Broad Target Accelerator Plasmid Collections
(Kit #
1000000103, 1000000104, 1000000105
)
Depositing Labs: Jesse Boehm, William Hahn, Matthew Meyerson, and David Root
Clinical sequencing of human cancers has revealed large numbers of rare, poorly characterized mutations. However, the traditional approach of elucidating disease allele function involves detailed one-by-one investigation. While these efforts have been incredibly powerful, the throughput of such investigation does not match the pace of genetic discovery today.
The Broad Institute’s Target Accelerator effort aims to tackle this challenge and make it possible to functionally characterize the cancer genome in a high throughput fashion. A series of pilot projects aims to evaluate whether integrating experimental and computational approaches indeed accelerates the process by which mutant alleles become nominated and provisionally credentialed as targets.
The wild-type and mutant alleles featured in this collection focus on somatic variants found in lung adenocarcinoma as well as variants found across cancer types as part of “pan-cancer” analyses. These plasmids can be purchased individually or as part of a series of kits, collectively called the Broad Target Accelerator Plasmid Collections.
These kits will be sent as bacterial glycerol stocks in a 96-well plate format.
These pilot studies were generously funded through the Cancer Target Discovery and Development (CTD2) network, U.S. NCI grant U01 CA176058, and via the Slim Initiative for Genomic Medicine, a project funded by the Carlos Slim Foundation in Mexico.
Ordering
| Item | Catalog # | Description | Quantity | Price (USD) | ||
|---|---|---|---|---|---|---|
| Kit | 1000000103 | Target Accelerator Pan-Cancer Mutant Collection | 1 | $2238 | Add to Cart | |
| Kit | 1000000104 | Target Accelerator Lung Cancer Mutant Collection | 1 | $1452 | Add to Cart | |
| Kit | 1000000105 | Target Accelerator Genetic Pathway Reference Set | 1 | $1149 | Add to Cart | |
Description
In the Target Accelerator Plasmid Collections, you can find the wild-type and mutant alleles described in the initial Target Accelerator publications, focused on somatic variants found in lung adenocarcinoma as well as variants found across cancer types as part of “pan-cancer” analyses.
Publication List for Kits:
- Systematic Functional Interrogation of Rare Cancer Variants Identifies Oncogenic Alleles (Kim E, et al., 2016)
- Kit #1000000103 - Target Accelerator Pan-Cancer Mutant Collection
- Kit #1000000105 - Target Accelerator Genetic Pathway Reference Set
- High-throughput Phenotyping of Lung Cancer Somatic Mutations (Berger AH, et al., 2016)
- Kit #1000000104 - Target Accelerator Lung Cancer Mutant Collection

Kit #1000000103: Target Accelerator Pan-Cancer Mutant Collection
The Target Accelerator Pan-Cancer Mutant Collection consists of a total of 631 plasmids used in the experiments reported in Kim et al., 2016. Overall, these include 454 mutant clones and 177 corresponding wild-type clones. For a description of the methods used to nominate these mutations and genes, please refer to the methods section of Kim et al., 2016 .
Each clone was derived from the hORFeome 8.1 collection of wild-type human ORFs (Yang X, et al., 2011 ). cDNA clones with perfect or near-perfect matches to REFseq templates were used in Suzuki-based stitching PCR reactions to create mutant variants in the Gateway compatible pDONR223 vector as depicted in the graphic below.
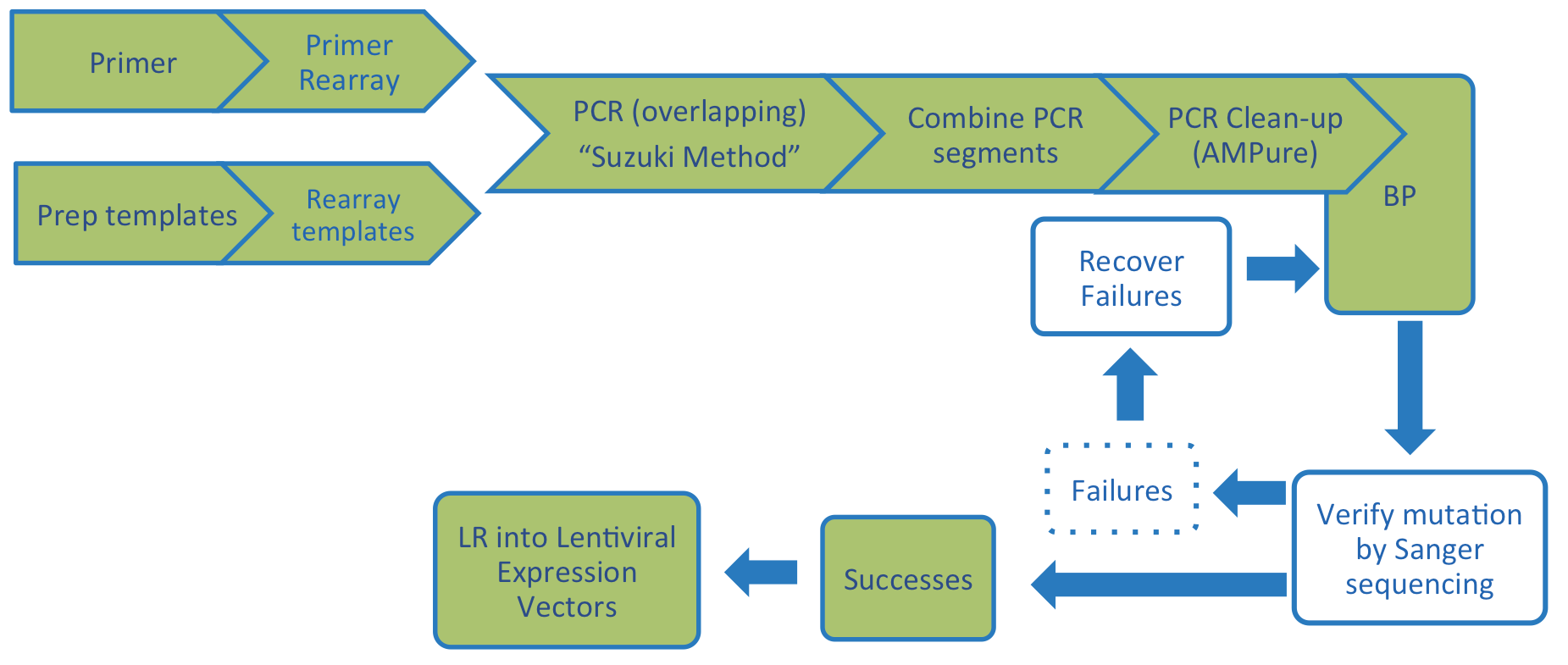
Variants were sequence confirmed using Sanger sequencing, and LR reactions transferred these into pLX-based destination vectors for the experiments described in the papers (see Control Vectors Tab above). Note: the Kim et al. paper also describes the creation of the Target Accelerator Genetic Pathway Reference Set (see Kit #1000000105 below).
pDONR223_CTNNB1_p.G34V, which is part of this collection, is distributed as an individual bacterial stab.
Kit #1000000104: Target Accelerator Lung Cancer Mutant Collection
The Lung Cancer Mutation Collection consists of a total of 307 plasmids used in the experiments reported in Berger et al., 2016. Overall, these include 180 mutant plasmids, 45 wild-type plasmids corresponding to these lung cancer genes and 82 wild-type plasmids serving as expression controls for the L1000 experiments reported in the paper. For more information regarding the rationale and selection process for the clones used, please see the methods section of Berger et al., 2016 .
Mutant plasmids were created and verified as described above for the pan-cancer clones and transferred into the pLX317 destination vector. This also applies to the 5 control plasmids encoding BFP, eGFP, HcRed, Luciferase, and LacZ ORFs listed above.
pDONR223_CTNNB1_p.G34V, which is part of this collection, is distributed as an individual bacterial stab.
Kit #1000000105: Target Accelerator Genetic Pathway Reference Set
The Genetic Pathway Reference Collection consists of 182 wild-type and 19 well-characterized mutant ORFs that represent over 25 cellular signaling pathways and processes. These clones are a high-quality, well annotated collection of genes and mutations that can be used as reference pathway signatures and phenotypes to compare with those derived from cancer alleles.
Control and Lentiviral Expression Vectors
These plasmids are not included in the kits and have to be ordered separately.
A set of 5 control ORFs are available to be used as negative controls for experiments with this collection:
A set of lentiviral vectors is available to express these ORFs in mammalian cells:
To learn more about the 15,000+ ORFs contained in the entry clone collection that was used to generate the mutant clones, as well as the entire lentiviral expression collection (15,000+ ORFs) described in the Yang X, et al. Nature Methods paper, please visit the ORFeome Collaboration Website . Plasmids from the full ORF collections can be obtained from ODF/HCC , DNASU , and Dharmacon .
Plasmid Sequences
Sequences for kit 1000000103 plasmids (CSV format) Sequences for kit 1000000104 plasmids (CSV format) Sequences for kit 1000000105 plasmids (CSV format)Broad Target Accelerator Plasmid Collection References
Target Accelerator References
Systematic functional interrogation of rare cancer variants identifies oncogenic alleles. Kim E, et al. Cancer Discovery, 2016, Jul;6(7):714-26. PubMed PMID 27147599
High-throughput phenotyping of lung cancer somatic mutations. Berger AH, et al. Cancer Cell, 2016, Aug 8;30(2):214-28. PubMed PMID 27478040
Supporting References
Systematic morphological profiling of human gene and allele function via Cell Painting. Rohban MH, et al. Elife, 2017, Mar 18;6. pii: e24060. PubMed PMID 28315521
Progress towards precision functional genomics in cancer. Johannessen CM and Boehm JS. Current Opinion in Systems Biology, 2017, 2:73–82. Link
A public genome-scale lentiviral expression library of human ORFs. Yang X, et al. Nature Methods, 2011, Jun 26;8(8):659-61. PubMed PMID 21706014
How to Cite these Kits
These plasmids were created by your colleagues. Please acknowledge the Principal Investigator, cite the article in which they were created, and include Addgene in the Materials and Methods of your future publications.
For your Materials and Methods section:
“The Target Accelerator Plasmid Collection was a gift from Jesse Boehm (Addgene kit # ).”
Target Accelerator Pan-Cancer Mutant Collection - #1000000103
- Resistance Color Key
Each circle corresponds to a specific antibiotic resistance in the kit plate map wells.
- Inventory
Searchable and sortable table of all plasmids in kit. The Well column links to its kit plate map on the right, as well as lists the plasmid well location in its plate. The Plasmid column links to a plasmid's individual web page.
- Kit Plate Map
96-well plate map for plasmid layout. Hovering over a well reveals the plasmid name, while clicking on a well opens the plasmid page.
Resistance Color Key
| Spectinomycin |
Inventory
Kit Plate Map - #1000000103
Target Accelerator Lung Cancer Mutant Collection - #1000000104
- Resistance Color Key
Each circle corresponds to a specific antibiotic resistance in the kit plate map wells.
- Inventory
Searchable and sortable table of all plasmids in kit. The Well column links to its kit plate map on the right, as well as lists the plasmid well location in its plate. The Plasmid column links to a plasmid's individual web page.
- Kit Plate Map
96-well plate map for plasmid layout. Hovering over a well reveals the plasmid name, while clicking on a well opens the plasmid page.
Resistance Color Key
| Spectinomycin |
Inventory
Kit Plate Map - #1000000104
Target Accelerator Genetic Pathway Reference Set - #1000000105
- Resistance Color Key
Each circle corresponds to a specific antibiotic resistance in the kit plate map wells.
- Inventory
Searchable and sortable table of all plasmids in kit. The Well column links to its kit plate map on the right, as well as lists the plasmid well location in its plate. The Plasmid column links to a plasmid's individual web page.
- Kit Plate Map
96-well plate map for plasmid layout. Hovering over a well reveals the plasmid name, while clicking on a well opens the plasmid page.
Resistance Color Key
| Spectinomycin |


