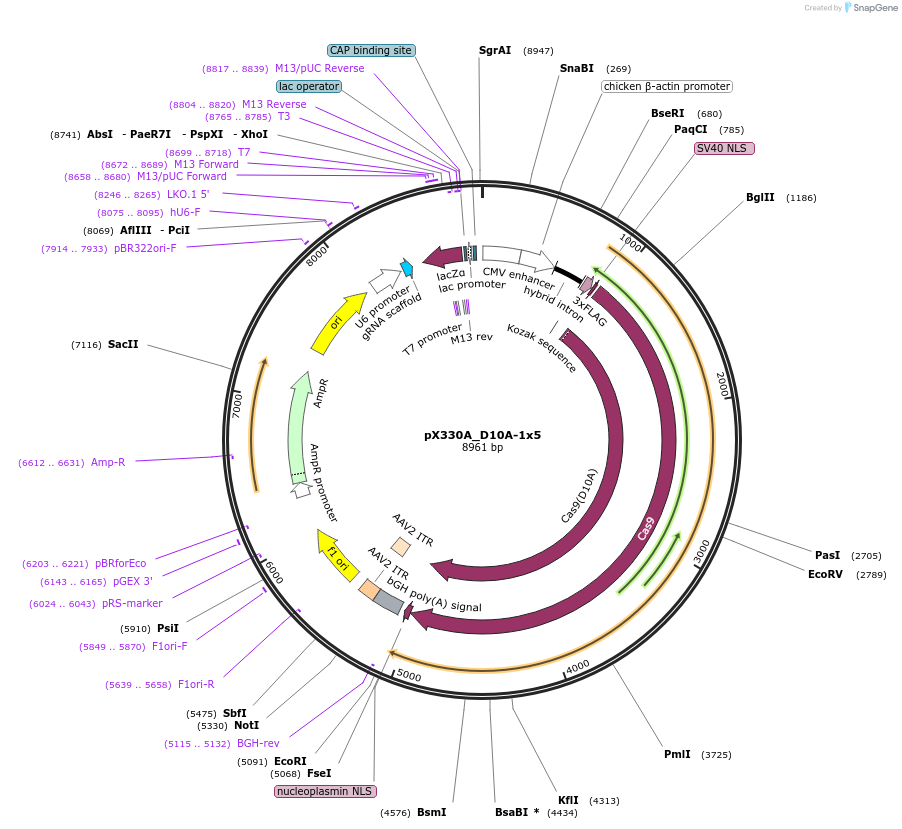
pX330A_D10A-1x5
Plasmid#58775PurposeExpresses Cas9 nickase and gRNADepositorInserthumanized S. pyogenes Cas9 (D10A) nickase
UseCRISPRTags3xFLAGExpressionMammalianMutationD10A nickase-converting mutationPromoterCBhAvailable SinceNov. 7, 2014AvailabilityAcademic Institutions and Nonprofits only -
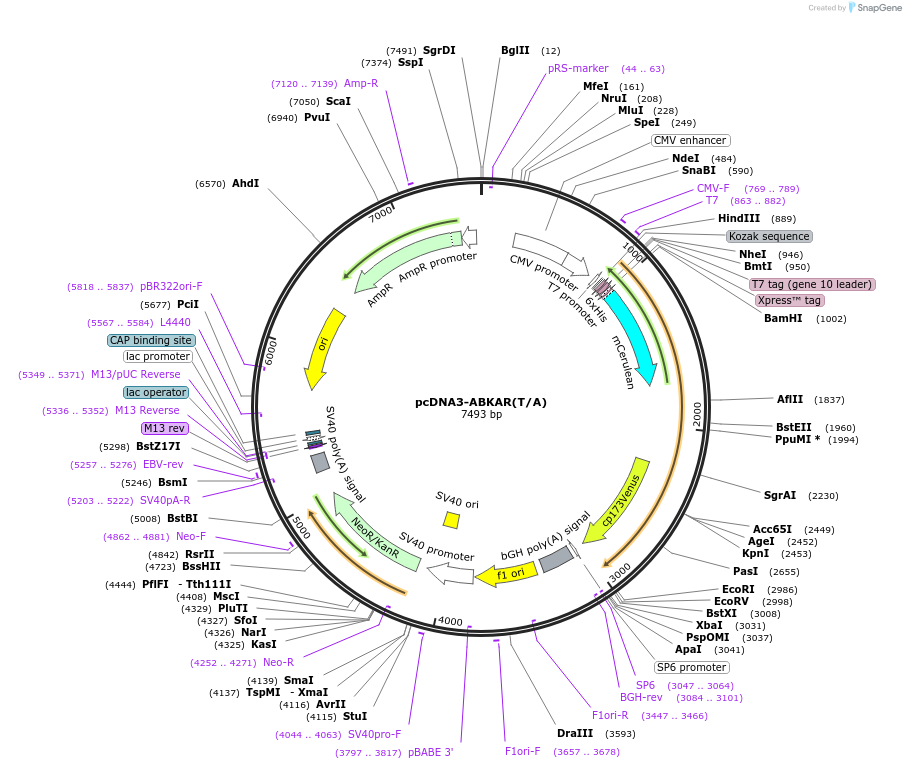
pcDNA3-ABKAR(T/A)
Plasmid#140052PurposeNegative control mutant for ABKAR biosensor.DepositorInsertABKAR(T/A)
Tags6xHIS - T7 tag (gene 10 leader) - Xpress (TM) tag…ExpressionMammalianMutationTarget Thr residue in substrate domain mutated to…PromoterCMVAvailable SinceNov. 21, 2020AvailabilityAcademic Institutions and Nonprofits only -
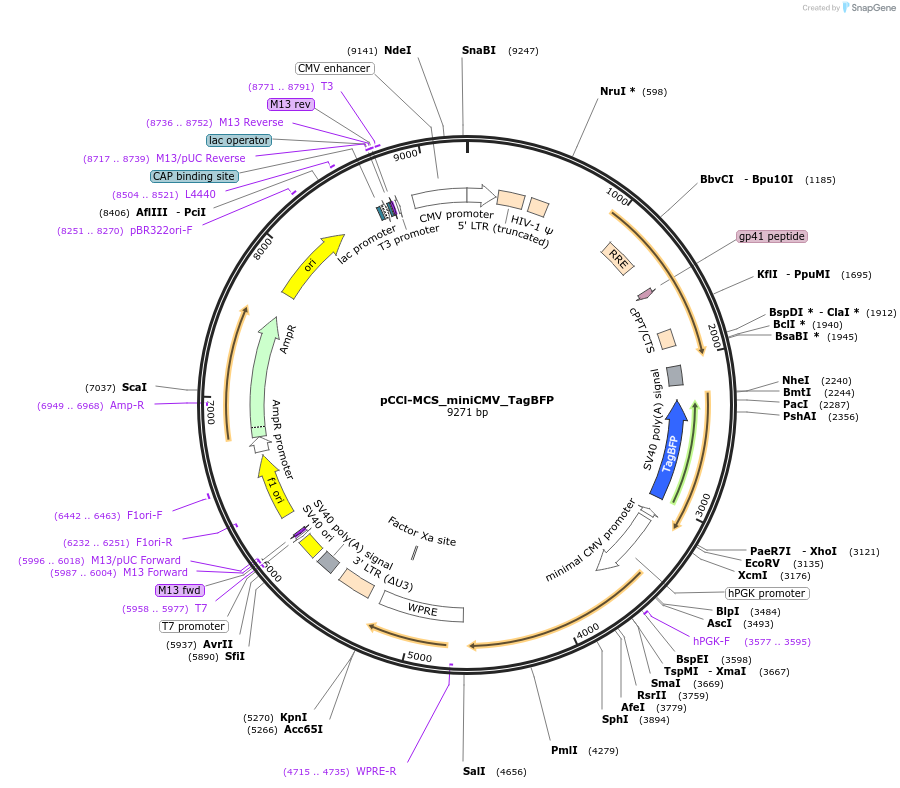
pCCl-MCS_miniCMV_TagBFP
Plasmid#134986PurposeThe plasmid includes a TagBFP gene driven by a minimal CMV promoter. Cis-regulatory elements can be cloned into the single EcoRV site. This plasmid can be used for generating a lentiviral vectorDepositorInsertTagBFP
UseLentiviralPromoterminCMVAvailable SinceJuly 15, 2020AvailabilityAcademic Institutions and Nonprofits only -
eGFP L138 gRNA
Plasmid#119133PurposegRNA that guides Cas9 and APOBEC complexes to L138 in eGFP.DepositorInserteGFP L138 gRNA
UseCRISPRMutationNonePromoterU6Available SinceJune 10, 2019AvailabilityAcademic Institutions and Nonprofits only -
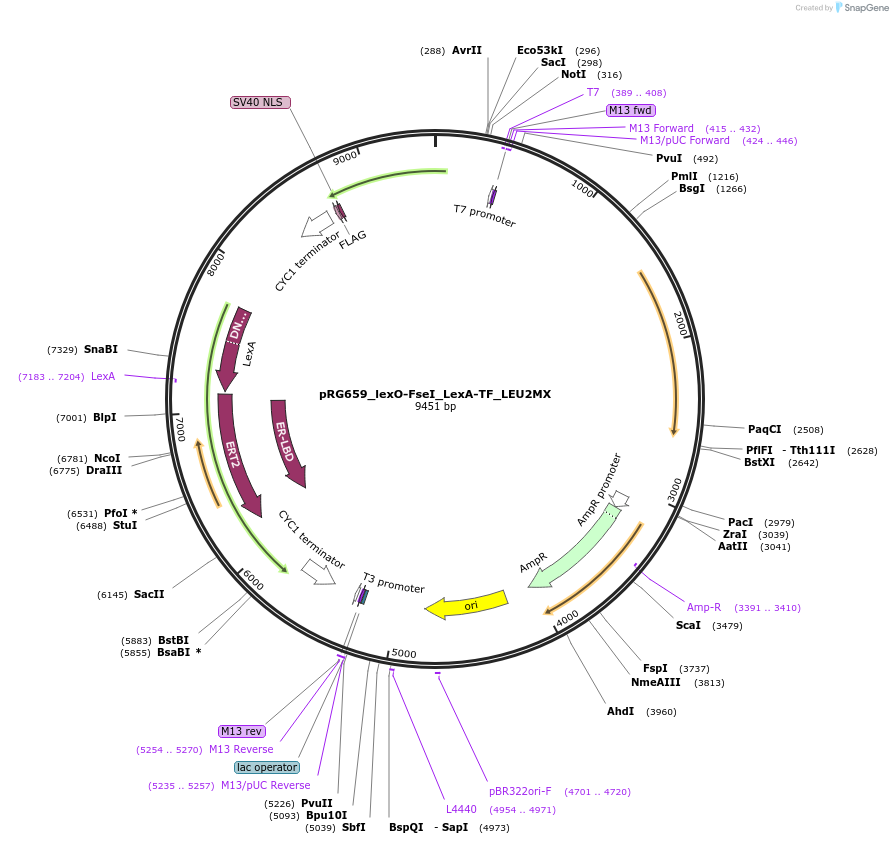
pRG659_lexO-FseI_LexA-TF_LEU2MX
Plasmid#154816PurposeInducible expression of FseI endonuclease in S. cerevisiaeDepositorInsertsFseI endonuclease
LexA-ER-B112
TagsFLAG and NLSExpressionYeastPromoterACT1 and lexO_4-CYC1_coreAvailable SinceOct. 26, 2020AvailabilityAcademic Institutions and Nonprofits only -
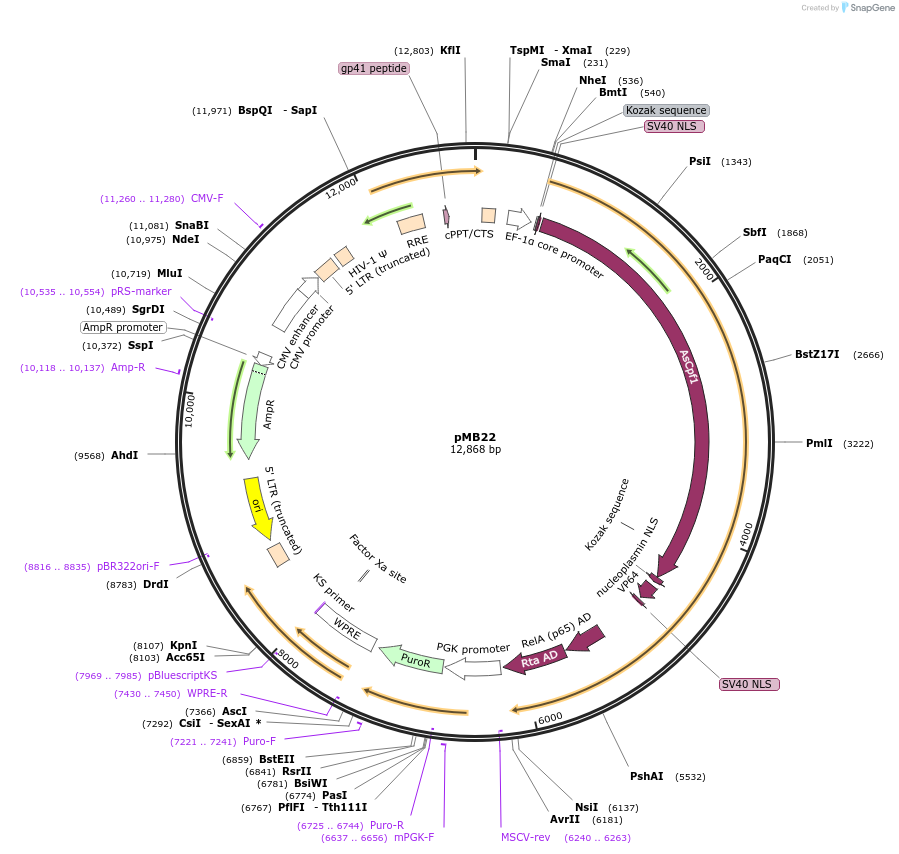
pMB22
Plasmid#122244PurposeLentivirus delivery for stable expression of AsCas12a-VPRDepositorInsertmammalian codon-optimized Acidaminococcus sp. Cas12a fused to VPR activator
UseLentiviralTagsNLS and VPR activatorPromoterEFsAvailable SinceMay 20, 2019AvailabilityAcademic Institutions and Nonprofits only -
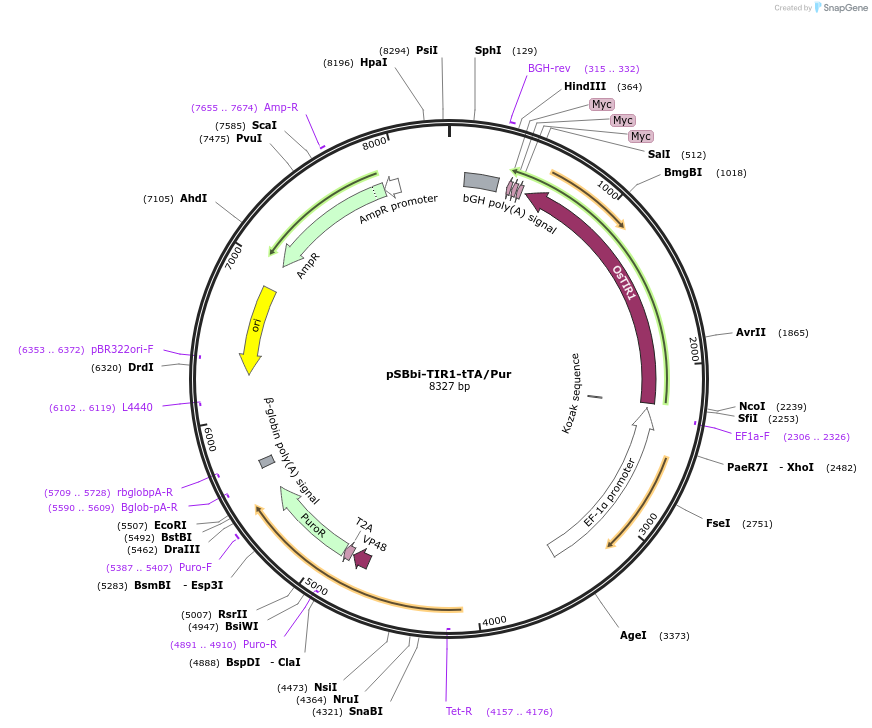
pSBbi-TIR1-tTA/Pur
Plasmid#171683Purposeconstitutive expression of TIR1-myc and tTA in sleeping beauty transposon vector with selection marker (puromycin)DepositorInsertstetracycline-controlled transactivator
Puromycin
EF-1a promoter
ExpressionMammalianPromoterEF-1a promoter and RPBSA promoterAvailable SinceOct. 26, 2021AvailabilityAcademic Institutions and Nonprofits only -
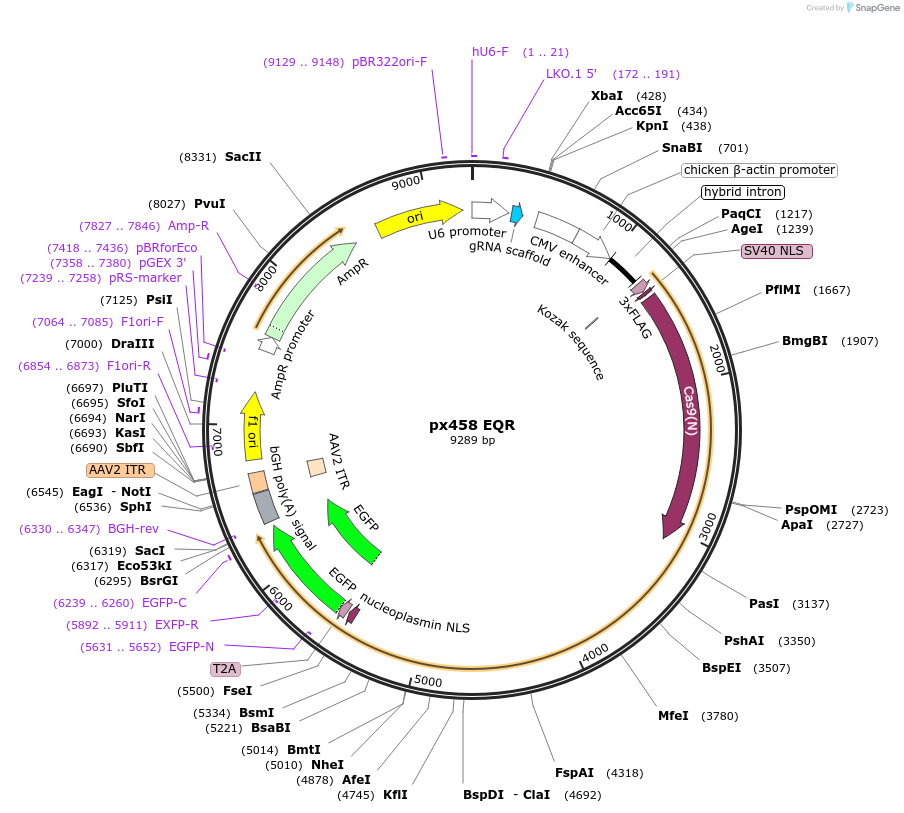
px458 EQR
Plasmid#101731PurposeExpresses a sgRNA and a Cas9 EQR variant that recognizes "NGAG" PAM motifsDepositorInsertSpCas9 EQR
Tags3xFLAG-NLS and NLSExpressionMammalianMutationEQR (D1135E, R1335Q, and T1337R mutations in Cas9)Available SinceOct. 17, 2017AvailabilityAcademic Institutions and Nonprofits only -
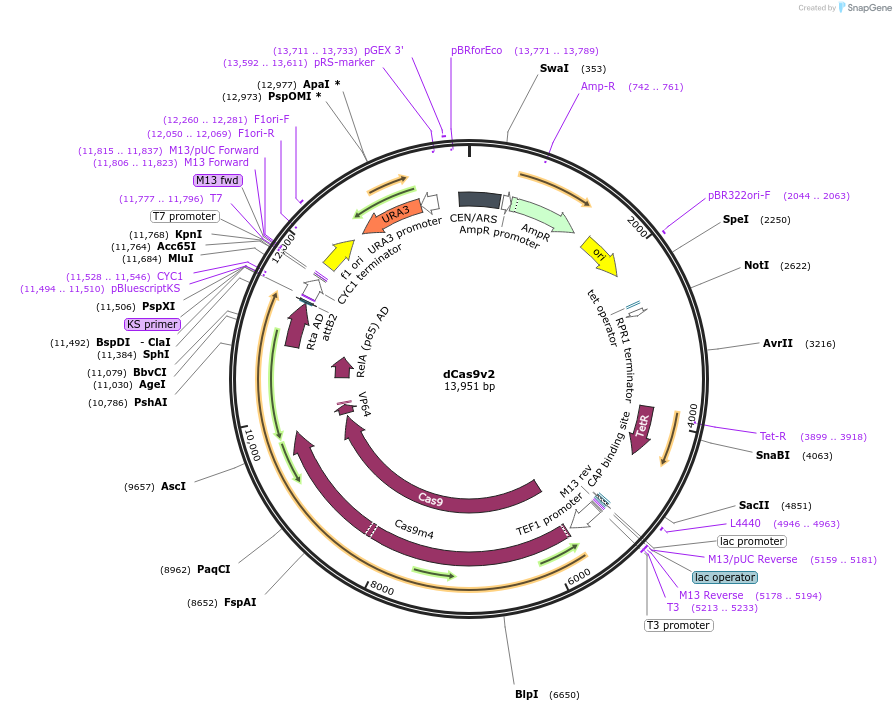
dCas9v2
Plasmid#103140PurposeTetR inducible dCas9-VPR plasmid with pRPR-NotI-tRPR1 Csy4-ready siteDepositorInsertdCas9-VPR
ExpressionYeastMutationRemoved scaffold gRNA from original plasmidAvailable SinceJan. 3, 2018AvailabilityAcademic Institutions and Nonprofits only -
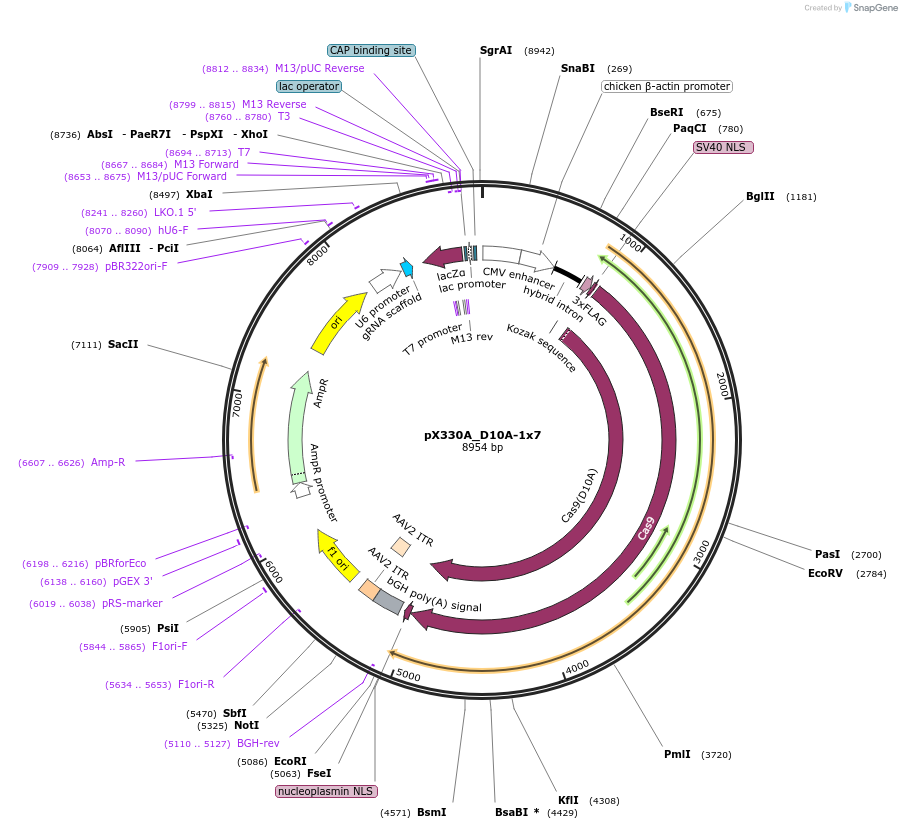
pX330A_D10A-1x7
Plasmid#58777PurposeExpresses Cas9 nickase and gRNADepositorInserthumanized S. pyogenes Cas9 (D10A) nickase
UseCRISPRTags3xFLAGExpressionMammalianMutationD10A nickase-converting mutationPromoterCBhAvailable SinceNov. 7, 2014AvailabilityAcademic Institutions and Nonprofits only